Abstract
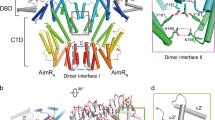
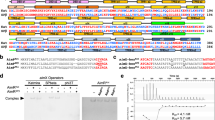
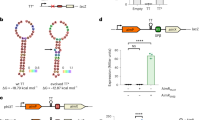
A bacteriophage can replicate and release virions from a host cell in the lytic cycle or switch to a lysogenic process in which the phage integrates itself into the host genome as a prophage. In Bacillus cells, some types of phages employ the arbitrium communication system, which contains an arbitrium hexapeptide, the cellular receptor AimR and the lysogenic negative regulator AimX. This system controls the decision between the lytic and lysogenic cycles. However, both the mechanism of molecular recognition between the arbitrium peptide and AimR and how downstream gene expression is regulated remain unknown. Here, we report crystal structures for AimR from the SPbeta phage in the apo form and the arbitrium peptide-bound form at 2.20 Å and 1.92 Å, respectively. With or without the peptide, AimR dimerizes through the C-terminal capping helix. AimR assembles a superhelical fold and accommodates the peptide encircled by its tetratricopeptide repeats, which is reminiscent of RRNPP family members from the quorum-sensing system. In the absence of the arbitrium peptide, AimR targets the upstream sequence of the aimX gene; its DNA binding activity is prevented following peptide binding. In summary, our findings provide a structural basis for peptide recognition in the phage lysis–lysogeny decision communication system.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Oppenheim, A. B., Kobiler, O., Stavans, J., Court, D. L. & Adhya, S. Switches in bacteriophage λ development. Annu. Rev. Genet. 39, 409–429 (2005).
Golding, I. Single-cell studies of phage λ: hidden treasures under Occam’s rug. Annu. Rev. Virol. 3, 453–472 (2016).
Zeng, L. et al. Decision making at a subcellular level determines the outcome of bacteriophage infection. Cell 141, 682–691 (2010).
Golding, I. Decision making in living cells: lessons from a simple system. Annu. Rev. Biophys. 40, 63–80 (2011).
Echols, H. & Green, L. Establishment and maintenance of repression by bacteriophage λ: the role of the cI, cII, and c3 proteins. Proc. Natl Acad. Sci. USA 68, 2190–2194 (1971).
Kobiler, O. et al. Quantitative kinetic analysis of the bacteriophage λ genetic network. Proc. Natl Acad. Sci. USA 102, 4470–4475 (2005).
Hoyt, M. A., Knight, D. M., Das, A., Miller, H. I. & Echols, H. Control of phage lambda development by stability and synthesis of cII protein: role of the viral cIII and host hflA, himA and himD genes. Cell 31, 565–573 (1982).
Erez, Z. et al. Communication between viruses guides lysis–lysogeny decisions. Nature 541, 488–493 (2017).
Davidson, A. R. Virology: phages make a group decision. Nature 541, 466–467 (2017).
Hynes, A. P. & Moineau, S. Phagebook: the social network. Mol. Cell 65, 963–964 (2017).
Perez-Pascual, D., Monnet, V. & Gardan, R. Bacterial cell-cell communication in the host via RRNPP peptide-binding regulators. Front. Microbiol. 7, 706 (2016).
Do, H. & Kumaraswami, M. Structural mechanisms of peptide recognition and allosteric modulation of gene regulation by the RRNPP family of quorum-sensing regulators. J. Mol. Biol. 428, 2793–2804 (2016).
Rocha-Estrada, J., Aceves-Diez, A. E., Guarneros, G. & de la Torre, M. The RNPP family of quorum-sensing proteins in Gram-positive bacteria. Appl. Microbiol. Biotechnol. 87, 913–923 (2010).
Neiditch, M. B., Capodagli, G. C., Prehna, G. & Federle, M. J. Genetic and structural analyses of RRNPP intercellular peptide signaling of Gram-positive bacteria. Annu. Rev. Genet. 51, 311–333 (2017).
Gallego del Sol, F. & Marina, A. Structural basis of Rap phosphatase inhibition by Phr peptides. PLoS Biol. 11, e1001511 (2013).
Parashar, V., Jeffrey, P. D. & Neiditch, M. B. Conformational change-induced repeat domain expansion regulates Rap phosphatase quorum-sensing signal receptors. PLoS Biol. 11, e1001512 (2013).
Zouhir, S. et al. Peptide-binding dependent conformational changes regulate the transcriptional activity of the quorum-sensor NprR. Nucleic Acids Res. 41, 7920–7933 (2013).
Grenha, R. et al. Structural basis for the activation mechanism of the PlcR virulence regulator by the quorum-sensing signal peptide PapR. Proc. Natl Acad. Sci. USA 110, 1047–1052 (2013).
Parashar, V., Mirouze, N., Dubnau, D. A. & Neiditch, M. B. Structural basis of response regulator dephosphorylation by Rap phosphatases. PLoS Biol. 9, e1000589 (2011).
Holm, L., Kaariainen, S., Rosenstrom, P. & Schenkel, A. Searching protein structure databases with DaliLite v.3. Bioinformatics 24, 2780–2781 (2008).
Krissinel, E. & Henrick, K. Inference of macromolecular assemblies from crystalline state. J. Mol. Biol. 372, 774–797 (2007).
Waters, C. M. & Bassler, B. L. Quorum sensing: cell-to-cell communication in bacteria. Annu. Rev. Cell Dev. Biol. 21, 319–346 (2005).
Bassler, B. L. & Losick, R. Bacterially speaking. Cell 125, 237–246 (2006).
Perego, M. Forty years in the making: understanding the molecular mechanism of peptide regulation in bacterial development. PLoS Biol. 11, e1001516 (2013).
