Abstract
Glioblastoma multiforme (GBM) is the most common and malignant type of primary brain tumor, and 95% of patients die within 2 years after diagnosis. In this study, aiming to overcome chemoresistance to the first-line drug temozolomide (TMZ), we carried out research to discover a novel alternative drug targeting the oncogenic NFAT signaling pathway for GBM therapy. To accelerate the drug’s clinical application, we took advantage of a drug repurposing strategy to identify novel NFAT signaling pathway inhibitors. After screening a set of 93 FDA-approved drugs with simple structures, we identified pimavanserin tartrate (PIM), an effective 5-HT2A receptor inverse agonist used for the treatment of Parkinson’s disease-associated psychiatric symptoms, as having the most potent inhibitory activity against the NFAT signaling pathway. Further study revealed that PIM suppressed STIM1 puncta formation to inhibit store-operated calcium entry (SOCE) and subsequent NFAT activity. In cellula, PIM significantly suppressed the proliferation, migration, division, and motility of U87 glioblastoma cells, induced G1/S phase arrest and promoted apoptosis. In vivo, the growth of subcutaneous and orthotopic glioblastoma xenografts was markedly suppressed by PIM. Unbiased omics studies revealed the novel molecular mechanism of PIM’s antitumor activity, which included suppression of the ATR/CDK2/E2F axis, MYC, and AuroraA/B signaling. Interestingly, the genes upregulated by PIM were largely associated with cholesterol homeostasis, which may contribute to PIM’s side effects and should be given more attention. Our study identified store-operated calcium channels as novel targets of PIM and was the first to systematically highlight the therapeutic potential of pimavanserin tartrate for glioblastoma.
Similar content being viewed by others
Introduction
Glioblastoma multiforme (GBM), a type of high-grade (grade IV) glioma originating from the neuroepithelium, is the most malignant primary brain tumor. The 5-year survival rate of GBM patients is only 5% [1]. Due to its extensive infiltrative growth behavior [2], it is almost impossible to remove GBM tumor lesions completely through surgical resection without affecting normal brain function. Therefore, effective chemotherapeutic drugs are important for improving patient survival. Currently, few drugs have been approved for GBM therapy. Temozolomide (TMZ) and several other alkylating agents, including lomustine and carmustine, are currently applied as first-line drugs in the clinic [1]. However, patients often acquire chemoresistance to these drugs after long-term administration, and these drugs become less effective especially in recurrent tumors. In the past few decades, intense efforts have been made to explore the molecular pathogenesis of GBM tumors, and many promising therapeutic targets for GBM, including MET signaling [3], AMP kinase, the PAX6/DLX5-WNT5 pathway [4], the PI3K-AKT pathway, EGFR, and MMPs, have been identified. Some agents targeting these proteins or pathways have entered clinical trials. However, patient survival still has not been substantially improved, possibly because of the high heterogeneity and invasive profile of GBM. Therefore, effective drugs with a novel mechanism of action remain an urgent need in the clinic.
The nuclear factor of activated T cells (NFAT) family is a crucial family of transcription factors originally described in immune T cells and subsequently found to be ubiquitously expressed in various types of immune and nonimmune cells [5]. NFAT signaling plays an essential role in diverse biological processes, including immune cell activation and differentiation, cell cycle progression, organogenesis, etc. However, accumulating evidence has revealed that overexpression or hyperactivation of NFATs is often correlated with poor prognosis in a wide range of cancer types, including lymphoma [6], ovarian cancer, and breast cancer [7]. In glioblastoma, after analysis of a large number of clinical samples, it was found that NFAT was also overexpressed in GBM tumors compared with low-grade gliomas [8], a property that may promote the invasion of glioblastoma cells. In addition, many oncogenic proteins associated with cancer cell proliferation (e.g., c-MYC and MDM2), metastasis (e.g., COX-2 and ATX), inflammation (e.g., COX-2) or angiogenesis (e.g., VEGFR1 and interleukins) can be upregulated by NFAT transcription factors [5], and most of these oncogenic proteins have also been observed to be overexpressed in glioblastoma cells. Additionally, upstream oncogenic signals, such as aberrant growth factor (VEGFA, autotaxin, and acetyl-CoA) [9] and Ca2+ signals [10], can also be integrated with NFAT signaling in glioblastoma cells to promote tumorigenesis and tumor progression. Therefore, the NFAT signaling pathway constitutes a promising therapeutic target for cancers, including glioblastoma.
To date, five NFATs (NFAT1–NFAT5) have been identified, and the activation of all NFATs except NFAT5 is tightly controlled by store-operated calcium entry (SOCE) [11]. Briefly, in the resting state, NFATs are hyperphosphorylated and located in the cytosol. When stimuli bind to their cognate receptors (G-protein coupled receptors or receptor tyrosine kinases), phospholipase C is activated and hydrolyzes phosphatidylinositol 4,5-bisphosphate (PIP2) to release 1,4,5-phosphoinositol (IP3). Then, IP3 binds to the ER (endoplasmic reticulum)-resident calcium ion channel IP3R (IP3 receptor) and triggers Ca2+ depletion from ER stores, which then evokes SOCE. The sustained increase in the cytosolic [Ca2+] activates calcineurin to dephosphorylate NFATs, and dephosphorylated NFATs then translocate into the nucleus and modulate gene transcription. Targeting any node can inhibit the activation of NFATs.
Although some inhibitors targeting the NFAT signaling pathway have been reported, none of them have entered clinical trials for cancer therapy because of toxicity or other reasons. FDA-approved drugs are of great value to humans, and their pharmacodynamic profiles and safety have been well characterized [12]. Hence, developing new drugs by repurposing these drugs for new indications can effectively reduce the research time frame (3–8 years), cost (~300 million dollars) and risk of failure. In this study, we adopted a drug repurposing strategy to identify novel NFAT signaling pathway inhibitors, hoping to discover a drug that could gain rapid use for clinical GBM therapy.
Materials and methods
Chemicals
All 93 compounds (Table S1) described in the drug screening library were obtained from TargetMol. All other reagents were purchased from TargetMol unless otherwise indicated.
Cell lines and culture conditions
The human GBM cell lines U87-MG, U118, A172, T98G, and LN-229 and the human normal breast cell line MCF-10A were purchased from Procell. The NFAT1(1-460)-GFP stably transfected HeLa cell line and stably transfected luciferase U87 cell line were established as described in a previous paper [13]. All cell lines were cultured in DMEM medium supplemented with 10% (v/v) FBS (Sigma) in an atmosphere of 5% CO2 and 95% air unless otherwise indicated.
Plasmids
The GCaMP6m-XC (#111543), Opto-CRAC (#101245), Human STIM1-CFP (#19755), ER-GCamp6-150 (#86918), and NFAT-GFP (#11107) plasmids were purchased from Addgene. Constitutive activation of mCherry-calcineurin was established via the method described in a previous paper [13]. To prepare the mCherry-STIM1 constructs, the full-length complementary DNA (cDNA) of human STIM1 was amplified from STIM1-CFP by PCR and was then subcloned into the pcDNA3.1+ vector between the XhoI and NotI sites, with the mCherry sequence inserted between two additional BamHI and Not1 sites introduced immediately downstream of residue Asn 38.
