Abstract
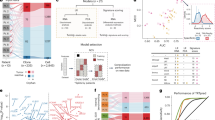
Negative regulation of antitumor T-cell-immune responses facilitates tumor-immune escape. Here, we show that deletion of CD147, a type I transmembrane molecule, in T cells, strongly limits in vivo tumor growth of mouse melanoma and lung cancer in a CD8+ T-cell-dependent manner. In mouse tumor models, CD147 expression was upregulated on CD8+ tumor-infiltrating lymphocytes (TILs), and CD147 was coexpressed with two immune-checkpoint molecules, Tim-3 and PD-1. Mining publicly available gene-profiling data for CD8+ TILs in tumor biopsies from metastatic melanoma patients showed a higher level of CD147 expression in exhausted CD8+ TILs than in other subsets of CD8+ TILs, along with expression of PD-1 and TIM-3. Additionally, CD147 deletion increased the abundance of TILs, cytotoxic effector function of CD8+ T cells, and frequency of PD-1+ CD8+ TILs, and partly reversed the dysfunctional status of PD-1+Tim-3+CD8+ TILs. The cytotoxic transcription factors Runx3 and T-bet mediation enhanced antitumor responses by CD147–/– CD8+ T cells. Moreover, CD147 deletion in T cells increased the frequency of TRM-like cells and the expression of the T-cell chemokines CXCL9 and CXCL10 in the tumor microenvironment. Analysis of tumor tissue samples from patients with non-small-cell lung cancer showed negative correlations between CD147 expression on CD8+ TILs and the abundance of CD8+ TILs, histological grade of the tumor tissue samples, and survival of patients with advanced tumors. Altogether, we found a novel function of CD147 as a negative regulator of antitumor responses mediated by CD8+ TILs and identified CD147 as a potential target for cancer immunotherapy.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
References
Martinez-Lostao, L., Anel, A. & Pardo, J. How do cytotoxic lymphocytes kill cancer cells? Clin. Cancer Res. 21, 5047–5056 (2015).
Thommen, D. S. & Schumacher, T. N. T cell dysfunction in cancer. Cancer Cell 33, 547–562 (2018).
Li, H. et al. Dysfunctional CD8 T cells form a proliferative, dynamically regulated compartment within human melanoma. Cell 176, 775–789 (2019).
Gubin, M. M. et al. High-dimensional analysis delineates myeloid and lymphoid compartment remodeling during successful immune-checkpoint cancer therapy. Cell 175, 1014–1030 (2018).
Chew, V. et al. Delineation of an immunosuppressive gradient in hepatocellular carcinoma using high-dimensional proteomic and transcriptomic analyses. Proc. Natl Acad. Sci. USA 114, E5900–E5909 (2017).
Khan, O. et al. TOX transcriptionally and epigenetically programs CD8(+) T cell exhaustion. Nature 571, 211–218 (2019).
Wang, J. et al. Fibrinogen-like protein 1 is a major immune inhibitory ligand of LAG-3. Cell 176, 334–347 (2019).
Li, J. et al. Co-inhibitory molecule B7 superfamily member 1 expressed by tumor-infiltrating myeloid cells induces dysfunction of anti-tumor CD8(+) T cells. Immunity 48, 773–786 (2018).
Sun, C., Mezzadra, R. & Schumacher, T. N. Regulation and function of the PD-L1 checkpoint. Immunity 48, 434–452 (2018).
Pal, S. K. et al. Clinical cancer advances 2019: annual report on progress against cancer from the American Society of Clinical Oncology. J. Clin. Oncol. 37, 834–849 (2019).
Hashimoto, M. et al. CD8 T cell exhaustion in chronic infection and cancer: opportunities for interventions. Annu. Rev. Med. 69, 301–318 (2018).
Ribas, A. & Wolchok, J. D. Cancer immunotherapy using checkpoint blockade. Science 359, 1350–1355 (2018).
Sharma, P., Hu-Lieskovan, S., Wargo, J. A. & Ribas, A. Primary, adaptive, and acquired resistance to cancer immunotherapy. Cell 168, 707–723 (2017).
Wei, S. C. et al. Distinct cellular mechanisms underlie Anti-CTLA-4 and anti-PD-1 checkpoint blockade. Cell 170, 1120–1133 (2017).
Li, J., Ni, L. & Dong, C. Immune checkpoint receptors in cancer: redundant by design? Curr. Opin. Immunol. 45, 37–42 (2017).
Koch, C. et al. T cell activation-associated epitopes of CD147 in regulation of the T cell response, and their definition by antibody affinity and antigen density. Int. Immunol. 11, 777–786 (1999).
