Abstract
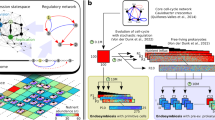
Endosymbiosis plays an important role in ecology and evolution, but fundamental aspects of the origin of intracellular symbionts remain unclear. The extreme age of many symbiotic relationships, lack of data on free-living ancestors and uniqueness of each event hinder investigations. Here, we describe multiple strains of the bacterium Polynucleobacter that evolved independently and under similar conditions from closely related, free-living ancestors to become obligate endosymbionts of closely related ciliate hosts. As these genomes reduced in parallel from similar starting states, they provide unique glimpses into the mechanisms underlying genome reduction in symbionts. We found that gene loss is contingently lineage-specific, with no evidence for ordered streamlining. However, some genes in otherwise disrupted pathways are retained, possibly reflecting cryptic genetic network complexity. We also measured substitution rates between many endosymbiotic and free-living pairs for hundreds of genes, which showed that genetic drift, and not mutation pressure, is the main non-selective factor driving molecular evolution in endosymbionts.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Change history
27 February 2018
The Supplementary Information file originally published with this Article was missing Supplementary Figs 1–7. This has now been corrected.
References
Jezberová, J. et al. Ubiquity of Polynucleobacter necessarius ssp. asymbioticus in lentic freshwater habitats of a heterogeneous 2000 km2 area. Environ. Microbiol. 12, 658–669 (2010).
Hahn, M. W. et al. The passive yet successful way of planktonic life: genomic and experimental analysis of the ecology of a free-living Polynucleobacter population. PLoS ONE 7, e32772 (2012).
Hahn, M. W., Jezberová, J., Koll, U., Saueressig-Beck, T. & Schmidt, J. Complete ecological isolation and cryptic diversity in Polynucleobacter bacteria not resolved by 16S rRNA gene sequences. ISME J. 10, 1642–1655 (2016).
Heckmann, K. & Schmidt, H. J. Polynucleobacter necessarius gen. nov., sp. nov., an obligately endosymbiotic bacterium living in the cytoplasm of Euplotes aediculatus. Int. J. Syst. Bacteriol. 37, 456–457 (1987).
Heckmann, K., Ten Hagen, R. & Görtz, H.-D. Freshwater Euplotes species with a 9 type 1 cirrus pattern depend upon endosymbionts. J. Protozool. 30, 284–289 (1983).
Vannini, C., Petroni, G., Verni, F. & Rosati, G. Polynucleobacter bacteria in the brackish-water species Euplotes harpa (Ciliata Hypotrichia). J. Eukaryot. Microbiol. 52, 116–122 (2005).
Vannini, C. et al. Endosymbiosis in statu nascendi: close phylogenetic relationship between obligately endosymbiotic and obligately free-living Polynucleobacter strains (Betaproteobacteria). Environ. Microbiol. 9, 347–359 (2007).
Hahn, M. W., Schmidt, J., Pitt, A., Taipale, S. J. & Lang, E. Reclassification of four Polynucleobacter necessarius strains as Polynucleobacter asymbioticus comb. nov., Polynucleobacter duraquae sp. nov., Polynucleobacter yangtzensis sp. nov., and Polynucleobacter sinensis sp. nov., and emended description of the species Polynucleobacter necessarius. Int. J. Syst. Evol. Microbiol. 66, 2883–2892 (2016).
Vannini, C., Ferrantini, F., Ristori, A., Verni, F. & Petroni, G. Betaproteobacterial symbionts of the ciliate Euplotes: origin and tangled evolutionary path of an obligate microbial association. Environ. Microbiol. 14, 2553–2563 (2012).
Gil, R. et al. The genome sequence of Blochmannia floridanus: comparative analysis of reduced genomes. Proc. Natl Acad. Sci. USA 100, 9388–9393 (2003).
Rio, R. V. M., Lefevre, C., Heddi, A. & Aksoy, S. Comparative genomics of insect-symbiotic bacteria: influence of host environment on microbial genome composition. Appl. Environ. Microbiol. 69, 6825–6832 (2003).
