Abstract
Efficient termination is required for robust gene transcription. Eukaryotic organisms use a conserved exoribonuclease-mediated mechanism to terminate the mRNA transcription by RNA polymerase II (Pol II)1,2,3,4,5. Here we report two cryogenic electron microscopy structures of Saccharomyces cerevisiae Pol II pre-termination transcription complexes bound to the 5′-to-3′ exoribonuclease Rat1 and its partner Rai1. Our structures show that Rat1 displaces the elongation factor Spt5 to dock at the Pol II stalk domain. Rat1 shields the RNA exit channel of Pol II, guides the nascent RNA towards its active centre and stacks three nucleotides at the 5′ terminus of the nascent RNA. The structures further show that Rat1 rotates towards Pol II as it shortens RNA. Our results provide the structural mechanism for the Rat1-mediated termination of mRNA transcription by Pol II in yeast and the exoribonuclease-mediated termination of mRNA transcription in other eukaryotes.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
Atomic coordinates have been deposited in the PDB under the accession codes 8JCH (Pol II Rat1–PTTC1) and 8K5P (Pol II Rat1–PTTC2). Corresponding cryo-EM density maps have been deposited in the Electron Microscopy Data Bank under the accession codes EMD-36162 (Pol II Rat1–PTTC1) and EMD-36908 (Pol II Rat1–PTTC2). Source data are provided with this paper.
References
Kim, M. et al. The yeast Rat1 exonuclease promotes transcription termination by RNA polymerase II. Nature 432, 517–522 (2004).
West, S., Gromak, N. & Proudfoot, N. J. Human 5′ -> 3′ exonuclease Xrn2 promotes transcription termination at co-transcriptional cleavage sites. Nature 432, 522–525 (2004).
Luo, W. & Bentley, D. A ribonucleolytic rat torpedoes RNA polymerase II. Cell 119, 911–914 (2004).
Fong, N. et al. Effects of transcription elongation rate and Xrn2 exonuclease activity on RNA polymerase II termination suggest widespread kinetic competition. Mol. Cell 60, 256–267 (2015).
Krzyszton, M. et al. Defective XRN3-mediated transcription termination in Arabidopsis affects the expression of protein-coding genes. Plant J. 93, 1017–1031 (2018).
Proudfoot, N. J. Transcriptional termination in mammals: stopping the RNA polymerase II juggernaut. Science 352, aad9926 (2016).
Girbig, M., Misiaszek, A. D. & Muller, C. W. Structural insights into nuclear transcription by eukaryotic DNA-dependent RNA polymerases. Nat. Rev. Mol. Cell Biol. 23, 603–622 (2022).
Porrua, O. & Libri, D. Transcription termination and the control of the transcriptome: why, where and how to stop. Nat. Rev. Mol. Cell Biol. 16, 190–202 (2015).
Eaton, J. D. & West, S. Termination of transcription by RNA polymerase II: BOOM! Trends Genet. 36, 664–675 (2020).
Richard, P. & Manley, J. L. Transcription termination by nuclear RNA polymerases. Genes Dev. 23, 1247–1269 (2009).
Kumar, A., Clerici, M., Muckenfuss, L. M., Passmore, L. A. & Jinek, M. Mechanistic insights into mRNA 3′-end processing. Curr. Opin. Struc. Biol. 59, 143–150 (2019).
Lunde, B. M. et al. Cooperative interaction of transcription termination factors with the RNA polymerase II C-terminal domain. Nat. Struct. Mol. Biol. 17, 1195–1201 (2010).
Carminati, M. et al. A direct interaction between CPF and RNA Pol II links RNA 3′ end processing to transcription. Mol. Cell 83, 4461–4478 (2023).
Brannan, K. et al. mRNA decapping factors and the exonuclease Xrn2 function in widespread premature termination of RNA polymerase II transcription. Mol. Cell 46, 311–324 (2012).
Nojima, T. et al. Mammalian NET-Seq reveals genome-wide nascent transcription coupled to RNA processing. Cell 161, 526–540 (2015).
Jimeno-Gonzalez, S., Haaning, L. L., Malagon, F. & Jensen, T. H. The yeast 5′-3′ exonuclease Rat1p functions during transcription elongation by RNA polymerase II. Mol. Cell 37, 580–587 (2010).
Cortazar, M. A. et al. Xrn2 substrate mapping identifies torpedo loading sites and extensive premature termination of RNA pol II transcription. Genes Dev. 36, 1062–1078 (2022).
