Abstract
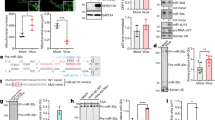
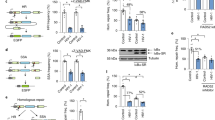
The human immunodeficiency virus type 1 (HIV-1) causes persistent infection in human and induces miR-146a expression in infected cells. miR-146a represses the innate immune response by inhibiting the expression of TRAF6 and IRAK1 genes, thus negatively controls the NF-κB-related cytokines and interferon stimulated genes. Here we reported that lentiviral CRISPR/Cas9 system was highly efficient in introducing mutations in the precursor miR-146a genomic sequences, resulting in a loss of miR-146a expression and function. miR-146a ablation led to increasing cytokines production in LPS-stimulated A549 cells. Moreover, miR-146a knockout in HIV-1 infected MT2 cells markedly increased the expression of cytokines and HIV-1 restriction factors and reversed T cell exhaustion markers expression, thus influencing HIV-1 replication. Our study indicates that lentiviral CRISPR/Cas9-mediated gene editing is an effective approach to abrogate miR-146a expression, which consequently inhibits HIV-1 replication as well as proviral reactivation by enhancing the expression of cytokines and HIV-1 restriction factors.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 6 digital issues and online access to articles
$119.00 per year
only $19.83 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Dong CS, Qu L, Wang HY, Wei L, Dong YS, Xiong SD. Targeting hepatitis B virus cccDNA by CRISPR/Cas9 nuclease efficiently inhibits viral replication. Antivir Res. 2015;118:110–17.
van Diemen FR, Kruse EM, Hooykaas MJG, Bruggeling CE, Schurch AC, van Ham PM, et al. CRISPR/Cas9-mediated genome editing of herpesviruses limits productive and latent infections. Plos Pathogens. 2016;12:e1005701.
Kaminski R, Chen Y, Fischer T, Tedaldi E, Napoli A, Zhang Y, et al. Elimination of HIV-1 genomes from human T-lymphoid cells by CRISPR/Cas9 gene editing. Sci Rep. 2016;6:22555.
Wang Q, Chen S, Xiao Q, Liu Z, Liu S, Hou P, et al. Genome modification of CXCR4 by Staphylococcus aureus Cas9 renders cells resistance to HIV-1 infection. Retrovirology. 2017;14:51.
Li C, Guan XM, Du T, Jin W, Wu B, Liu YL, et al. Inhibition of HIV-1 infection of primary CD4(+) T-cells by gene editing of CCR5 using adenovirus-delivered CRISPR/Cas9. J General Virol. 2015;96:2381–93.
Nguyen EV, Gharib SA, Crothers K, Chow YH, Park DR, Goodlett DR, et al. Proteomic landscape of bronchoalveolar lavage fluid in human immunodeficiency virus infection. Am J Physiol-Lung Cell Mol Physiol. 2014;306:L35–L42.
Foldi J, Kozhaya L, McCarty B, Mwamzuka M, Marshed F, Ilmet T, et al. HIV-infected children have elevated levels of PD-1(+) memory CD4 T cells with low proliferative capacity and high inflammatory cytokine effector functions. J Infect Dis. 2017;216:641–50.
Trautmann L, Janbazian L, Chomont N, Said EA, Gimmig S, Bessette B, et al. Upregulation of PD-1 expression on HIV-specific CD8(+) T cells leads to reversible immune dysfunction. Nat Med. 2006;12:1198–202.
Velu V, Shetty RD, Larsson M, Shankar EM. Role of PD-1 co-inhibitory pathway in HIV infection and potential therapeutic options. Retrovirology. 2015;12:14.
Hunt PW. HIV and inflammation: mechanisms and consequences. Curr HIV/AIDS Rep. 2012;9:139–47.
Stacey AR, Norris PJ, Qin L, Haygreen EA, Taylor E, Heitman J, et al. Induction of a striking systemic cytokine cascade prior to peak viremia in acute human immunodeficiency virus type 1 infection, in contrast to more modest and delayed responses in acute hepatitis B and C virus infections. J Virol. 2009;83:3719–33.