NFAT nuclear translocation assay with 93 compounds selected from the FDA-approved Drug Library
HeLa cells stably transfected with NFAT1-GFP were seeded in 35-mm glass-bottom dishes at a density of ~20,000 cells per dish and were then cultured overnight. For preliminary screening, 93 compounds selected from the TargetMol FDA-approved drug library were added to each dish at a final concentration of 100 μM and incubated for 30 min. Subsequently, 10 μM TG (thapsigargin, a noncompetitive inhibitor of sarco/ER Ca2+ ATPase) and 1 mM Ca2+ were added to each dish and incubated with the cells for another 20 min. Finally, the dishes were gently washed with PBS three times, and the cells were fixed with 4% paraformaldehyde (PFA) solution (1000 μL per dish) for 15 min at room temperature in the dark and gently washed twice with PBS. Then, 1000 μL of PBS was added to each dish for imaging in a Lecia SP8 confocal laser scanning microscope system (objective lens: ×63). The ratio of the green fluorescence intensities in the nucleus and cytoplasm of each cell was calculated by ImageJ analysis software. For normalization, this ratio in cells treated with TG/Ca2+ and DMSO was set as 1, while that in cells without TG treatment was set as 0. A lower ratio indicated higher inhibitory activity against NFAT nuclear translocation.
The inhibition curves of compounds that completely inhibited NFAT nuclear translocation at 100 μM were further determined by the procedures described above. However, compounds were prepared at a series of concentrations (200, 100, 50, 25, 12.5, 6.25, 3.13, 1.56, 0.78, and 0.39 μM) and added to each dish separately. In addition, after fixation with 4% PFA, cells were permeabilized with 0.5% Triton X-100 (1000 μL per well) and stained with DAPI (1 μg/mL) for 5 min at room temperature. Finally, dose-response data were exported into GraphPad Prism 7 to calculate the IC50 values.
Thapsigargin-induced and light-operated Ca2+ influx assay
U87 cells or HeLa cells were grown in double antibody-free MEM supplemented with 10% FBS at 37 °C in a 5% CO2 atmosphere. Before transfection, cultured cells were seeded in a 35 mm glass-bottom dish and cultured overnight. The changes in cytoplasmic [Ca2+] induced by TG or light were measured with the green fluorescent genetically encoded calcium indicator GCaMP6m-XC. GCaMP6m-XC fluorescence intensities, reflecting Ca2+ levels, were recorded on a Lecia SP8 confocal laser scanning microscope with a 488-nm laser. The resulting data collected with Leica Application Suite X were plotted and analyzed with GraphPad Prism. All experiments were conducted at 37 °C unless otherwise indicated and were repeated at least three times.
To monitor the influence of pimavanserin tartrate (PIM) on thapsigargin (TG)-induced Ca2+ influx in U87 and HeLa cells, 4 μg GCaMP6m-XC was transfected into U87 or HeLa cells, and Ca2+ signals were imaged 24 h after transfection. Prior to Ca2+ measurement, cells were incubated with DMSO, 10 μM PIM (U87 cells) or 40 μM PIM (HeLa cells) for 30 min at 37 °C and were then maintained in Ca2+-free D-Hanks balanced salt solution (HBSS, 400 mg/L KCl, 60 mg/L KH2PO4, 350 mg/L NaHCO3, 8000 mg/L NaCl, 121 mg/L Na2HPO4·12H2O, and 1000 mg/L D-glucose). Cytoplasmic Ca2+ levels were recorded for 1 min every 1.29 s after adding 1 μM TG. Then, 1 mM CaCl2 was added, and Ca2+ signals were recorded for another 3 min.
To measure the kinetic properties of PIM-mediated inhibition of SOCE, U87 cells were first treated with TG/Ca2+ to induce SOCE. When SOCE was stabilized, PIM (20 μM) was added, and Ca2+ levels were recorded by measuring the GCaMP6m-XC fluorescence intensity changes.
The experiment to evaluate the reversibility of PIM-mediated inhibition of SOCE was conducted as follows: After inhibition of SOCE by PIM, PIM was removed, and the cells were incubated in culture medium for 25 min. Then, the cells were maintained in Ca2+-free D-Hanks balanced salt solution as described above. Cytoplasmic Ca2+ levels were recorded after adding 10 μM TG followed by 1 mM CaCl2.
To monitor Ca2+ release induced by PIM, cells were maintained in Ca2+-free D-Hanks balanced salt solution as described above, and PIM was then added. Cytoplasmic Ca2+ levels were recorded for at least 2 min.
To test the effect of PIM on ER Ca2+ levels, 4 μg ER-GCamp6 was transfected into U87 cells using Lipofectamine 2000 (Thermo Fisher Scientific). After 24 h of transfection, PIM (20 μM) was added, and ER Ca2+ levels were recorded for at least 3 min by measuring the green ER-GCamp6 fluorescence intensity.
To measure the influence of temperature on PIM-mediated inhibition of SOCE, U87 cells were pretreated with 20 μM PIM at different temperatures (37, 33, 30, and 25 °C) and were then maintained in Ca2+-free D-Hanks balanced salt solution as described above. Cytoplasmic Ca2+ levels were recorded at the indicated temperatures after adding 1 μM TG followed by 1 mM CaCl2.
To evaluate the inhibitory activity of PIM against the ORAI channel, 5 μg Opto-CRAC and 4 μg GCaMP-XC were cotransfected into U87 cells using Lipofectamine 2000 (Thermo Fisher Scientific). Twenty-four hours after transfection, cells were incubated with DMSO or 25 μM PIM for 30 min. Because the excitation wavelength used to monitor GCaMP6m-XC fluorescence (488 nm) overlaps with the wavelength for photoactivation of Opto-CRAC, a 488-nm laser served as the blue light source to stimulate the ORAI channel and elicit Ca2+ influx. The mCherry-positive cells were selected for statistical analysis, and cytoplasmic Ca2+ levels were recorded every 1.29 s until the fluorescence was stabilized.
Confocal imaging
The cell lines used for imaging included HeLa cells stably transfected with NFAT1-GFP. Before imaging, cells were seeded in 35-mm glass-bottom confocal dishes and cultured overnight. Confocal imaging was performed on a Lecia SP8 confocal laser scanning microscope. The data were analyzed using GraphPad Prism 7 software. All experiments were repeated at least three times.
A competitive binding assay was applied to verify whether PIM inhibits TG-induced NFAT nuclear translocation directly by targeting the 5-HT2A receptor. Briefly, PIM (25 μM) was incubated with HeLa cells stably transfected with NFAT1-GFP for 30 min, and 10 μM TG and 1 mM Ca2+ were then added. Subsequently, the 5-HT2A receptor agonists 5-hydroxytryptamine (5-HT, 50 μM, Aladdin) and 2,5-dimethoxy-4-iodoamphetamine (DOI, Sigma-Aldrich, 50 μM) and the 5-HT2A receptor antagonist ketanserin (50 μM) were added to compete with PIM for binding to the 5-HT2A receptor for 20 min at 37 °C. The level of NFAT nuclear translocation to be restored was monitored with the Lecia SP8 microscope, and the data were analyzed with ImageJ and GraphPad Prism 7. The inhibitory activity of the inverse agonist nelotanserin (50 μM) against TG-induced NFAT nuclear translocation was also evaluated by the procedure described above.
To explore the influence of temperature on PIM-mediated inhibition of NFAT nuclear translocation, PIM (25 μM) was incubated with HeLa cells stably transfected with NFAT1-GFP at 25 °C for 30 min, and 10 μM TG and 1 mM Ca2+ were then added. BTP2 was used as a control. In addition, after inhibition of TG-induced NFAT nuclear translocation by PIM (25 μM) at 37 °C, the experimental dish was moved to a 25 °C environment and incubated for at least 15 min. The level of rescued NFAT nuclear translocation was recorded with the Lecia SP8 microscope.
To test whether PIM’s inhibitory effect on TG-induced NFAT nuclear translocation is reversible, 40 μM PIM was used to inhibit TG-induced NFAT nuclear translocation in HeLa cells stably transfected with NFAT1-GFP. Then, PIM was removed, and the cells were incubated in culture medium for 25 min. Finally, the cells were restimulated with 10 μM TG and 1 mM CaCl2. The level of rescued NFAT nuclear translocation was recorded with the Lecia SP8 microscope.
