Abstract
Multiple sclerosis (MS) is a disease of the central nervous system treated with disease-modifying therapies, including the biologic, interferon-β (IFN-β). Up to 60% of IFN-β-exposed MS patients develop abnormal biochemical liver test results1,2, and 1 in 50 experiences drug-induced liver injury3. Since genomic variation contributes to other forms of drug-induced liver injury4,5, we aimed to identify biomarkers of IFN-β-induced liver injury using a two-stage genome-wide association study. The rs2205986 variant, previously linked to differential expression of IRF6, surpassed genome-wide significance in the combined two-stage analysis (P = 2.3 × 10–8, odds ratio = 8.3, 95% confidence interval = 3.6–19.2). Analysis of an independent cohort of IFN-β-treated MS patients identified via electronic medical records showed that rs2205986 was also associated with increased peak levels of aspartate aminotransferase (P = 7.6 × 10–5) and alkaline phosphatase (P = 4.9 × 10-4). We show that these findings may be applicable to predicting IFN-β-induced liver injury, offering insight into its safer use.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout

Similar content being viewed by others
References
Francis, G. S. et al. Hepatic reactions during treatment of multiple sclerosis with interferon-β-1a: incidence and clinical significance. Drug Saf. 26, 815–827 (2003).
Chan, S., Kingwell, E., Oger, J., Yoshida, E. & Tremlett, H. High-dose frequency beta-interferons increase the risk of liver test abnormalities in multiple sclerosis: a longitudinal study. Mult. Scler. 17, 361–367 (2011).
Kowalec, K. et al. Characteristics associated with drug-induced liver injury from interferon beta in multiple sclerosis patients. Expert Opin. Drug Saf. 13, 1305–1317 (2014).
Daly, A. K. et al. HLA-B*5701 genotype is a major determinant of drug-induced liver injury due to flucloxacillin. Nat. Genet. 41, 816–819 (2009).
Singer, J. B. et al. A genome-wide study identifies HLA alleles associated with lumiracoxib-related liver injury. Nat. Genet. 42, 711–714 (2010).
Ostapowicz, G. et al. Results of a prospective study of acute liver failure at 17 tertiary care centers in the United States. Ann. Intern. Med. 137, 947–954 (2002).
Navarro, V. J. & Senior, J. R. Drug-related hepatotoxicity. N. Engl. J. Med. 354, 731–739 (2006).
Spraggs, C. F. et al. HLA-DQA1*02:01 is a major risk factor for lapatinib-induced hepatotoxicity in women with advanced breast cancer. J. Clin. Oncol. 29, 667–673 (2011).
Aithal, G. P. et al. Case definition and phenotype standardization in drug-induced liver injury. Clin. Pharmacol. Ther. 89, 806–815 (2011).
Nelson, M. R. et al. Genome-wide approaches to identify pharmacogenetic contributions to adverse drug reactions. Pharm. J. 9, 23–33 (2009).
GTEx Consortium. The Genotype-Tissue Expression (GTEx) pilot analysis: multitissue gene regulation in humans. Science 348, 648–660 (2015).
Honda, K. & Taniguchi, T. IRFs: master regulators of signalling by Toll-like receptors and cytosolic pattern-recognition receptors. Nat. Rev. Immunol. 6, 644–658 (2006).
Wang, P. X. et al. Interferon regulatory factor 9 is a key mediator of hepatic ischemia/reperfusion injury. J. Hepatol. 62, 111–120 (2015).
Baranzini, S. E. et al. Prognostic biomarkers of IFNβ therapy in multiple sclerosis patients. Mult. Scler. 21, 894–904 (2015).
Mosedale, M. et al. Candidate risk factors and mechanisms for tolvaptan-induced liver injury are identified using a Collaborative Cross approach. Toxicol. Sci. 156, 438–454 (2017).
Lin, Y. et al. Upregulation of interferon regulatory factor 6 promotes neuronal apoptosis after traumatic brain injury in adult rats. Cell Mol. Neurobiol. 36, 27–36 (2016).
Byrnes, V., Afdhal, N., Challies, T. & Greenstein, P. E. Drug induced liver injury secondary to interferon-beta (IFN-β) in multiple sclerosis. Ann. Hepatol. 5, 56–59 (2006).