Weidle, U. H., Scheuer, W., Eggle, D., Klostermann, S. & Stockinger, H. Cancer-related issues of CD147. Cancer Genomics Proteom. 7, 157–169 (2010).
Solstad, T. et al. CD147 (Basigin/Emmprin) identifies FoxP3+CD45RO+CTLA4+-activated human regulatory T cells. Blood 118, 5141–5151 (2011).
Landskron, J. & Tasken, K. CD147 in regulatory T cells. Cell. Immunol. 282, 17–20 (2013).
Renno, T. et al. A role for CD147 in thymic development. J. Immunol. 168, 4946–4950 (2002).
Arora, K. et al. Extracellular cyclophilins contribute to the regulation of inflammatory responses. J. Immunol. 175, 517–522 (2005).
Gwinn, W. M. et al. Novel approach to inhibit asthma-mediated lung inflammation using anti-CD147 intervention. J. Immunol. 177, 4870–4879 (2006).
Damsker, J. M. et al. Targeting the chemotactic function of CD147 reduces collagen-induced arthritis. Immunology 126, 55–62 (2009).
Agrawal, S. M., Silva, C., Wang, J., Tong, J. P. & Yong, V. W. A novel anti-EMMPRIN function-blocking antibody reduces T cell proliferation and neurotoxicity: relevance to multiple sclerosis. J. Neuroinflammation. 9, 64 (2012).
Nabeshima, K. et al. Emmprin, a cell surface inducer of matrix metalloproteinases (MMPs), is expressed in T-cell lymphomas. J. Pathol. 202, 341–351 (2004).
Chen, X. et al. Inhibition of CD147 gene expression via RNA interference reduces tumor cell proliferation, activation, adhesion, and migration activity in the human Jurkat T-lymphoma cell line. Cancer Investig. 26, 689–697 (2008).
Agrawal, S. M. et al. Extracellular matrix metalloproteinase inducer shows active perivascular cuffs in multiple sclerosis. Brain 136, 1760–1777 (2013).
Guo, N. et al. CD147 and CD98 complex-mediated homotypic aggregation attenuates the CypA-induced chemotactic effect on Jurkat T cells. Mol. Immunol. 63, 253–263 (2015).
Staffler, G. et al. Selective inhibition of T cell activation via CD147 through novel modulation of lipid rafts. J. Immunol. 171, 1707–1714 (2003).
Hu, J. et al. Involvement of HAb18G/CD147 in T cell activation and immunological synapse formation. J. Cell Mol. Med. 14, 2132–2143 (2010).
Ruiz, S., Castro-Castro, A. & Bustelo, X. R. CD147 inhibits the nuclear factor of activated T-cells by impairing Vav1 and Rac1 downstream signaling. J. Biol. Chem. 283, 5554–5566 (2008).
Supper, V. et al. Association of CD147 and calcium exporter PMCA4 uncouples IL-2 expression from early TCR signaling. J. Immunol. 196, 1387–1399 (2016).
Yao, H. et al. Important functional roles of basigin in thymocyte development and T cell activation. Int. J. Biol. Sci. 10, 43–52 (2013).
Miller, B. C. et al. Subsets of exhausted CD8(+) T cells differentially mediate tumor control and respond to checkpoint blockade. Nat. Immunol. 20, 326–336 (2019).
Sade-Feldman, M. et al. Defining T cell states associated with response to checkpoint immunotherapy in melanoma. Cell 175, 998–1013 (2018).
Thommen, D. S. et al. A transcriptionally and functionally distinct PD-1(+) CD8(+) T cell pool with predictive potential in non-small-cell lung cancer treated with PD-1 blockade. Nat. Med. 24, 994–1004 (2018).
Singer, M. et al. A distinct gene module for dysfunction uncoupled from activation in tumor-infiltrating T cells. Cell 166, 1500–1511 (2016).
Cruz-Guilloty, F. et al. Runx3 and T-box proteins cooperate to establish the transcriptional program of effector CTLs. J. Exp. Med. 206, 51–59 (2009).
Milner, J. J. et al. Runx3 programs CD8(+) T cell residency in non-lymphoid tissues and tumours. Nature 552, 253–257 (2017).
Hahn, J. N., Kaushik, D. K. & Yong, V. W. The role of EMMPRIN in T cell biology and immunological diseases. J. Leukoc. Biol. 98, 33–48 (2015).
Zhu, X., Song, Z., Zhang, S., Nanda, A. & Li, G. CD147: a novel modulator of inflammatory and immune disorders. Curr. Med. Chem. 21, 2138–2145 (2014).