Kim, M., Ahn, S. H., Krogan, N. J., Greenblatt, J. F. & Buratowski, S. Transitions in RNA polymerase II elongation complexes at the 3′ ends of genes. EMBO J. 23, 354–364 (2004).
Zhang, Z., Fu, J. & Gilmour, D. S. CTD-dependent dismantling of the RNA polymerase II elongation complex by the pre-mRNA 3′-end processing factor, Pcf11. Genes Dev. 19, 1572–1580 (2005).
Zhang, H., Rigo, F. & Martinson, H. G. Poly(A) signal-dependent transcription termination occurs through a conformational change mechanism that does not require cleavage at the poly(A) site. Mol. Cell 59, 437–448 (2015).
Logan, J., Falck-Pedersen, E., Darnell, J. E. Jr & Shenk, T. A poly(A) addition site and a downstream termination region are required for efficient cessation of transcription by RNA polymerase II in the mouse beta maj-globin gene. Proc. Natl Acad. Sci. USA 84, 8306–8310 (1987).
Eaton, J. D., Francis, L., Davidson, L. & West, S. A unified allosteric/torpedo mechanism for transcriptional termination on human protein-coding genes. Genes Dev. 34, 132–145 (2020).
Luo, W., Johnson, A. W. & Bentley, D. L. The role of Rat1 in coupling mRNA 3′-end processing to transcription termination: implications for a unified allosteric–torpedo model. Genes Dev. 20, 954–965 (2006).
Cortazar, M. A. et al. Control of RNA Pol II speed by PNUTS-PP1 and Spt5 dephosphorylation facilitates termination by a “sitting duck torpedo” mechanism. Mol. Cell 76, 896–908 e894 (2019).
Xue, Y. et al. Saccharomyces cerevisiae RAI1 (YGL246c) is homologous to human DOM3Z and encodes a protein that binds the nuclear exoribonuclease Rat1p. Mol. Cell. Biol. 20, 4006–4015 (2000).
Xiang, S. et al. Structure and function of the 5′→3′ exoribonuclease Rat1 and its activating partner Rai1. Nature 458, 784–788 (2009).
Mayer, A. et al. Uniform transitions of the general RNA polymerase II transcription complex. Nat. Struct. Mol. Biol. 17, 1272–1278 (2010).
Narain, A. et al. Targeted protein degradation reveals a direct role of SPT6 in RNAPII elongation and termination. Mol. Cell 81, 3110–3127 (2021).
Baejen, C. et al. Genome-wide analysis of RNA polymerase II termination at protein-coding genes. Mol. Cell 66, 38–49 (2017).
Jinek, M., Coyle, S. M. & Doudna, J. A. Coupled 5′ nucleotide recognition and processivity in Xrn1-mediated mRNA decay. Mol. Cell 41, 600–608 (2011).
Stevens, A. & Poole, T. L. 5′-exonuclease-2 of Saccharomyces cerevisiae. Purification and features of ribonuclease activity with comparison to 5′-exonuclease-1. J. Biol. Chem. 270, 16063–16069 (1995).
Ehara, H. et al. Structure of the complete elongation complex of RNA polymerase II with basal factors. Science 357, 921–924 (2017).
Parua, P. K. et al. A Cdk9–PP1 switch regulates the elongation–termination transition of RNA polymerase II. Nature 558, 460–464 (2018).
Kecman, T. et al. Elongation/termination factor exchange mediated by PP1 phosphatase orchestrates transcription termination. Cell Rep. 25, 259–269 (2018).
Grosso, A. R., de Almeida, S. F., Braga, J. & Carmo-Fonseca, M. Dynamic transitions in RNA polymerase II density profiles during transcription termination. Genome Res. 22, 1447–1456 (2012).
Orozco, I. J., Kim, S. J. & Martinson, H. G. The poly(A) signal, without the assistance of any downstream element, directs RNA polymerase II to pause in vivo and then to release stochastically from the template. J. Biol. Chem. 277, 42899–42911 (2002).
Gromak, N., West, S. & Proudfoot, N. J. Pause sites promote transcriptional termination of mammalian RNA polymerase II. Mol. Cell. Biol. 26, 3986–3996 (2006).
James, K., Gamba, P., Cockell, S. J. & Zenkin, N. Misincorporation by RNA polymerase is a major source of transcription pausing in vivo. Nucleic Acids Res. 45, 1105–1113 (2017).