Zhao GX, Liu LF, Su B, Zhang T, Chen P, Li W, et al. The dynamic changes of interferon lambdas relatedgenes and proteins in JAK/STAT pathway in both acute and chronic HIV-1 infected patients. AIDS Res Therapy. 2017;14:31.
Honda K, Takaoka A, Taniguchi T. Type I inteferon gene induction by the interferon regulatory factor family of transcription factors. Immunity. 2006;25:349–60.
Anderson P. Post-transcriptional control of cytokine production. Nat Immunol. 2008;9:353–9.
Savan R. Post-transcriptional regulation of interferons and their signaling pathways. J Interferon Cytokine Res. 2014;34:318–29.
Taganov KD, Boldin MP, Chang KJ, Baltimore D. NF-kappaB-dependent induction of microRNA miR-146, an inhibitor targeted to signaling proteins of innate immune responses. Proc Natl Acad Sci USA. 2006;103:12481–6.
Perry MM, Moschos SA, Williams AE, Shepherd NJ, Larner-Svensson HM, Lindsay MA. Rapid changes in microRNA-146a expression negatively regulate the IL-1beta-induced inflammatory response in human lung alveolar epithelial cells. J Immunol. 2008;180:5689–98.
Wang S, Zhang X, Ju Y, Zhao B, Yan X, Hu J, et al. MicroRNA-146a feedback suppresses T cell immune function by targeting Stat1 in patients with chronic hepatitis B. J Immunol. 2013;191:293–301.
Hou Z, Zhang J, Han Q, Su C, Qu J, Xu D, et al. Hepatitis B virus inhibits intrinsic RIG-I and RIG-G immune signaling via inducing miR146a. Sci Rep. 2016;6:26150.
Duskova K, Nagilla P, Le HS, Iyer P, Thalamuthu A, Martinson J, et al. MicroRNA regulation and its effects on cellular transcriptome in Human Immunodeficiency Virus-1 (HIV-1) infected individuals with distinct viral loadand CD4 cell counts. BMC Infectious Dis. 2013;13:250.
Rom S, Rom I, Passiatore G, Pacifici M, Radhakrishnan S, Del Valle L, et al. CCL8/MCP-2 is a target for mir-146a in HIV-1-infected human microglial cells. FASEB J.2010;24:2292–300.
Egana-Gorrono L, Escriba T, Boulanger N, Guardo AC, Leon A, Bargallo ME, et al. Differential MicroRNA expression profile between stimulated PBMCs from HIV-1 infected elite controllers and viremic progressors. PLoS ONE. 2014;9:e106360.
Feng Y, Zhang XX, Song QF, Li TB, Zeng Y. Drosha processing controls the specificity and efficiency of global microRNA expression. Biochim Et Biophys Acta-Gene Regul Mech. 2011;1809:700–7.
Feng Y, Zhang XX, Graves P, Zeng Y. A comprehensive analysis of precursor microRNA cleavage by human Dicer. Rna-a Publ Rna Soc. 2012;18:2083–92.
Qu XY, Wang PF, Ding DL, Li L, Wang HB, Ma L, et al. Zinc-finger-nucleases mediate specific and efficient excision of HIV-1 proviral DNA from infected and latently infected human T cells. Nucleic Acids Res. 2013;41:7771–82.
D’Elia RV, Harrison K, Oyston PC, Lukaszewski RA, Clark GC. Targeting the “cytokine storm” for therapeutic benefit. Clin Vaccin Immunol. 2013;20:319–27.
Chousterman BG, Swirski FK, Weber GF. Cytokine storm and sepsis disease pathogenesis. Semin Immunopathol. 2017;39:517–28.
Tisoncik JR, Korth MJ, Simmons CP, Farrar J, Martin TR, Katze MG. Into the eye of the cytokine storm. Microbiol Mol Biol Rev. 2012;76:16–32.
Kassu A, Marcus RA, D’Souza MB, Kelly-McKnight EA, Golden-Mason L, Akkina R, et al. Regulation of virus-specific CD4(+) T cell function by multiple costimulatory receptors during chronic HIV infection. J Immunol. 2010;185:3007–18.