Lucena, M. I. et al. Glutathione S-transferase m1 and t1 null genotypes increase susceptibility to idiosyncratic drug-induced liver injury. Hepatology 48, 588–596 (2008).
Daly, A. K. et al. Genetic susceptibility to diclofenac-induced hepatotoxicity: contribution of UGT2B7, CYP2C8, and ABCC2 genotypes. Gastroenterology 132, 272–281 (2007).
Kieseier, B. C. The mechanism of action of interferon-β in relapsing multiple sclerosis. CNS Drugs 25, 491–502 (2011).
Chen, P. et al. Carbamazepine-induced toxic effects and HLA-B*1502 screening in Taiwan. N. Engl. J. Med. 364, 1126–1133 (2011).
Milo, R. & Kahana, E. Multiple sclerosis: geoepidemiology, genetics and the environment. Autoimmun. Rev. 9, A387–A394 (2010).
Poser, C. M. et al. New diagnostic criteria for multiple sclerosis: guidelines for research protocols. Ann. Neurol. 13, 227–231 (1983).
Polman, C. H. et al. Diagnostic criteria for multiple sclerosis: 2010 revisions to the McDonald criteria. Ann. Neurol. 69, 292–302 (2011).
Fontana, R. J. et al. Presentation and outcomes with clinically apparent interferon beta hepatotoxicity. Dig. Dis. Sci. 58, 1766–1775 (2013).
Fontana, R. J. et al. Drug-Induced Liver Injury Network (DILIN) prospective study: rationale, design and conduct. Drug Saf. 32, 55–68 (2009).
De Jager, P. L. et al. Meta-analysis of genome scans and replication identify CD6, IRF8 and TNFRSF1A as new multiple sclerosis susceptibility loci. Nat. Genet. 41, 776–782 (2009).
Chang, C. C. et al. Second-generation PLINK: rising to the challenge of larger and richer datasets. Gigascience 4, 7 (2015).
Deelen, P. et al. Genotype harmonizer: automatic strand alignment and format conversion for genotype data integration. BMC Res. Notes 7, 901 (2014).
1000 Genomes Project Consortium. et al. A global reference for human genetic variation. Nature 526, 68–74 (2015).
Jia, X. et al. Imputing amino acid polymorphisms in human leukocyte antigens. PLoS ONE 8, e64683 (2013).
Price, A. L. et al. Principal components analysis corrects for stratification in genome-wide association studies. Nat. Genet. 38, 904–909 (2006).
Pruim, R. J. et al. LocusZoom: regional visualization of genome-wide association scan results. Bioinformatics 26, 2336–2337 (2010).
Roden, D. M. et al. Development of a large-scale de-identified DNA biobank to enable personalized medicine. Clin. Pharmacol. Ther. 84, 362–369 (2008).
Davis, M. F. et al. Automated extraction of clinical traits of multiple sclerosis in electronic medical records. J. Am. Med. Inform. Assoc. 20, e334–e340 (2013).
Guo, Y. et al. Illumina human exome genotyping array clustering and quality control. Nat. Protoc. 9, 2643–2662 (2014).
International Multiple Sclerosis Genetics Consortium. et al. Genetic risk and a primary role for cell-mediated immune mechanisms in multiple sclerosis. Nature 476, 214–219 (2011).
Lucena, M. I. et al. Phenotypic characterization of idiosyncratic drug-induced liver injury: the influence of age and sex. Hepatology 49, 2001–2009 (2009).
Robin, X. et al. pROC: an open-source package for R and S+ to analyze and compare ROC curves. BMC Bioinforma. 12, 77 (2011).
DeLong, E. R., DeLong, D. M. & Clarke-Pearson, D. L. Comparing the areas under two or more correlated receiver operating characteristic curves: a nonparametric approach. Biometrics 44, 837–845 (1988).