Molodtsov, V., Wang, C., Firlar, E., Kaelber, J. T. & Ebright, R. H. Structural basis of Rho-dependent transcription termination. Nature 14, 367–374 (2023).
Murayama, Y. et al. Structural basis of the transcription termination factor Rho engagement with transcribing RNA polymerase from Thermus thermophilus. Sci. Adv. 9, eade7093 (2023).
Chengyuan, W. et al. Structural basis of archaeal FttA-dependent transcription termination. Preprint at bioRxiv https://doi.org/10.1101/2023.08.09.552649 (2023).
Fianu, I. et al. Structural basis of integrator-mediated transcription regulation. Science 374, 883–887 (2021).
Zheng, H. et al. Structural basis of INTAC-regulated transcription. Protein Cell 14, 698–702 (2023).
Higo, T. et al. Development of a hexahistidine-3x FLAG-tandem affinity purification method for endogenous protein complexes in Pichia pastoris. J. Struct. Funct. Genomics 15, 191–199 (2014).
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017).
Zhang, K. Gctf: real-time CTF determination and correction. J. Struct. Biol. 193, 1–12 (2016).
Zivanov, J. et al. New tools for automated high-resolution cryo-EM structure determination in RELION-3. Elife 7, e42166 (2018).
Punjani, A., Rubinstein, J. L., Fleet, D. J. & Brubaker, M. A. cryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290–296 (2017).
Xu, J. et al. Structural basis for the initiation of eukaryotic transcription-coupled DNA repair. Nature 551, 653–657 (2017).
Farnung, L., Ochmann, M., Engeholm, M. & Cramer, P. Structural basis of nucleosome transcription mediated by Chd1 and FACT. Nat. Struct. Mol. Biol. 28, 382–387 (2021).
Jumper, J. et al. Highly accurate protein structure prediction with AlphaFold. Nature 596, 583–589 (2021).
Pettersen, E. F. et al. UCSF Chimera-a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D 60, 2126–2132 (2004).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D 66, 213–221 (2010).
Ehara, H., Kujirai, T., Shirouzu, M., Kurumizaka, H. & Sekine, S. I. Structural basis of nucleosome disassembly and reassembly by RNAPII elongation complex with FACT. Science 377, eabp9466 (2022).
Acknowledgements
The work was supported by the National Key Research and Development Program of China 2018YFA0900701 (Y. Zhang) and the Basic Research Zone Program of Shanghai JCYJ-SHFY-2022-012 (Y. Zhang). We thank L. Kong, F. Wang, G. Li, J. Duan at electron microscopy system at the National Facility for Protein Science in Shanghai (NFPS), M. Zhang at the electron microscopy center in Interdisciplinary Research Center on Biology and Chemistry (IRCBC), and staffs X. Men, F. Liu, and S. Wang at electron microscopy center at Shuimu Biosciences in Hangzhou for providing technical support and assistance in cryo-EM data collection. We thank M. Zhang and Z. Zhang at the core facility of the Center for Excellence in Molecular Plant Sciences (CEMPS) for technical support and assistance in screening cryo-EM samples. The S. cerevisiae BJ2168 strain was a gift from G. Cai. The pESC-his vector was a gift from Z. Zhou.
Author information
Authors and Affiliations
Contributions
Y. Zeng collected the cryo-EM data, solved the cryo-EM structures and carried out biochemical experiments. H.-W.Z. purified yeast Pol II. X.-X.W. assisted in structure determination. Y. Zhang designed experiments, analysed data and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature thanks Seth Darst and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 The assembly of the Pol II Rat1-PTTC1 and Pol II Rat1-PTTC2.
a-b, Elution peaks and SDS-PAGE results of Pol II Rat1-PTTC1 from a size-exclusion chromatography. c-d, Elution peaks and SDS-PAGE result of Pol II Rat1-PTTC2 from a size-exclusion chromatography. Asterisks indicate degradation products of Rpb1. e, The Pol II release result shows the Rat1-Rai1 complex releases Pol II from the elongation complex. Three biological repeats were done, see Supplementary Fig. 1. f, The RNA cleavage result shows the Rat1-Rai1 complex has exoribonuclease activity. Three biological repeats were done, see Supplementary Fig. 1.