Kaufmann DE, Kavanagh DG, Pereyra F, Zaunders JJ, Mackey EW, Miura T, et al. Upregulation of CTLA-4 by HIV-specific CD4+ T cells correlates with disease progression and defines a reversible immune dysfunction. Nat Immunol. 2007;8:1246–54.
Sun T, Li X, Song H, Gao F, Zhou G, Li X, et al. MiR-146a aggravates LPS-induced inflammatory injury by targeting CXCR4 in the articular chondrocytes. Cell Physiol Biochem. 2017;44:1282–94.
Chen QZ, Luo F, Lu MX, Li N, Teng Y, Huang QL, et al. HTNV-induced upregulation of miR-146a in HUVECs promotes viral infection by modulating pro-inflammatory cytokine release. Biochem Biophys Res Commun. 2017;493:807–13.
Fu Y, Zhang L, Zhang F, Tang T, Zhou Q, Feng C, et al. Exosome-mediated miR-146a transfer suppresses type I interferon response and facilitates EV71 infection. PLoS Pathog. 2017;13:e1006611.
Deng SM, Wang HL, Jia CL, Zhu SK, Chu XM, Ma Q, et al. MicroRNA-146a induces lineage-negative bone marrow cell apoptosis and senescence by targeting polo-like kinase 2 expression. Arterioscler Thromb Vasc Biol. 2017;37:280.
Angelini F, Pagano F, Bordin A, Picchio V, De Falco E, Chimenti I. Getting old through the blood: circulating molecules in aging and senescence of cardiovascular regenerative cells. Front Cardiovasc Med. 2017;4:62.
Olivieri F, Lazzarini R, Recchioni R, Marcheselli F, Rippo MR, Di Nuzzo S, et al. MiR-146a as marker of senescence-associated pro-inflammatory status in cells involved in vascular remodelling. Age. 2013;35:1157–72.
Li D, Duan MY, Feng Y, Geng LL, Li XQ, Zhang WG. MiR-146a modulates macrophage polarization in systemic juvenile idiopathic arthritis by targeting INHBA. Mol Immunol. 2016;77:205–12.
Huang C, Liu XJ, QunZhou, Xie J, Ma TT, Meng XM, et al. MiR-146a modulates macrophage polarization by inhibiting Notch1 pathway in RAW264.7 macrophages. Int Immunopharmacol. 2016;32:46–54.
Fu YF, Sander JD, Reyon D, Cascio VM, Joung JK. Improving CRISPR-Cas nuclease specificity using truncated guide RNAs. Nat Biotechnol. 2014;32:279–84.
Li L, Feng HM, Da Q, Jiang HL, Chen L, Xie LL, et al. Expression of HIV-encoded microRNA-TAR and its inhibitory effect on viral replication in human primary macrophages. Arch Virol. 2016;161:1115–23.
Acknowledgements
This research was supported by research grants from the National Natural Science Foundation of China (Nos. 81271818 and 81471940), project of Hubei Provincial Science & Technology (2015CFB184) to Y.F. And we appreciate Research grants from the National Natural Science Foundation of China (No. 81471941), the Science and Technology Ministry of China as part of a major project of infectious disease control and prevention carried out by W.H. (2014ZX10001003).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Rights and permissions
About this article
Cite this article
Teng, Y., Luo, M., Yu, T. et al. CRISPR/Cas9-mediated deletion of miR-146a enhances antiviral response in HIV-1 infected cells. Genes Immun 20, 327–337 (2019). https://doi.org/10.1038/s41435-018-0036-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41435-018-0036-x
This article is cited by
-
MicroRNAs and long non-coding RNAs during transcriptional regulation and latency of HIV and HTLV
Retrovirology (2024)
-
Novel interferon-sensitive genes unveiled by correlation-driven gene selection and systems biology
Scientific Reports (2021)
-
Elevated expression of miR-146a correlates with high levels of immune cell exhaustion markers and suppresses cellular immune function in chronic HIV-1-infected patients
Scientific Reports (2019)
-
Message from the new Editors-in-Chief
Genes & Immunity (2019)