Monnet, V. & Gardan, R. Quorum-sensing regulators in Gram-positive bacteria: ‘cherchez le peptide’. Mol. Microbiol. 97, 181–184 (2015).
Monnet, V., Juillard, V. & Gardan, R. Peptide conversations in Gram-positive bacteria. Crit. Rev. Microbiol. 42, 339–351 (2016).
Core, L. & Perego, M. TPR-mediated interaction of RapC with ComA inhibits response regulator-DNA binding for competence development in Bacillus subtilis. Mol. Microbiol. 49, 1509–1522 (2003).
Shi, K. et al. Structure of peptide sex pheromone receptor PrgX and PrgX/pheromone complexes and regulation of conjugation in Enterococcus faecalis. Proc. Natl Acad. Sci. USA 102, 18596–18601 (2005).
Kozlowicz, B. K. et al. Molecular basis for control of conjugation by bacterial pheromone and inhibitor peptides. Mol. Microbiol. 62, 958–969 (2006).
Declerck, N. et al. Structure of PlcR: Insights into virulence regulation and evolution of quorum sensing in Gram-positive bacteria. Proc. Natl Acad. Sci. USA 104, 18490–18495 (2007).
Bouillaut, L. et al. Molecular basis for group-specific activation of the virulence regulator PlcR by PapR heptapeptides. Nucleic Acids Res. 36, 3791–3801 (2008).
Baker, M. D. & Neiditch, M. B. Structural basis of response regulator inhibition by a bacterial anti-activator protein. PLoS Biol. 9, e1001226 (2011).
Parashar, V., Aggarwal, C., Federle, M. J. & Neiditch, M. B. Rgg protein structure-function and inhibition by cyclic peptide compounds. Proc. Natl Acad. Sci. USA 112, 5177–5182 (2015).
Makthal, N. et al. Structural and functional analysis of RopB: a major virulence regulator in Streptococcus pyogenes. Mol. Microbiol. 99, 1119–1133 (2016).
Talagas, A. et al. Structural insights into streptococcal competence regulation by the cell-to-cell communication system ComRS. PLoS Pathog. 12, e1005980 (2016).
Deng, Y. et al. ApnI, a transmembrane protein responsible for subtilomycin immunity, unveils a novel model for lantibiotic immunity. Appl. Environ. Microbiol. 80, 6303–6315 (2014).
Zahler, S. A., Korman, R. Z., Rosenthal, R. & Hemphill, H. E. Bacillus subtilis bacteriophage SPbeta: localization of the prophage attachment site, and specialized transduction. J. Bacteriol. 129, 556–558 (1977).
Wang, Q. S. et al. The macromolecular crystallography beamline of SSRF. Nucl. Sci. Tech. 26, 12–17 (2015).
Wang, Q.-S. et al. Upgrade of macromolecular crystallography beamline BL17U1 at SSRF. Nucl. Sci. Tech. 29, 68 (2018).
Otwinowski, Z. & Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–326 (1997).
Winn, M. D. et al. Overview of the CCP4 suite and current developments. Acta Crystallogr. D 67, 235–242 (2011).
McCoy, A. J. et al. Phaser crystallographic software. J. Appl. Crystallogr. 40, 658–674 (2007).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D 66, 213–221 (2010).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D 60, 2126–2132 (2004).
Wach, A. PCR-synthesis of marker cassettes with long flanking homology regions for gene disruptions in S. cerevisiae. Yeast 12, 259–265 (1996).
Acknowledgements
We thank the staff of the BL17U1/BL19U1/BL19U2 beamline of the NCPSS at the Shanghai Synchrotron Radiation Facility for assistance during data collection and research associates at the Center for Protein Research, Huazhong Agricultural University, for technical support. We thank Z. Dong and Y. Deng for helpful discussions and gene disruption in prophages. We also thank Q. Tang for help with the DNA I footprinting assay. This work was supported by funds from the National Key R&D Program of China (2018YFA0507700), the National Natural Science Foundation of China (grant no. 81601742 for T.Z.) and the Fundamental Research Funds for the Central Universities (Program No. 2662014BQ028 for T.Z. and 2662017PY031 for P.Y.).
Author information
Authors and Affiliations
Contributions
Q.W., Z.G., P.Y., Z.L., D.P. and T.Z. designed the project and analysed data. Q.W. and Z.G. contributed to protein production and crystallization. Q.W. and K.P. performed EMSA, AUC, SLS, ITC, size-exclusion chromatography, and crosslinking and DNase I footprinting assays. Q.W., J.W. and T.Z performed gene disruption and in vivo experiments. Z.L. performed SAXS measurements. Q.W., Z.G. and T.Z. wrote the paper. All authors contributed to editing of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figures 1–15, Supplementary Tables 1–4.
Rights and permissions
About this article
Cite this article
Wang, Q., Guan, Z., Pei, K. et al. Structural basis of the arbitrium peptide–AimR communication system in the phage lysis–lysogeny decision. Nat Microbiol 3, 1266–1273 (2018). https://doi.org/10.1038/s41564-018-0239-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41564-018-0239-y
This article is cited by
-
Arbitrium communication controls phage lysogeny through non-lethal modulation of a host toxin–antitoxin defence system
Nature Microbiology (2024)
-
Insights into the mechanism of action of the arbitrium communication system in SPbeta phages
Nature Communications (2022)
-
Dormant phages communicate via arbitrium to control exit from lysogeny
Nature Microbiology (2021)
-
Structural insights into DNA recognition by AimR of the arbitrium communication system in the SPbeta phage
Cell Discovery (2019)
-
The secret social lives of viruses
Nature (2019)