Acknowledgements
We gratefully acknowledge the participation of all patients and families who took part in this study across Canada, Europe, and the USA. This work would not have been possible without their generous participation. We also acknowledge the contributions of the Canadian Pharmacogenomics Network for Drug Safety Consortium (C. Hildebrand, A. Borrie, T. Wong, T. Bendyshe-Walton, N. Massah, F. Miao, M. Higginson, M. Staub, K. Shaw, B. Malkin, and J. Stortz), Winnipeg Health Sciences Centre MS Clinic (N. Hall and B. Stranger), London Health Sciences Centre MS Clinic (H. Rosehart), Dalhousie MS Clinic (K. Stadnyk, K. Sabourin, and B. DeCoste), and Centre Hospitalier de L’Université de Montréal (É. Roger). We thank all the MS clinic neurologists who contributed to the study through patient examination and data collection. This study makes use of data generated by the Wellcome Trust Case Control Consortium. A full list of the investigators who contributed to the generation of the data is available from www.wtccc.org.uk. Funding for the original Wellcome Trust Case Control Consortium project was provided by the following Wellcome Trust awards: 076113, 085475 and 090355. This investigation was supported by a pilot grant from the British Columbia Clinical Genomics Network (to H.T. (the principal investigator)). Doctoral studentships provided additional funding for K.K. from the Canadian Institutes of Health Research (CIHR), CIHR Drug Safety and Effectiveness Cross-Disciplinary Training Program (DSECT), MS Society of Canada, University of British Columbia, and Canadian Pharmacogenomics Network for Drug Safety. G.E.B.W. received fellowships from the CIHR and CIHR DSECT programs. B.I.D. received stipends from the British Columbia Children’s Hospital Research Institute, CIHR DSECT, CIHR, and the Michael Smith Foundation for Health Research during the period of this study. The Drug-Induced Liver Injury Network is supported by the National Institute of Diabetes and Digestive and Kidney Diseases of the National Institutes of Health (NIH) as part of the Cooperative Agreements (U01s) project under grants U01-DK065176 (Duke), U01-DK065201 (UNC), U01-DK065184 (Michigan), U01-DK065211 (Indiana), U01DK065193 (UConn), U01-DK065238 (UCSF/CPMC), U01-DK083023 (UTSW), U01-DK083027 (TJH/UPenn), U01-DK082992 (Mayo), U01-DK083020 (USC), and U01-DK100928 (Icahn). Additional funding was provided by CTSA grants UL1 RR025761 (Indiana), UL1 RR025747 (UNC), and UL1 UL1 RR024986 (UMich), and the Intramural Research Program of the National Cancer Institute. Vanderbilt University Medical Center BioVU is supported by institutional funding and Vanderbilt CTSA grant ULTR000445 from NCATS/NIH. Genome-wide genotyping was funded by P50GM115305 from NIGMS. H.T. and B.C.C. had full access to all the data in the study, and take full responsibility for the integrity of the data and the accuracy of the data analysis. The study funders had no role in the study design, data collection, data analysis, data interpretation, or writing of the report.
Author information
Authors and Affiliations
Contributions
K.K. performed the baseline epidemiological analyses of the clinical characteristics and genetic association analyses, compiled the corresponding sections, performed literature reviews, and wrote the first draft of the manuscript. G.E.B.W. performed whole-genome imputation association analyses and population control analyses, and compiled the corresponding sections. B.I.D. performed the HLA imputation and related association analyses, and compiled the corresponding sections. M.F.D., J.C.D., G.E.B.W., and B.C.C. performed the analysis and interpretation of the BioVU cohorts. K.K., G.E.B.W., B.I.D., F.A., A.P.B., E.K., and C.J.D.R. provided further data interpretation. C.J.D.R., B.C.C., and H.T. were responsible for study design and interpretation of the data. K.K., E.M.Y., A.T., R.A.M., M.K., T.L.C., P.D., N.C., M.W., P.H., Z.X., and P.L.D.J. were responsible for patient recruitment. All authors contributed to drafting and revising the work for important intellectual content, and made substantial contributions to the concept and design of the study and analysis and interpretation of the data.