Guo, N. et al. A critical epitope in CD147 facilitates memory CD4(+) T-cell hyper-activation in rheumatoid arthritis. Cell. Mol. Immunol. 16, 568–579 (2019).
Geng, J. J. et al. Targeting CD147 for T to NK lineage reprogramming and tumor therapy. EBioMedicine 20, 98–108 (2017).
Chen, R. et al. CD147 deficiency in T cells prevents thymic involution by inhibiting the EMT process in TECs in the presence of TGFbeta. Cell. Mol. Immunol. https://doi.org/10.1038/s41423-019-0353-7 (2020).
Carow, B. et al. lck-driven Cre expression alters T cell development in the thymus and the frequencies and functions of peripheral T cell subsets. J. Immunol. 197, 2261–2268 (2017).
Shan, Q. et al. The transcription factor Runx3 guards cytotoxic CD8(+) effector T cells against deviation towards follicular helper T cell lineage. Nat. Immunol. 18, 931–939 (2017).
Ciucci, T., Vacchio, M. S. & Bosselut, R. A STAT3-dependent transcriptional circuitry inhibits cytotoxic gene expression in T cells. Proc. Natl Acad. Sci. USA 114, 13236–13241 (2017).
Ganesan, A. P. et al. Tissue-resident memory features are linked to the magnitude of cytotoxic T cell responses in human lung cancer. Nat. Immunol. 18, 940–950 (2017).
Nagarsheth, N., Wicha, M. S. & Zou, W. Chemokines in the cancer microenvironment and their relevance in cancer immunotherapy. Nat. Rev. Immunol. 17, 559–572 (2017).
Sharpe, A. H. & Pauken, K. E. The diverse functions of the PD1 inhibitory pathway. Nat. Rev. Immunol. 18, 153–167 (2018).
Chow, M. T. et al. Intratumoral activity of the CXCR3 chemokine system is required for the efficacy of anti-PD-1 therapy. Immunity 50, 1498–1512 (2019).
Schalper, K. A. et al. Objective measurement and clinical significance of TILs in non-small cell lung cancer. J. Natl. Cancer Inst. 107, dju435 (2015).
Kurachi, M. et al. Optimized retroviral transduction of mouse T cells for in vivo assessment of gene function. Nat. Protoc. 12, 1980–1998 (2017).
Kluger, H. M. et al. PD-L1 studies across tumor types, its differential expression and predictive value in patients treated with immune checkpoint inhibitors. Clin. Cancer Res. 23, 4270–4279 (2017).
Acknowledgements
The authors thank Xiwen Dong, Lijuan Wang, Qian He, and Xiaomin Li for their technical assistance and Jingmin Yu for assisting with mouse genotyping. This work was supported by grants from the National Natural Science Foundation of China (81572802), the National Basic Research Program of China (2015CB553700), and the Fourth Military Medical University Foundation for Development of Science and Technology (2019XB005).
Author information
Authors and Affiliations
Contributions
Y.C., J.X., X.W., H.Y., Z.Y., W.W., P.W., T.G., Y.L., X.Y., and H.L. conducted the experiments and data analysis; J.X. and Z.-N.C. conceived the study and designed the experiments; Y.C., J.X., H.B., and Z.-N.C. discussed and interpreted the data; Y.C. and J.X. wrote the paper. All authors read and approved the final paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Rights and permissions
About this article
Cite this article
Chen, Y., Xu, J., Wu, X. et al. CD147 regulates antitumor CD8+ T-cell responses to facilitate tumor-immune escape. Cell Mol Immunol 18, 1995–2009 (2021). https://doi.org/10.1038/s41423-020-00570-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41423-020-00570-y
Keywords
This article is cited by
-
CD8+ T cell-based cancer immunotherapy
Journal of Translational Medicine (2024)
-
124I-labeled anti-CD147 antibody for noninvasive detection of CD147-positive pan-cancers: construction and preclinical studies
Acta Pharmacologica Sinica (2024)
-
Intervention with extracellular matrix metalloproteinase inducer in osteoclasts attenuates periodontitis-induced bone resorption
Odontology (2024)
-
Pharmacokinetic Parameters of Recombinant Human Cyclophilin A in Mice
European Journal of Drug Metabolism and Pharmacokinetics (2024)
-
Impact of BSG/CD147 gene expression on diagnostic, prognostic and therapeutic strategies towards malignant cancers and possible susceptibility to SARS-CoV-2
Molecular Biology Reports (2023)