Moran, N. A. Accelerated evolution and Muller’s rachet in endosymbiotic bacteria. Proc. Natl Acad. Sci. USA 93, 2873–2878 (1996).
Lamelas, A. et al. Serratia symbiotica from the aphid Cinara cedri: a missing link from facultative to obligate insect endosymbiont. PLoS Genet. 7, e1002357 (2011).
Oakeson, K. F. et al. Genome degeneration and adaptation in a nascent stage of symbiosis. Genome Biol. Evol. 6, 76–93 (2014).
Gould, S. J. Wonderful Life (W. W. Norton & Co., New York, 1989).
Wernegreen, J. J., Richardson, A. O. & Moran, N. A. Parallel acceleration of evolutionary rates in symbiont genes underlying host nutrition. Mol. Phylogenet. Evol. 19, 479–485 (2001).
O’Fallon, B. Population structure, levels of selection, and the evolution of intracellular symbionts. Evolution 62, 361–373 (2008).
Pettersson, M. E. & Berg, O. G. Muller’s ratchet in symbiont populations. Genetica 130, 199–211 (2007).
Moran, N. A., McLaughlin, H. J. & Sorek, R. The dynamics and time scale of ongoing genomic erosion in symbiotic bacteria. Science 323, 379–382 (2009).
McCutcheon, J. P. & Moran, N. A. Extreme genome reduction in symbiotic bacteria. Nat. Rev. Microbiol. 10, 13–26 (2012).
Burke, G. R. & Moran, N. A. Massive genomic decay in Serratia symbiotica, a recently evolved symbiont of aphids. Genome Biol. Evol. 3, 195–208 (2011).
Clayton, A. L., Jackson, D. G., Weiss, R. B. & Dale, C. Adaptation by deletogenic replication slippage in a nascent symbiont. Mol. Biol. Evol. 33, 1957–1966 (2016).
Husnik, F. & McCutcheon, J. P. Repeated replacement of an intrabacterial symbiont in the tripartite nested mealybug symbiosis. Proc. Natl Acad. Sci. USA 113, E5416–E5424 (2016).
Wernegreen, J. J. & Moran, N. A. Evidence for genetic drift in endosymbionts (Buchnera): analyses of protein-coding genes. Mol. Biol. Evol. 16, 83–87 (1999).
Itoh, T., Martin, W. & Nei, M. Acceleration of genomic evolution caused by enhanced mutation rate in endocellular symbionts. Proc. Natl Acad. Sci. USA 99, 12944–12948 (2002).
Nei, M. Selectionism and neutralism in molecular evolution. Mol. Biol. Evol. 22, 2318–2342 (2005).
Boscaro, V. et al. Polynucleobacter necessarius, a model for genome reduction in both free-living and symbiotic bacteria. Proc. Natl Acad. Sci. USA 110, 18590–18595 (2013).
Hao, Z. et al. Genome sequence of a freshwater low-nucleic-acid-content bacterium, betaproteobacterium strain CB. Genome Announc. 1, e0013513 (2013).
Syberg-Olsen, M. J. et al. Biogeography and character evolution of the ciliate genus Euplotes (Spirotrichea, Euplotia), with description of Euplotes curdsi sp. nov. PLoS ONE 11, e0165442 (2016).
Manzano-Marín, A. & Latorre, A. Settling down: the genome of Serratia symbiotica from the aphid Cinara tujafina zooms in on the process of accommodation to a cooperative intracellular life. Genome Biol. Evol. 6, 1683–1698 (2014).
Hoetzinger, M., Schmidt, J., Jezberová, J., Koll, U. & Hahn, M. W. Microdiversification of a pelagic Polynucleobacter species is mainly driven by acquisition of genomic islands from a partially interspecific gene pool. Appl. Environ. Microbiol. 83, e02266-16 (2017).
Moran, N. A., McCutcheon, J. P. & Nakabachi, A. Genomics and evolution of heritable bacterial symbionts. Annu. Rev. Genet. 42, 165–190 (2008).
Clayton, A. L. et al. A novel human-infection-derived bacterium provides insights into the evolutionary origins of mutualistic insect-bacterial symbioses. PLoS Genet. 8, e1002990 (2012).