To explore whether PIM/Ca2+ can induce NFAT nuclear translocation, HeLa cells stably transfected with NFAT1-GFP were incubated with 40 μM PIM and 1 mM Ca2+ at room temperature and 37 °C. The levels of NFAT nuclear translocation were recorded with the Lecia SP8 microscope.
To explore the effect of PIM on STIM1 puncta formation after TG stimulation, 4 μg STIM1-CFP or mCherry-STIM1 was transfected using Lipofectamine 2000 (Thermo Fisher Scientific) into HeLa cells stably transfected with NFAT1-GFP. After 24 h of transfection, PIM (40 μM) was added and incubated with the cells for 30 min, and 10 μM TG was then added and incubated with the cells for another 20 min at 37 °C. Finally, the cells were fixed with 4% PFA for imaging with the Lecia SP8 microscope.
Cell proliferation assay
U87, U118, A172, T98G, and LN-229 cells and MCF-10A human normal breast cells (2 × 103 cells per well) were seeded in 96-well plates in a total volume of 100 μL. After culturing overnight, cells were treated with a series of PIM concentrations for 48 h at 37 °C in 5% CO2. The medium was then discarded, and 100 μL of fresh medium containing WST-8 (375 μM) and PMS (15 μM) was added to each well and incubated with the cells for another 1–4 h. The absorbance of each well was measured at 490 nm in a microplate reader. The survival rate was calculated with the following equation: survival rate = 100% × (ODtreated well − ODblank well)/(ODcontrol well − ODblank well). Survival curves were plotted using GraphPad Prism 7 software, and IC50 values (defined as the drug concentration required to inhibit growth by 50% relative to that of control cells) were calculated.
Cell cycle and cell apoptosis analysis
The cell cycle was analyzed using a Cell Cycle Assay Kit Plus (US Everbright® Inc., C6078-50T). U87 cells were treated with 3.5 or 7 μM PIM or left untreated at 37 °C in 5% CO2. After incubation for 24 h, cells were collected, and the cell suspension was centrifuged at 1000 r/min for 4 min and washed twice with ice-cold staining buffer. The cell pellets were resuspended in 1 mL of PBS. Then, 4 μL of RedNucleus I (a reagent in the kit) was added, and the cells were stained for 20 min at room temperature in the dark. The cellular DNA content was analyzed with an Image StreamX Mark II flow cytometer (Merck). The resulting data were then analyzed using IDEAS software version 6.2 and GraphPad Prism 7 software.
Apoptosis was evaluated using an Annexin V-FITC/PI (propidium iodide) Apoptosis Detection Kit (Beyotime, C1062-20T). Briefly, U87 cells were treated with PIM (7, 14 μM) or an equal amount of DMSO (control) for 24 h at 37 °C in 5% CO2. Then, the cells were collected and washed twice with ice-cold PBS. The cell suspension was centrifuged at 1000 r/min for 4 min and resuspended in 100 μL of 1× binding buffer supplied in the kit. Subsequently, 5 μL of Annexin V-FITC and 5 μL of propidium iodide were added, and the cell suspension was stained at room temperature for 15 min in the dark. After that, 400 μL of 1× binding buffer was added, and the sample was analyzed in the Image StreamX Mark II flow cytometer (Merck). The resulting data were analyzed using IDEAS software version 6.2 and GraphPad Prism 7 software.
Wound-healing assay
The ability of PIM to inhibit cell migration was assessed by a wound-healing assay. In brief, U87 glioma cells were seeded in a plastic-bottom six-well plate. After culturing overnight, the cell monolayer was scratched with a 200 μL pipette tip to generate a wound gap. Then, U87 cells were further cultured in the presence or absence of PIM (7 μM), and images were acquired at 0 and 24 h. The number of cells that migrated into the wound area was calculated with ImageJ software, and the data were analyzed with GraphPad Prism 7.
Real-time monitoring of cell motility
U87 cells were seeded in a black, glass-bottom six-well plate. Then, the plate was mounted for imaging in a PerkinElmer high-content imaging system in phase contrast mode (objective lens: ×40), and cells were cultured with 7 μM PIM or an equivalent volume of DMSO (control) at 37 °C in 5% CO2. Images were acquired every 30 min for 4 h. The displacement values and the generations of cells at each time point were analyzed using Harmony 4.9 software, and the data were exported as a txt file. Each cell migration trajectory was plotted using MATLAB software, and the resulting data were statistically analyzed with GraphPad Prism 7 software. All cells in one imaging field were included in the statistical analysis, and at least five fields of view were recorded per group.
RNA-sequencing and data analysis
U87 cells were treated with 7 μM PIM or the same volume of MEM containing an equal amount of DMSO for 24 h at 37 °C in 5% CO2. Then, cells (>1 × 106) were collected, and total RNA was extracted using TRIzol reagent (Invitrogen) according to the manufacturer’s instructions. To guarantee that the following experiments proceeded smoothly, the quality of the extracted total RNA was further evaluated using a Fragment Analyzer (Thermo Fisher Scientific) with a Standard Sensitivity RNA Analysis Kit (15 nt). Subsequently, mRNA molecules were purified with oligo(dT)-coupled magnetic beads (Invitrogen Thermo Fisher Scientific, USA) and fragmented into small pieces. A cDNA library was constructed with a BGIEasy DNA Library Prep Kit (BGI, Shenzhen). mRNA-seq was performed on the BGISEQ 500 platform (Shenzhen, China). Each sample was analyzed in biological triplicate.
The original sequencing data were filtered using SOAPnuke (version 1.5.2) to remove reads with sequencing adapters, low-quality base ratios and unknown bases with high N ratios. HISAT2 (version 2.0.4) was applied to map the clean reads against the reference genome (GCF_000001405.38_GRCh38.p12). The clean data were aligned with the reference coding gene set using Bowtie2 (version 2.2.5), and the expression of genes was quantified with RSEM (version 1.2.12). The differential gene expression levels between the PIM treatment group and DMSO control group were statistically analyzed with DESeq2 (v1.4.5), and genes with a log2 | fold change | ≥ 0.4 and P value < 0.05 were considered significantly differentially expressed. The expression fold changes of all identified genes (17,797) were plotted with P values using MATLAB. A heatmap scaled by the Z-scores of the expression levels of significantly differentially expressed genes was also plotted in MATLAB. Functional and pathway enrichment analyses of differentially expressed genes were performed using online GSEA tools with the Hallmark, KEGG_pathway and PID_pathway gene set categories (https://www.gsea-msigdb.org/gsea/msigdb/annotate.jsp), and the analysis results were visualized on bubble or bar charts with Origin (version 2019) and GraphPad Prism 7 software.
iTRAQ proteomics and data analysis
U87 cells were treated with 7 μM PIM or the same volume of MEM containing an equal amount of DMSO for 24 h at 37 °C in 5% CO2. Then, the cells were collected and gently washed with PBS at least three times. Total protein was extracted using the following procedure. First, the cell pellets were incubated on ice for 5 min with L3 lysis buffer (8 M urea and 40 mM Tris-HCl (pH 8.5)) containing the protease inhibitor PMSF (1 mM) and 2 mM EDTA. Then, 10 mM DTT was added, and the cells were lysed by ultrasonication (200 W, 15 min). The obtained cell lysate was centrifuged at 25,000 × g and 4 °C for 15 min. To further reduce disulfide bonds, the supernatant was incubated with 10 mM DTT at 56 °C for 1 h and was then mixed with 55 mM IAM and incubated for 45 min in the dark to block cysteine thiol groups. Protein was precipitated using chilled acetone (2 h, −20 °C). The precipitated protein was air-dried and redissolved in 0.5 M TEAB. The protein sample solution was then centrifuged at 25,000 × g and 4 °C for 15 min. The protein concentration in the supernatant was determined using a Bradford Assay Kit (Bio-Rad).