Extended Data Fig. 2 Cryo-EM data processing of Pol II Rat1-PTTC1.
a, The flowchart of data processing for Pol II Rat1-PTTC1. b, Representative cryo-EM micrograph. c, Representative 2D class averages. d-f, Gold-standard FSC curves of map 1a, map 1b, and map 1c. g, The 3D FSC plot. The dotted line represents 0.143 cutoff of the global FSC curve. h, Nominal 2.7 Å-resolution cryo-EM reconstruction of map 2a filtered by local resolution. i-j, Map classified from Pol II Rat1-PTTC1. Map 1a represents the Pol II Rat1-PTTC1. Map 1c represents Pol II Spt4-Spt5 bound TEC. The Rat1-Rai1 map is colored green, and the Spt4-Spt5 map is colored blue.
Extended Data Fig. 3 Cryo-EM data processing of Pol II Rat1-PTTC2.
a, The flowchart of data processing for Pol II Rat1-PTTC2. b-c, Gold-standard FSC curves of map 2a, map 2b. d, The 3D FSC plot. The dotted line represents the 0.143 cutoff of the global FSC curve. e, Nominal 2.8 Å-resolution cryo-EM reconstruction of map 2a filtered by local resolution. f, Local refined map (map 2b) of Rat1, Rai1, Rpb4-Rpb7, and structure elements around the RNA exit channel of Pol II.
Extended Data Fig. 4 The interface between Rat1 and Rai1 in Pol II Rat1-PTTC1.
a, Overview of the interface between Rat1 and Rai1. The dashed boxes highlight the structure elements of Rat1 that contact the two concaves of Rai1. Green represents the Rat1 regions not observed in the crystal structure of S. pombe Rat1-Rai1 complex. b, Structural comparison of the Rat1-Rai1 complex in Pol II Rat1-PTTC1 and the crystal structure of S. pombe Rat1-Rai1 complex (PDB:3FQD26). c-d, The cryo-EM maps (map 1b) of interfaces 1 and 2.
Extended Data Fig. 5 The cryo-EM map of interfaces between Pol II and the Rat1-Rai1 complex in the structure of Pol II Rat1-PTTC1.
The view orientation and cartoon color are the same as in Fig. 2d. The cryo-EM map is colored in blue.
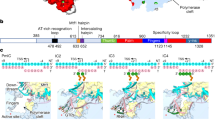
Extended Data Fig. 6 Sequence alignment of Rat1 homologs.
Sequence alignment shows that Rat1 residues in arms 1 and 2 contacting Pol II are generally conserved (red circles). Other Rat1 residues contacting Pol II are shown in blue circles.
Extended Data Fig. 8 Rat1 can be recruited to Pol II independent of RNA.
a-b, Elution peaks and SDS-PAGE result of Pol II-RR from a size-exclusion chromatography without a nucleic-acid scaffold. Two biological repeats were done, see source data and Supplementary Fig. 1. c, The native-page result shows that the Rat1-Rai1(RR) complex slows the migration of Pol II in a concentration-dependent way, suggesting that the Rat1-Rai1 complex forms a stable complex with Pol II. Three biological repeats were done, see Supplementary Fig. 1. d, The native-page result shows that removing arms 1 and 2 weakens the interaction with Pol II. Three biological repeats were done, see Supplementary Fig. 1. e. The flag pull-down result shows Spt5 does not affect the interaction between Rat1-Rai1 and Pol II. Three biological repeats were done, see Supplementary Fig. 1.
Extended Data Fig. 9 The interaction of nascent RNA with Rat1 in Pol II Rat1-PTTC1.
a, The interaction between the 5′ phosphate groups of the first two nucleotides with the active site of Rat1. Black dash, the coordination bonds. Blue dash, the H-bonds. b, The interaction of nascent RNA with Rat1 and Pol II. The cryo-EM map for the RNA-contacting residues is shown in blue. C, The modeled triphosphate group of the RNA 5′ terminus clashes with Rat1. Left, RNA with a monophosphate group in the structure of Pol II Rat1-PTTC1; right, RNA with a modeled triphosphate group in the structure of Pol II Rat1-PTTC1.
Supplementary information
Supplementary Fig. 1
Uncropped gels from Extended Data Figs. 1 and 8.
Source data
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Zeng, Y., Zhang, HW., Wu, XX. et al. Structural basis of exoribonuclease-mediated mRNA transcription termination. Nature 628, 887–893 (2024). https://doi.org/10.1038/s41586-024-07240-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-024-07240-3
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.