Corresponding authors
Ethics declarations
Competing interests
K.K. has a consulting agreement with Emerald Lake Safety. E.M.Y. is an investigator of clinical trials sponsored by Gilead, Merck, Vertex, Hoffman LaRoche, Abbvie, Janssen, Boeringher Ingelheim, Intercept, and Genfit. He has also received honoraria for continuing medical education lectures from Merck Canada and Gilead Canada, spoken at Advisory Board Meetings of Boeringher Ingelheim Canada, Hoffman LaRoche Canada, Abbvie Canada, and Celgene Canada (for which he received an honorarium), and is a member of the Gilead Canada compassionate release program adjudication committee, for which he received an honorarium. A.T. has received grant support from Hoffman la Roche, Sanofi Genzyme, Chugai, Novartis, and Biogen, as well as performing consultancy work for Biogen, EMD Serono, Hoffman la Roche, Sanofi Genzyme, and Teva Neuroscience. R.A.M. has conducted clinical trials for Sanofi-Aventis, and receives research funding from the CIHR, National MS Society, MS Society of Canada, Research Manitoba, Consortium of MS Centers, Crohn’s and Colitis Canada, and Waugh Family Chair in Multiple Sclerosis. In the past two years, M.K. or M.K.’s institution (London Health Sciences Centre and Department of Neurological Sciences, Western University) has received research support or grants from CIHR. T.L.C. was a principal investigator for a research study for Biogen Idec Canada, has received honoraria for lectures from EMD Merck Serono Canada, Biogen Idec Canada, Teva Canada Innovation, and Genzyme, has participated in or spoken at Advisory Board Meetings of EMD Merck Serono, Teva Canada Innovation, Biogen Idec Canada, Genzyme, and Novartis (for which an honorarium and travel support were received in exchange for attendance). P.D. has received support as a consultant, membership on advisory councils, and grants from the pharmaceutical industry, and is funded by the MS Society of Canada and CIHR. N.C. has consulting agreements with Abbvie, Lilly, DS Biopharma (Afimmune), Tobira (Allergan), NuSirt, Celgene, Axovant, Shire, and Madrigal, and has received research grants from Cumberland, Galectin, and Intercept. M.W. and P.H. receive funding from the Swedish Research Council (Medicine 521-2011-2440 and 521-2014-3370) and Clinical Research Support (ALF) at Uppsala University. M.W. has also received grants from the Swedish Heart and Lung Foundation (20120557 and 20140291). Z.X. was a recipient of the Clinician Scientist Development Award from the National Multiple Sclerosis Society and American Academy of Neurology, and is supported by NINDS NIH K08-NS079493 and NINDS R01 NS098023. P.L.D.J. has received research funding from Sanofi/Genzyme and Biogen, and has consulted for Teva Pharmaceuticals and Sanofi/Genzyme. C.J.D.R. receives funding support from the CIHR, Canadian Foundation for Innovation, Canadian Hearing Foundation, BC Children’s Hospital Foundation, Child & Family Research Institute, Canadian Gene Cure Foundation, Teva Pharmaceutical Industries, Genome BC, and CIHR Drug Safety and Effectiveness Network. H.T. is the Canada Research Chair for Neuroepidemiology and Multiple Sclerosis, and currently receives research support from the National MS Society, CIHR, MS Society of Canada, and MS Scientific Research Foundation. In the past five years, H.T. has received support from the Multiple Sclerosis Society of Canada (Don Paty Career Development Award), Michael Smith Foundation for Health Research (Scholar Award), and UK MS Trust. H.T. has received speaker honoraria and/or travel expenses to attend conferences from the Consortium of MS Centres (2013), National MS Society (2014 and 2016), ECTRIMS (2013–2018), Biogen Idec (2014), and American Academy of Neurology (2013–2016). All speaker honoraria were either declined or donated to an MS charity or unrestricted grant for use by H.T.’s research group. B.C.C. currently receives research funding from the CIHR, Genome Canada, Genome British Columbia, and British Columbia Children’s Hospital Research Institute (Vancouver, Canada), and has previously received matching funds support for Genome Canada funding from Pfizer Canada (unrestricted).
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 Genetic ancestry determined by principal-components analysis of patients genotyped on the Illumina MEGA array.
The first two principal components are plotted to visualize the distribution of genetic ancestry. Each point represents one patient, colored by their self-reported ancestry (n = 170 in stage 1 and n = 10 in stage 2). The circled patients correspond to those that were considered to be of European genetic ancestry for analyses.