The extracted proteins were digested with trypsin. The obtained peptides were dried by vacuum centrifugation and redissolved in 0.5 M TEAB. Then, the peptides were labeled with isobaric tags according to the protocol for the 8-plex iTRAQ reagent (Applied Biosystems). The iTRAQ-labeled peptide mixtures were fractionated using an LC system (Shimadzu, LC-20AB). Each fraction was further analyzed by HPLC-ESI-MS/MS (UltiMate 3000 UHPLC, Thermo Fisher Scientific; Q-Exactive HFX, Thermo Fisher Scientific, San Jose, CA). The raw MS data were converted into MGF (Mascot Generic File) format files using Proteome Discoverer version 1.4 (Thermo). Subsequently, the MGF files were searched and aligned against the 6997_ncbi_homo database (81064 sequences). Only unique peptides were used for protein quantification, and iTRAQ quantitation was performed with IQuant software developed by BGI (Shenzhen, China). Each sample was analyzed in biological duplicate. For each replicate, proteins were considered significantly up- or downregulated when the ratio of their expression level in the drug treatment group to that in the control group was ≥1.2 or ≤0.83 (q value < 0.05). Proteins that were considered differentially expressed in at least one replicate with a mean ratio ≥1.1 or ≤0.9 were defined as the final set of differentially expressed proteins. Functional enrichment analysis of the differentially expressed proteins was performed using online GSEA tools with the hallmark gene set.
Immunoblotting
After treatment with 7 or 10 μM PIM or with saline solution for 24 h, cells were collected and lysed with RIPA buffer on ice for 10 min. The lysate was then centrifuged at 12,000 × g for 20 min at 4 °C. The protein concentrations in the supernatants were determined using a BCA kit (Beyotime, China). The lysates were separated by SDS-PAGE, transferred to PVDF membranes and probed with antibodies against different proteins (anti-PCNA, BOSTER, Cat No. #BM0104; anti-NPC2, BOSTER, #A1058-2).
Animals
All animal experiments were approved by the Ethics Committee and IACUC of the Third Military University and were conducted according to the guidelines for laboratory animal care and use. BALB/c nude mice 6–8 weeks of age and of either sex were supplied by the Animal Center of the Third Military University (Chong Qing). A subcutaneous glioblastoma xenograft model was established by implanting 100 μL of U87 cells (a density of 5 × 104 cells/μL) in the posterior limbs of mice. Mice were then randomized into the vehicle control (n = 15) and PIM treatment groups (n = 15). Pimavanserin tartrate (Shanghai Biochempartner Co., Ltd.) was administered via gavage every day for 3 weeks at a dose of 10 mg/kg body weight, and mice in the control group received an equal amount of saline solution. Each week, five mice were sacrificed, and solid tumors were isolated. The tumor dimensions were measured with a Vernier caliper, and the tumor volume was calculated with the following equation: TV (cm3) = L × W2 × 1/2, where L is the largest tumor diameter and W is the perpendicular tumor diameter. H&E staining and immunohistochemical staining of DNA fragments (TUNEL, Abcam, #ab206386), activated caspase-3 (red fluorescence-positive, Cell Signaling, #9664), Cyclin B2 (green fluorescence-positive, Abcam, #ab185622), and NFATc3 (CST, #4998) were performed in subcutaneous GBM xenograft tumor tissues harvested from mice treated with vehicle or PIM for 3 weeks.
To generate the orthotopic GBM model in the brain, 10 μL (5 × 104 cells/μL) of a U87-luc cell suspension was injected into the brain with a Hamilton syringe (180 μm needle) using a micropump system. Then, animals with established tumors were divided into two groups: the pimavanserin tartrate treatment group (n = 5, 10 mg/kg per day, gavage administration) and the control group (n = 5, equal amount of saline solution, gavage administration). Tumor growth was monitored each week using bioluminescence imaging as follows: mice were anesthetized with isoflurane and injected with D-luciferin (120 mg/kg, i.p.), and after 10 min, bioluminescence signals in tumors were recorded on a Xenogen IVIS System.
Statistical analysis
To determine the significance of differences between groups, comparisons were made with Student’s t test. For all statistical tests, a significance level of 0.05 was considered statistically significant. In all figures, *P < 0.05 and **P < 0.01 by unpaired Student’s t test.
Results
Identification of pimavanserin tartrate as a potent NFAT pathway inhibitor
To date, thousands of drugs have been approved for clinical use, and screening all drugs is a burdensome task. Considering that identified drugs may be used as lead compounds for further structural optimization, we selected 93 compounds from the TargetMol FDA-approved drug library that had simple and easily modified structures to establish the screening library (Table S1). Thapsigargin (TG), an effective inhibitor of the Ca2+-ATPase (SERCA) on the ER, can passively deplete Ca2+ from ER stores, triggering SOCE and subsequent NFAT nuclear translocation. Therefore, to evaluate the inhibitory activity of selected compounds against the NFAT signaling pathway, we established a TG-induced NFAT nuclear translocation assay using the NFAT1-GFP stably transfected HeLa cell line; the screening process is shown in Fig. S1f. The ratio of the green fluorescence intensity in the nucleus to that in the cytoplasm reflects NFAT activity, and a lower ratio indicates higher inhibitory activity against NFAT nuclear translocation. Preliminary screening results revealed that 8 compounds completely inhibited TG-induced NFAT nuclear translocation at 100 μM (Figs. 1a and S1a) and that only sertaconazole and pimavanserin tartrate (PIM) showed partial inhibitory activity at 10 μM. Subsequently, the half-maximum inhibitory concentration (IC50) values of these eight drugs were determined using the same method, and it was found that PIM displayed the highest inhibitory activity, with an IC50 value of 6.47 μM (Fig. 1c), and that 25 μM PIM completely inhibited NFAT nuclear translocation (Fig. S1c). In addition, PIM suppressed NFAT nuclear translocation triggered by the upstream signaling molecule ATP (Fig. S1b). Moreover, PIM inhibited NFAT nuclear translocation in U87 cells in a concentration-dependent manner (Fig. 1d). Therefore, we selected PIM for further study (Fig. 1b). Interestingly, the inhibitory activity of PIM against the NFAT signaling pathway was temperature sensitive. PIM inhibited NFAT nuclear translocation at 37 °C, but this inhibition was reduced or abolished at colder temperatures (25 °C, Figs. 1e and S1d). By contrast, the reported NFAT signaling pathway inhibitor BTP2 (Fig. S1e) had no such properties, implying that PIM inhibited NFAT nuclear translocation through a unique mechanism and that subsequent experiments should be conducted at 37 °C.
a The primary inhibitory activity of 93 compounds (at 100 μM) selected from the FDA-approved drug library against NFAT nuclear translocation. The ratio of green fluorescence intensity in the nucleus to that in the cytoplasm was normalized as follows: NC, the negative control without thapsigargin (TG) stimulation, was set as 0; PC, the positive control with TG stimulation, was set as 1. A lower ratio indicated higher inhibitory activity. b Chemical structure of pimavanserin tartrate. c Inhibition curves of the eight identified potent NFAT signaling pathway inhibitors. Relative to the other compounds, PIM showed the most potent inhibitory effect on NFAT nuclear translocation, with an IC50 value of 6.47 μM. The data are shown as the mean ± S.D. values (n = 4). d Representative confocal images of NFAT1-GFP-transfected U87 cells after incubation with DMSO (control) or PIM (20 μM). TG (10 μM) was added to induce store depletion and subsequent NFAT nuclear translocation. Scale bar, 25 μm. The data are shown as the mean ± S.D. values (n = 3). e Representative confocal images of HeLa cells stably transfected with NFAT1-GFP after incubation with PIM (25 μM) at 37 or 25 °C. The inhibitory activity of PIM against NFAT nuclear translocation was lost at 25 °C.