Bennett, G. M., McCutcheon, J. P., McDonald, B. R. & Moran, N. A. Lineage-specific patterns of genome deterioration in obligate symbionts of sharpshooter leafhoppers. Genome Biol. Evol. 8, 296–301 (2016).
Andersson, J. O. & Andersson, S. G. E. Insights into the evolutionary process of genome degradation. Curr. Opin. Genet. Dev. 9, 664–671 (1999).
Nilsson, A. I. et al. Bacterial genome size reduction by experimental evolution. Proc. Natl Acad. Sci. USA 102, 12112–12116 (2005).
Hershberg, R., Tang, H. & Petrov, D. A. Reduced selection leads to accelerated gene loss in Shigella. Genome Biol. 8, R164 (2007).
McCutcheon, J. P. & von Dohlen, C. D. An interdependent metabolic patchwork in the nested symbiosis of mealybugs. Curr. Biol. 21, 1366–1372 (2011).
Ghignone, S. et al. The genome of the obligate endobacterium of the AM fungus reveals an interphylum network of nutritional interactions. ISME J. 6, 136–145 (2012).
Nakabachi, A. et al. The 160-kilobase genome of the bacterial endosymbiont Carsonella. Science 314, 267 (2006).
Smith, W. A. et al. Phylogenetic analysis of symbionts in feather-feeding lice of the genus Columbicola: evidence for repeated symbiont replacements. BMC Evol. Biol. 13, 109 (2013).
Rosati, G., Modeo, L., Melai, M., Petroni, G. & Verni, F. A multidisciplinary approach to describe protists: a morphological, ultrastructural, and molecular study on Peritromus kahli Villeneuve-Brachon, 1940 (Ciliophora, Heterotrichea). J. Eukaryot. Microbiol. 51, 49–59 (2004).
Katoh, K. & Standley, D. M. MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol. Biol. Evol. 30, 772–780 (2013).
Capella-Gutiérrez, S., Silla-Martínez, J. M. & Gabaldón, T. trimAl: a tool for automated alignment trimming in large-scale phylogenetic analyses. Bioinformatics 25, 1972–1973 (2009).
Nguyen, L.-T., Schmidt, H. A., von Haeseler, A. & Minh, B. Q. IQ-TREE: a fast and effective stochastic algorithm for estimating maximum-likelihood phylogenies. Mol. Biol. Evol. 32, 268–274 (2015).
Ronquist, F. et al. MrBayes 3.2: efficient Bayesian phylogenetic inference and model choice across a large model space. Syst. Biol. 61, 539–542 (2012).
Prescott, D. M. Evolution of DNA organization in hypotrichous ciliates. Ann. NY Acad. Sci. 870, 301–313 (1999).
Tritt, A., Eisen, J. A., Facciotti, M. T. & Darling, A. E. An integrated pipeline for de novo assembly of microbial genomes. PLoS ONE 7, e42304 (2012).
Aziz, R. K. et al. The RAST server: rapid annotations using subsystems technology. BMC Genomics 9, 75 (2008).
Emms, D. M. & Kelly, S. OrthoFinder: solving fundamental biases in whole genome comparisons dramatically improves orthogroup inference accuracy. Genome Biol. 16, 157 (2015).
Garcia, S. L. et al. Metabolic potential of a single cell belonging to one of the most abundant lineages in freshwater bacterioplankton. ISME J. 7, 137–147 (2013).
Tatusova, T., Ciufo, S., Fedorov, B., O’Neill, K. & Tolstoy, I. RefSeq microbial genomes database: new representation and annotation strategy. Nucleic Acids Res. 42, D553–D559 (2014).
Criscuolo, A. & Gribaldo, S. BMGE (block mapping and gathering with entropy): a new software for selection of phylogenetic informative regions from multiple sequence alignments. BMC Evol. Biol. 10, 210 (2010).
Stamatakis, A. RAxML version 8: a tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 30, 1312–1313 (2014).
Le, S. Q., Dang, C. C. & Gascuel, O. Modeling protein evolution with several amino acid replacement matrices depending on site rates. Mol. Biol. Evol. 29, 2921–2936 (2012).