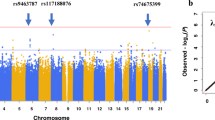
Supplementary Figure 2 Genome-wide scan of interferon-β-induced liver injury in multiple sclerosis.
a, Manhattan plot showing the observed distribution of –log10 (P values) against the chromosomal location (GRCh37) of genetic variants in the imputed stage 1 cohort of European genetic ancestry (n = 151). P values are from logistic regression employing an additive model with adjustment for MS disease course at the start of IFN-β, with the variant most strongly associated with liver injury from IFN-β (rs2205986) circled. The red horizontal line (P = 1.0 × 10–6) indicates the screening threshold employed to prioritize variants for subsequent stage 2 analyses. b, Quantile–quantile plot showing the distributions of adjusted observed –log10 (P values) plotted against expected –log10 (P values) in the stage 1 cohort of patients with European genetic ancestry (n = 151). A genomic inflation factor (λGC) of 1.06 indicates no obvious population stratification. The blue lines indicate the 95% confidence interval, and P values were obtained in the same manner as stated in a.
Supplementary Figure 3 Box plots of peak biochemical liver test results from the BioVU multiple sclerosis cohort, stratified by rs2205986 genotype status.
a–d, Peak alkaline phosphatase (ALP) (a) and aspartate aminotransferase (AST) (b) values were statistically significant (P = 4.9 × 10–4 for ALP and P = 7.6 × 10–5 for AST) with carriers displaying elevated levels, while no association was detected for mean peak alanine aminotransferase (ALT) (c) and total bilirubin (TBIL) (d) values. Each box plot summarizes the relevant biochemical liver test result by rs2205986 genotype for MS patients (n = 87) exposed to interferon-β. Biochemical liver enzyme test results are either displayed as units per liter (U/l, i.e., ALP, ALT and AST) or milligrams per deciliter (mg/dl, i.e., TBIL). These plots display the interquartile range, with the medians depicted as black lines and whiskers indicating 1.5 times the interquartile range. Dashed lines indicate the upper limit of normal values for the respective biochemical liver test. Linear regression was performed using an additive genetic model for highest values, adjusted for age at biochemical liver test date, sex, and the first two principal components.
Supplementary Figure 4 Multi-tissue eQTL analyses for rs2205986 and IRF6 from the GTEx Project.
m values represent the posterior probability for an eQTL effect (PLoS Genet. 8, e1002555, 2012), and tissues with values > 0.9 (i.e., cerebellum, tibial nerve, aorta, cerebellar hemisphere, whole blood and frontal cortex) are predicted to be eQTLs.
Supplementary Figure 5 Receiver operating characteristic (ROC) curve of the clinical and clinical/genomic model for predicting interferon-β-induced liver injury in the combined cohort.
The area under the curves and corresponding 95% confidence intervals are presented for the combined cohort (n = 182). Clinical variables were age at start of IFN-β, IFN-β product and sex. The genomic variable included was rs2205986. The ROC curves of these two prediction models were compared using DeLong’s test.
Supplementary Figure 6 Genotyping cluster plots of rs2205986, by genotyping method.
a, Genotyped by the Illumina MEGA array, plot generated using Illumina GenomeStudio software (n = 170 stage 1 and 10 stage 2 samples, all ancestries). One sample that presented as a G/G genotype was removed during quality control owing to low array call rate. b, TaqMan rs2205986 allelic discrimination plot generated using Thermo Fisher TaqMan Genotyper software (Methods) in stage 1 of European-ancestry patients (n = 151).
Supplementary Figure 7 Principal-component analysis of disease-matched population controls.
Multiple sclerosis population controls from the United States ascertained to be of northern European genetic ancestry (blue points, n = 1,319) were included. Cases used in stage 1 analyses (red points) and excluded southern European samples (gray points) are included for reference.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–7 and Supplementary Tables 1–5
Rights and permissions
About this article
Cite this article
Kowalec, K., Wright, G.E.B., Drögemöller, B.I. et al. Common variation near IRF6 is associated with IFN-β-induced liver injury in multiple sclerosis. Nat Genet 50, 1081–1085 (2018). https://doi.org/10.1038/s41588-018-0168-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41588-018-0168-y
This article is cited by
-
SWEDEGENE—a Swedish nation-wide DNA sample collection for pharmacogenomic studies of serious adverse drug reactions
The Pharmacogenomics Journal (2020)
-
Pharmacogenomics of COVID-19 therapies
npj Genomic Medicine (2020)
-
Drug-induced liver injury
Nature Reviews Disease Primers (2019)
-
Drug-Induced Liver Injury: Highlights of the Recent Literature
Drug Safety (2019)
-
Replication study of GWAS risk loci in Greek multiple sclerosis patients
Neurological Sciences (2019)