Pimavanserin tartrate inhibits NFAT activity independent of the 5-HT2A receptor
Pimavanserin tartrate is an atypical antipsychotic that was approved for the treatment of psychosis accompanied by Parkinson’s disease, including hallucinations and delusions, and it achieves its therapeutic effects mainly through its potent and highly selective inverse agonist activity against the 5-HT2A receptor [14]. Therefore, we first hypothesized that PIM may target the 5-HT2A receptor to suppress NFAT nuclear translocation and that the reduced intrinsic activity of the 5-HT2A receptor may result in lower IP3 levels in the cytosol, which further elevate the [Ca2+] in ER stores and inhibit SOCE. To confirm this hypothesis, we performed a competitive binding assay in which the potent 5-HT2A receptor antagonist ketanserin and agonists DOI and 5-HT, which have no inhibitory activity against NFAT translocation (Fig. S2b), were used individually to compete with PIM for 5-HT2A receptor binding. The results showed that inhibition of NFAT nuclear translocation induced by PIM could not be rescued by these high-affinity ligands of the 5-HT2A receptor (Fig. 2a). In addition, another reported 5-HT2A receptor reverse agonist, nelotanserin (Nelo), was incapable of inhibiting NFAT nuclear translocation induced by TG (Fig. 2b). The screening library above also contains several 5-HT receptor ligands, and similarly, not all of these 5-HT receptor antagonists could inhibit TG-initiated NFAT translocation (Fig. S2a). Collectively, these results demonstrated that blocking the 5-HT receptor might not be sufficient to inhibit TG-induced NFAT translocation and that PIM inhibited NFAT translocation via a mechanism other than targeting the 5-HT2A receptor. In addition, pimavanserin tartrate also exhibited binding affinity for sigma1 receptors and adrenergic receptors. However, competitive binding assays also revealed that NFAT nuclear translocation inhibition induced by PIM was not rescued by these receptors’ high-affinity ligands (cutamesine and methoxamine) (Fig. S2c), indicating that PIM inhibited NFAT activity through a novel mechanism different from that through which it inhibits the reported targets.
a Competitive binding assay with PIM (25 μM), the 5-HT receptor antagonist ketanserin (50 μM) and the 5-HT receptor agonists DOI (50 μM) and 5-HT (50 μM). Substitution with a high-affinity 5-HT receptor ligand could not restore TG-induced NFAT nuclear translocation in the presence of PIM. Scale bar, 20 μm. The data are shown as the mean ± S.D. values (n = 3). b Representative confocal images of HeLa cells stably transfected with NFAT-GFP and treated with PIM (25 μM) or nelotanserin (Nelo; 50 μM), a potent inverse agonist of the 5-HT2A receptor that could not inhibit TG-induced NFAT nuclear translocation. Scale bar, 20 μm.
Pimavanserin tartrate inhibits store-operated calcium entry and subsequent NFAT nuclear translocation
Activation of NFATs requires a series of upstream signals. The above results prompted us to further explore the mechanism of action of PIM by assessing how it might affect each critical node in the NFAT signaling pathway. We first generated a construct with constitutively activated calcineurin (mCherry-caCnA), which could spontaneously lead to NFAT nuclear translocation independent of upstream signaling. After treatment with PIM, spontaneous NFAT nuclear translocation was not observed in stable NFAT1-GFP-transfected HeLa cells transfected with mCherry-caCnA (Fig. 3a), implicating that PIM might target the signaling node upstream of calcineurin. Therefore, we then assessed the impact of PIM on an upstream signal—the SOCE current—using the green fluorescent genetically encoded calcium indicator GCaMP6m-XC [15], and the results showed that TG-triggered SOCE calcium influx was obviously inhibited by PIM in both U87 and HeLa cells (Figs. 3b and S3a). As described above, SOCE is essential for NFAT activation, and a decrease in SOCE can contribute to PIM-mediated inhibition of NFAT activity.
a The effect of PIM on NFAT nuclear translocation (green) spontaneously initiated by a constitutively active form of calcineurin (caCnA; red). mCherry-tagged caCnA was transfected into HeLa cells stably transfected with NFAT-GFP to induce nuclear entry of NFAT-GFP. Transfected cells were treated with either DMSO or PIM (25 μM). Scale bar, 20 μm. b The effect of PIM on thapsigargin (TG)-induced Ca2+ influx. GCaMP6m-XC was used as the calcium indicator and was transfected into U87 cells. Transfected cells were treated with either DMSO (control) or PIM (10 μM). Empty bar (−), 0 mM Ca2+ in the external medium. Orange bar (+), 1 mM Ca2+ was added. Blue bar (+), 10 μM TG was added to trigger calcium depletion from ER stores. The data are shown as the mean ± S.D. values (n = 24). ***P < 0.001. c The kinetics of PIM’s inhibitory effects on SOCE. Empty bar (−), 0 mM Ca2+ in the external medium. Orange bar (+), 1 mM Ca2+ was added. Blue bar (+), 10 μM TG was added to trigger calcium depletion from ER stores. Gray bar (+), 20 μM PIM was added to inhibit SOCE. Histograms showing the time needed to start to decrease SOCE (Gray) and the time for inhibition to plateau. The data are shown as the mean ± S.D. values (n = 14). d The Ca2+ levels at the Ca2+ phase peak are shown as the mean ± S.D. values (n > 17). ***P < 0.001, **P < 0.01. e Representative curves of PIM-induced calcium release from intracellular Ca2+ stores. Empty bar (−), 0 mM Ca2+ in the external medium. Gray bar (+), 20 μM PIM was added to trigger calcium release. f The effect of PIM on ER Ca2+ levels. PIM (20 μM) gradually lowered ER Ca2+ levels. g Representative curves of thapsigargin (TG)-induced Ca2+ influx after pretreatment with PIM (20 μM) at different temperatures (37, 33, 30, and 25 °C). GCaMP6m-XC was used as the calcium indicator and was transfected into U87 cells. Empty bar (−), 0 mM Ca2+ in the external medium. Orange bar (+), 1 mM Ca2+ was added. Blue bar (+), 10 μM TG was added to trigger calcium depletion from ER stores. h The Ca2+ levels at the SOCE peak at different temperatures are shown as the mean ± S.D. values (n > 7). ***P < 0.001, **P < 0.01.