Le, S. Q. & Gascuel, O. An improved general amino acid replacement matrix. Mol. Biol. Evol. 25, 1307–1320 (2008).
Le, S. Q., Gascuel, O. & Lartillot, N. Empirical profile mixture models for phylogenetic reconstruction. Bioinformatics 24, 2317–2323 (2008).
Lartillot, N. & Philippe, H. A Bayesian mixture model for across-site heterogeneities in the amino-acid replacement process. Mol. Biol. Evol. 21, 1095–1109 (2004).
Yang, Z. PAML4: phylogenetic analysis by maximum likelihood. Mol. Biol. Evol. 24, 1586–1591 (2007).
R Core Team R: A Language and Environment for Statistical Computing. (R Foundation for Statistical Computing, Vienna, 2013).
Kanehisa, M., Furumichi, M., Tanabe, M., Sato, Y. & Morishima, K. KEGG: new perspectives on genomes, pathways, diseases and drugs. Nucleic Acids Res. 45, D353–D361 (2017).
Caspi, R. et al. The MetaCyc database of metabolic pathways and enzymes and the BioCyc collection of Pathway/Genome databases. Nucleic Acids Res. 42, D459–D471 (2014).
Finn, R. D. et al. InterPro in 2017—beyond protein family and domain annotations. Nucleic Acids Res. 45, D190–D199 (2017).
Placzek, S. et al. BRENDA in 2017: new perspectives and new tools in BRENDA. Nucleic Acids Res. 45, D380–D388 (2017).
Hahn, M. W., Stadler, P., Wu, Q. L. & Pöckl, M. The filtration-acclimatization method for isolation of an important fraction of the not readily cultivable bacteria. J. Microbiol. Methods 57, 379–390 (2004).
Acknowledgements
We thank S. Gabrielli for helping with the artwork. This work was supported by grants from the Natural Sciences and Engineering Research Council of Canada (227301 and 6544-2013 awarded to P.J.K. and D.H.L, respectively). V.B. and M.K. were supported by fellowships from the Tula Foundation to the Centre for Microbial Diversity and Evolution.
Author information
Authors and Affiliations
Contributions
V.B., D.H.L. and P.J.K. designed the study. V.B. sampled and isolated the ciliates. V.B. and C.V. cultured, screened and identified the Euplotes strains and Polynucleobacter symbionts. C.V. performed the isolation experiments on the symbionts. V.B. and C.V. optimized and performed the genomic DNA extractions. D.H.L. prepared the libraries. V.B. assembled and annotated the genomes. V.B. and M.F. conducted the functional analysis. M.K. performed the phylogenomic inference, clustering analysis and dN/dS calculations. V.B., M.K. and P.J.K. wrote the paper. All authors participated in the drafting process.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
A correction to this article is available online at https://doi.org/10.1038/s41559-018-0484-8.
Electronic supplementary material
Supplementary Information
Supplementary Figures 1–7 and Supplementary Discussion
Supplementary Data 1
Functional modules
Supplementary Data 2
dS and dN/dS comparisons
Rights and permissions
About this article
Cite this article
Boscaro, V., Kolisko, M., Felletti, M. et al. Parallel genome reduction in symbionts descended from closely related free-living bacteria. Nat Ecol Evol 1, 1160–1167 (2017). https://doi.org/10.1038/s41559-017-0237-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41559-017-0237-0
This article is cited by
-
Symbionts of Ciliates and Ciliates as Symbionts
Indian Journal of Microbiology (2024)
-
Contrasting outcomes of genome reduction in mikrocytids and microsporidians
BMC Biology (2023)
-
Single-cell Microbiomics Unveils Distribution and Patterns of Microbial Symbioses in the Natural Environment
Microbial Ecology (2023)
-
Insights into ‘Symbiodiniaceae phycosphere’ in a coral holobiont
Symbiosis (2021)
-
Serial horizontal transfer of vitamin-biosynthetic genes enables the establishment of new nutritional symbionts in aphids’ di-symbiotic systems
The ISME Journal (2020)