In the experiments described above, we determined the inhibitory activity of PIM against TG-induced NFAT nuclear translocation or SOCE by pretreating cells with PIM for 30 min. Interestingly, if NFAT nuclear translocation was first triggered by TG/Ca2+, and PIM was then added to the cell culture medium, NFAT nuclear translocation was still abolished by PIM (Fig. S3h). Therefore, we further characterized the kinetics of PIM-mediated inhibition of SOCE (Fig. 3c). The results showed that PIM did not immediately induce a decrease in SOCE; instead, ~4.2 min was required for PIM to begin to decrease SOCE. After ~8.5 min, the inhibition plateaued. This result implied that the binding between PIM and its target might be a slow process. In addition, PIM’s inhibitory effects on SOCE and TG-induced NFAT nuclear translocation were restored after removal of PIM, indicating that PIM’s inhibitory effect on SOCE was reversible (Fig. S3f, g). In the SOCE inhibition experiments, the level of TG-induced Ca2+ release was lower after treatment with PIM than that in the control group (Fig. 3d). To confirm whether this abolition resulted from Ca2+ release induced by PIM, we recorded Ca2+ levels after adding PIM to Ca2+-free cell culture medium. The results revealed that PIM increased cytosolic Ca2+ to a level similar to that after TG treatment, indicating that PIM-induced Ca2+ release from intracellular Ca2+ stores (Fig. 3d, e). However, the ability of PIM to induce Ca2+ release was weaker than that of TG. To further test whether this Ca2+ originated from the ER Ca2+ store, we transfected the ER calcium ion indicator ER-GCamp6 into U87 cells and then treated the transfected cells with PIM. The results showed that PIM gradually lowered the ER Ca2+ level (Fig. 3f), verifying our hypothesis. In particular, at colder temperatures, which abolished the inhibitory activity of PIM against SOCE, PIM/Ca2+ induced NFAT nuclear translocation similar to TG/Ca2+. However, at 37 °C, PIM/Ca2+ did not induce NFAT nuclear translocation, an effect that may be attributed to inhibitory activity downstream of ER Ca2+ depletion. As PIM’s inhibitory effect on TG-induced NFAT nuclear translocation was temperature sensitive, PIM’s inhibitory effect on SOCE was also temperature sensitive (Fig. 3g, h). With decreasing temperature, the inhibitory effect of PIM on SOCE was gradually reduced.
Pimavanserin tartrate prevents STIM1 puncta formation to inhibit store-operated calcium entry
The major store-operated channel (SOC) is the calcium release-activated calcium (CRAC) channel, which consists of two critical components: ER-resident stromal interaction molecules (STIM1) and ORAI calcium channels on the cell membrane [16]. STIM1 can sense the decrease in the calcium concentration in ER stores, forming oligomers and translocating toward ER-PM junctions to activate ORAI channels [17, 18]. Hence, we subsequently used light-operated CRAC constructs generated in our previous laboratory [19], which can allow ORAI channels to be opened by blue light without requiring ER store calcium depletion, to test the inhibitory activity against ORAI channels. However, after illumination with blue light, there was no significant difference in the calcium concentration increase between the control and PIM treatment groups (Fig. 4a). In addition, PIM did not affect light-induced NFAT nuclear translocation (Fig. 4b), indicating that the signaling node that PIM targets may be upstream of ORAI. Ionomycin, a calcium ionophore, can actively transport calcium from ER stores to the cytosol [20], triggering STIM1 oligomerization and subsequent CRAC channel-mediated calcium influx. The following assay revealed that PIM also inhibited ionomycin-induced NFAT translocation (Fig. 4c), implying that PIM might affect the STIM1 oligomerization process but not the upstream signals of STIM1. To verify this hypothesis, we expressed STIM1-CFP in HeLa cells stably transfected with NFAT1-GFP. Transfected cells were first incubated with PIM or an equal amount of DMSO and were then treated with TG. As expected, in cells in which NFAT nuclear translocation was inhibited, the formation of STIM1 puncta was significantly suppressed by PIM (Figs. 4d and S3b). Interestingly, it was serendipitously found that in some cells, PIM’s inhibitory effect on NFAT translocation was substantially rescued by overexpression of STIM1. In particular, when amino terminal mCherry-labeled STIM1 (mCherry-STIM1) was expressed, TG-induced NFAT nuclear translocation inhibition was hardly observed in mCherry-STIM1-positive cells (Fig. S3d). In addition, in the high-resolution imaging field, annular STIM1 puncta structures were commonly observed in the control group but not in the PIM-treated group (Fig. S3e). Chen et al. reported that ESYT1 was involved in the formation of these annular structures [21], and the direct target of PIM for inhibiting SOCE might be ESYT1. However, many studies have reported that knocking out ESYT1 can significantly reduce the formation of STIM puncta and ER-PM junctions but does not significantly influence SOCE in any cells except T cells [21,22,23]. Since PIM inhibited SOCE in a temperature-sensitive manner, we speculated that these contradictory results may result from the experimental conditions, a possibility that will be studied in detail in our subsequent work.
a The effect of PIM on light-operated ORAI-mediated calcium influx. Opto-CRAC (mCherry red) and GCaMP6m-XC (green) were cotransfected into U87 cells, which were then treated with DMSO (control) or PIM (20 μM). Scale bar, 10 μm. The data are shown as the mean ± S.D. values (n = 3). b The effect of PIM on NFAT nuclear translocation triggered by light-operated calcium influx. The Opto-CRAC plasmid labeled with mCherry was transfected into HeLa cells stably expressing NFAT1-GFP, which were then treated with DMSO (control) or PIM (25 μM). Blue light was used to trigger calcium influx and subsequent NFAT nuclear translocation. Scale bar, 20 μm. c The effect of PIM on ionomycin-induced NFAT nuclear translocation. HeLa cells stably expressing NFAT1-GFP were treated with DMSO or PIM (25 μM), and 3 μM ionomycin was added to induce Ca2+ depletion and subsequent nuclear translocation of NFAT. The data are shown as the mean ± S.D. values (n = 3). Scale bar, 20 μm. d The effect of PIM on the formation of STIM1 puncta. STIM1-CFP was transfected into HeLa cells stably expressing NFAT1-GFP. Transfected cells were treated with DMSO or PIM (40 μM), and TG was then added to induce Ca2+ depletion and subsequent STIM1 puncta formation. Scale bar, 10 μm. Note that in cells in which NFAT nuclear translocation was inhibited, the formation of STIM1 puncta was significantly suppressed by PIM. e Schematic diagram of the mechanism underlying PIM’s inhibitory effect on SOCE and subsequent NFAT translocation.
Pimavanserin tartrate impedes the growth, migration, and motility of glioblastoma cells in vitro
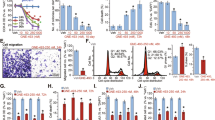
Next, we evaluated the antitumor activity of PIM using a GBM cell line. After treatment of U87 glioblastoma cells with 10 μM PIM for 48 h, a pronounced reduction in cell growth was observed (Fig. 5a). Subsequently, the proliferation inhibition curves of PIM in different glioblastoma cell lines were further measured via a WST-8-based proliferation assay. The results showed that PIM significantly suppressed the growth of GBM cells with an IC50 ranging from 1.46 to 8.07 μM, while it had little effect on the growth of normal MCF-10A cells, thereby attesting to its lower toxicity to normal cells (Fig. 5b) and broad-spectrum antiglioblastoma activity. To investigate the mechanism underlying the antiproliferative activity of PIM, we further performed flow cytometric cell cycle and apoptosis assays using the U87 cell line. As shown in Fig. 5c, PIM-induced significant G1/S phase arrest, and the percentage of G1/S phase cells increased from 64% to 84% and 81% upon treatment with 3.5 and 7 μM PIM, respectively, for 24 h. Apoptosis was also obvious, and the percentage of late apoptotic cells increased from 7.54% to 43.77% (Fig. 5d). The flow cytometer that we used was equipped with an imaging system; therefore, images of cells in each quadrant could be acquired. The acquired images indicated that cells in the third quadrant underwent apoptosis (Figs. 5d and S4c). Apoptotic cells have a small nuclear area and high bright-field contrast [24], and further analysis indicated that PIM promoted apoptosis (Fig. S4a). In addition, after treatment with 14 μM PIM for 24 h, almost all the cells were apoptotic (Fig. S4b). A high infiltration profile signifies one of the most malignant phenotypes; therefore, we further evaluated the effect of PIM on cell migration using a wound-healing assay. As depicted in Fig. 5e, PIM significantly inhibited the migration ability of U87 cells compared with control cells, and the average number of cells in the wound area was decreased by ~62.43%. In parallel, the migration trajectory of U87 cells in the presence and absence of PIM was monitored with a PerkinElmer high-content imaging system. The migration velocity and displacement were significantly reduced by PIM (Fig. 5f). In addition, we used Harmony software to analyze the inhibitory effect of PIM on cell division, and the ratio of dividing cells was markedly reduced by an average of 6% in the presence of PIM within the measurement period (Fig. 5g). Collectively, these results indicate that PIM displayed potent antitumor activity against GBM cells in vitro.
a Comparison of the growth rate of U87 cells after treatment with DMSO or 10 μM PIM for 48 h. Scale bars, 25 μm. b The dose-dependent inhibition curve of PIM on the proliferation of different glioblastoma cell lines (U87, T98G, A172, LN-229, and U118) or normal MCF-10A cells after incubation for 48 h. Cell proliferation inhibition was determined by a WST-8-based assay. PIM significantly suppressed the proliferation of these glioblastoma cell lines with IC50 values ranging from 1.46 to 8.07 μM, while PIM showed little effect on normal MCF-10A cells (mean ± S.D.; n = 3). c Cell cycle analysis of U87 cells after treatment with DMSO (control), 3.5 μM PIM or 7 μM PIM for 24 h. DNA was stained with RedNucleus I and was then analyzed by flow cytometry. Note that PIM-induced G1/S phase arrest. Inset: representative images of cells in different cell cycle phases. d Apoptosis analysis of U87 cells after treatment with DMSO (control) and 7 μM PIM for 24 h. Apoptosis was assessed by flow cytometry with Annexin V-FITC/PI double staining. The representative flow cytometry plots in the top row show the cell population distribution: Annexin V/PI double-negative cells (live cells), Annexin V single-positive cells (early apoptotic cells), Annexin V-FITC/PI double-positive cells (late apoptotic cells) and PI single-positive cells (dead cells). Histograms showing the results of quantitative analysis of apoptotic and dead cells in the total cell population (mean ± S.D.; n = 3), ***P < 0.001. Images of late apoptotic cells. e The effect of PIM on cell migration. After treatment with DMSO (control) or 7 μM PIM for 24 h, the cell migration rate was assessed by a wound-healing assay. Scale bars, 100 μm. The histogram shows the percentage of cells that migrated into the wound gap area (mean ± S.D., n = 4). **P < 0.01. f The effect of PIM on cell mobility. The diagram shows the representative migration trajectory of U87 cells over 4 h in the presence of DMSO (control) or 7 μM PIM. The top bar graph shows the normalized cell migration distance (mean ± S.D., n = 5). ***P < 0.001, **P < 0.01. The bottom bar graph shows the migration velocity (mean ± S.D., n = 5), *P < 0.05. g The effect of PIM on cell division. Compared with that in the DMSO control group, the number of dividing cells was obviously decreased by treatment with 7 μM PIM (mean ± S.D., n = 5). **P < 0.01.
Molecular signature of pimavanserin tartrate in glioblastoma cells as assessed by transcriptomic and proteomic profiling
PIM was approved for market release in 2016, and few studies have revealed its antitumor activity. The molecular mechanism underlying its antitumor activity has not been well elucidated. To gain an unbiased understanding of the mechanism of PIM’s action in GBM cells, we performed transcriptomic and proteomic analyses in U87 cells using RNA sequencing (RNA-Seq) and iTRAQ proteomics methods, respectively. Given that 7 μM PIM showed potent antitumor activity, as described above, this concentration was also used in the following omics studies. After treatment with 7 μM PIM, 1046 differentially expressed genes (DEGs) (log2 | fold change | ≥ 0.4 and P value < 0.05) were detected at the mRNA level; specifically, 667 genes were upregulated and 379 genes were downregulated relative to their levels in control cells (Fig. 6a/b). Subsequently, the functional enrichment of these DEGs was analyzed using GSEA online tools with its hallmark gene set category. The results showed that the downregulated genes were mainly correlated with E2F targets, G2/M checkpoint, MYC targets_V1/2, angiogenesis, mitotic spindle, etc. (Fig. 6c) and that the functions of the upregulated genes were largely associated with cholesterol homeostasis, fatty acid metabolism, androgen response, TNFα signaling via NF-κB, bile acid metabolism, etc. (Fig. 6d). In the iTRAQ proteomics experiment, we detected a total of 7249 proteins, 585 of which were identified as differentially expressed proteins (DEPs), with 272 downregulated and 313 upregulated (Fig. 6e). Subsequent analysis showed that only 66 genes were upregulated and 42 genes were downregulated at both the mRNA and protein levels and that the expression of most of the genes changed significantly at only one level (Fig. 6f). Although the gene expression correlation was low between the transcriptomic and proteomic data, the functions of the DEPs were highly consistent with those of the DEGs. In other words, the functions of downregulated proteins were also related to E2F targets, G2/M checkpoint, and MYC targets, while upregulated proteins were largely involved in cholesterol homeostasis (Fig. S5a/b).
a Volcano plot of mRNA-seq results. Data for all genes were plotted as log2 fold change values versus −log10 adjusted P values. The differentially expressed genes (DEGs) (log2 | fold change | ≥ 0.4 and P value < 0.05) detected after treatment with 7 μM PIM compared with after DMSO control treatment are presented as blue (379 downregulated genes) or red (667 upregulated genes) dots. b Heatmap showing the results of cluster analysis based on DEG expression levels. The gene expression levels were normalized to Z-score values. The bubble chart showing the results of functional enrichment analysis of downregulated (c) or upregulated (d) genes detected by RNA-seq using the GSEA hallmark gene set category. The blue box highlights the top-ranked enriched terms, while the red box highlights the top-ranked enriched cholesterol and fatty acid synthesis-related terms. e Pie chart of proteomic analysis results. A total of 7249 proteins were detected, among which 6764 had no significant expression change, 272 were downregulated, and 313 were upregulated. f Setup diagrams showing the expression correlations between the genes whose expression changed significantly in the proteomic and mRNA-seq results. The line between the two black dots indicates genes that were identified as differentially expressed by both proteomic analysis and mRNA-seq analysis. The blue line indicates downregulation in both analyses, and the red line indicates upregulation in both analyses. g The integrated functional enrichment analysis of downregulated genes identified by proteomic and mRNA-seq profiling using the GSEA PID_Pathways gene set category. h The integrated functional enrichment analysis of downregulated genes identified by proteomic and mRNA-seq profiling using the GSEA KEGG_Pathways gene set category. i Heatmap showing the downregulated genes important in the cell cycle, DNA replication, and base metabolism detected by transcriptomic (left) and proteomic (right) profiling. Left: The color value represents the Z-score of the gene expression level (n = 3); Right: The color value represents the q value of the related protein expression change ratio (n = 2). j Immunoblot analysis of PCNA and NPC2 in U87 cells treated with PIM. k Summary of the major signaling pathways that were downregulated by PIM in the U87 glioblastoma cell line. The RB-E2F, MYC, ATR and AuroraA/B signaling pathways were significantly suppressed.
To further explore the effect of PIM on intracellular processes in cancer cells, we performed integrated analysis of the transcriptomic and proteomic data. Integrated analysis of the downregulated genes was performed using the GSEA KEGG gene set category. The results revealed that PIM markedly reduced the expression of many genes important in DNA replication (MCM2-7, RFCs, POLA2, POLD1, PRIM2, etc.), cell cycle regulation (CCNA2, CCNB1/2, CDC20, CDK2/6, E2F2, etc.), and base metabolism (TYMS, RRM1/2, UCK2, etc.) (Fig. 6g/i). Most of these genes were downstream targets of E2F and MYC transcription factors and were categorized into the E2F targets and MYC targets_V1/2 groups in the hallmark gene set analysis (Fig. S5e/f). It has been clearly revealed that E2F and MYC play vital roles in cell proliferation, especially in controlling G1/S-phase transition [25,26,27]. PIM-induced impairment of E2F and MYC function can cause replication fork collapse and insufficient base supply for DNA replication, which ultimately results in G1 phase arrest and apoptosis, as shown above. As indicated by the Western blot results, PCNA, a downstream effector of E2F important in DNA replication, was obviously downregulated, confirming the impairment of E2F function (Fig. 6j). However, it was still difficult to find a signaling pathway network that PIM influenced. Hence, we conducted further integrated PID pathway enrichment analysis of downregulated genes. The ATR, AuroraA/B, and E2F signaling pathways were strongly inhibited (Fig. 6h). ATR (ataxia telangiectasia and Rad3-related protein) is an important regulator of the cell cycle and DNA damage repair (DDR) response [28] (Fig. 6i). ATR can phosphorylate cell cycle checkpoint kinase 1 (Chk1), which then inhibits CDC25A. CDK2, a key cyclin-dependent kinase, can be activated by CDC25A and along with CycA to phosphorylate RB, which finally releases E2F to regulate gene expression. In addition, RB can also be phosphorylated by the PDGFR/MAPK axis downstream effectors CDK4/6. Omics studies have revealed that components in this ATR/RB axis (CDK2, PDGFR, CycA, and CDK6) are greatly downregulated, which might lead to suppression of E2F activity [29]. Moreover, downregulation of CDK2 might lower the activity of the transcription factor FOXM1, which could reduce the expression of many G2/M checkpoint genes [30], such as CCNB1/2 and BIRC5. The chromosomal passenger (CPC) complex has a very important role in metaphase arrangement, sister chromatid separation, cytokinesis, etc. [31]. Our omics analysis demonstrated that all four of its components—AURKB (AuroraB), CDCA8, CPC, and BIRC5—were downregulated, indicating that mitosis might be severely inhibited. In addition, the expression of some regulators of the CPC complex, including PLK1, MKLP2, and CDK1, was suppressed, which further reduced CPC complex activity. The TPX2/AuroraA axis, another key regulator of cell mitosis, was also suppressed [32]. The S phase arrest observed above might result from inhibition of FOXM1, the CPC complex, and AuroraA/B [31]. In the PID pathway analysis, the MYC binding partners TIP48/49 and ACTL6A were found to be repressed, which may reduce the activity of MYC [33, 34].
Interestingly, our integrated omics studies revealed that the upregulated genes, including ALDOC, DHCR7, FADS2, FASN, FDFT1, HMGCS1, MVD, and SCD, were mainly associated with cholesterol homeostasis (Fig. S5c). All upregulated genes correlated with cholesterol homeostasis in the proteomic data were upregulated at the mRNA level, indicating that the increase in cholesterol metabolism may result from regulation of upstream molecules but not from a direct influence on cholesterol synthesis. Cholesterol esters are transported to lysosomes through endocytosis [35]. In lysosomes, acid lipases degrade cholesteryl esters to release cholesterol, and then via the Niemann-Pick C1/2 protein (NPC1/2), cholesterol is transported to the ER [36]. In proteomic studies, NPC2 expression was found to be downregulated by PIM at the protein level, which was also confirmed by immunoblotting. Downregulation of NPC2 might impede the transport of cholesterol to the ER from lysosomes and decrease the ER cholesterol level. This cholesterol depletion in the ER membrane can dissociate SREBP cleavage activation protein (SCAP) from the Insig1 complex, which inappropriately promotes the translocation of SREBPs to the Golgi apparatus [37]. Golgi-resident S1P/S2P proteases then cleaved SREBPs and allowed them to enter the nucleus to regulate a series of cholesterol metabolism genes [37] (Fig. S5d). Moreover, metabolism and signaling associated with cholesterol derivatives, including bile acids, androgen and estrogen, was also increased, possibly because of the elevated cholesterol. In addition, mTORC1 signaling was significantly upregulated (Fig. S5g). Very interestingly, most of the upregulated genes in mTORC1 signaling were closely related to cholesterol or fatty acid synthesis. Currently, unambiguous evidence has confirmed the critical role of mTORC1 signaling in regulating cholesterol homeostasis. High mTORC1 activity can suppress cholesterol trafficking to the ER, elevating the expression of genes involved in cholesterol synthesis [38, 39]. Therefore, the enhancement of cholesterol metabolism might partially result from the upregulation of mTORC1 signaling activity. In addition, it should be noted that an increase in cholesterol or fatty acid synthesis may be detrimental to PIM’s antitumor activity and contribute to its toxic side effects, and this possibility should be given special attention in future clinical applications.
Pimavanserin tartrate inhibits the growth of subcutaneous and orthotopic glioblastoma xenografts in vivo
To assess the antitumor efficacy of PIM in vivo, we first established a GBM xenograft model by implanting U87 cells subcutaneously into nude mice. The mice were randomized into two groups. The experimental group was treated with 10 mg/kg PIM daily by gavage for 3 weeks, while the control group was treated with an equal amount of saline solution. The results showed that PIM substantially suppressed tumor growth with prolonged treatment time (Fig. 7a/b). After 3 weeks of treatment, the tumor size was reduced by ~70% compared with that in the control group (Fig. 7b). There was no significant difference in mouse body weights between the control and PIM-treated groups (Fig. 7c), implying that PIM may have low toxicity. Subsequently, hematoxylin and eosin (H&E) staining of the harvested tumor tissues was performed. Compared with the control treatment, PIM-induced massive tumor tissue necrosis, showing high toxicity to tumor cells in vivo. Consistent with the flow cytometry results in cellula, immunohistochemical staining revealed that PIM can enhance caspase-3 activity and reduce cyclinB2 expression, thereby validating the apoptosis promotion and cell cycle inhibition effects in vivo (Figs. 7d/e and S4d). Moreover, after treatment with PIM, the number of nuclear NFAT-positive cells was decreased compared with that in the control group, indicating that PIM effectively inhibited NFAT nuclear translocation in vivo (Fig. 7e). Immunoblot analysis of NFATc3 in the cytosolic fraction extracted from tumor tissues demonstrated that the NFATc3 level was significantly higher in the PIM-treated group than the control group (Fig. S4e), which resulted from PIM’s inhibitory effect on NFAT translocation from the cytosol to the nucleus. It is known that the blood–brain barrier (BBB) is a key factor that determines the therapeutic efficacy of drugs for brain diseases, and many effective antitumor drugs fail for GBM treatment because of their inability to cross the BBB. Therefore, to further validate the therapeutic potential of PIM for in situ GBM tumors, we established an orthotopic xenograft GBM model by implanting luciferase-expressing U87 cells (U87-luciferase cells) in the intracalvarium. As described above, PIM was also administered via gavage at a dose of 10 mg/kg PIM daily for 3 weeks, and tumor growth was monitored weekly by bioluminescence imaging. As shown in Fig. 7f/g, after 2 weeks of treatment, the area of luminescence was significantly decreased, indicating that PIM effectively crossed the BBB to suppress orthotopic GBM tumor growth. Therefore, PIM exhibited promising therapeutic potential for GBM tumors and deserves further study.
a Representative images of subcutaneous glioblastoma xenografts obtained weekly from vehicle-treated or PIM-treated nude mice. Nude mice were subcutaneously implanted with U87 glioblastoma cells and treated with 10 mg/kg PIM or an equal amount of saline solution (vehicle control) daily for 3 weeks via gavage. b The volumes of the tumor tissues obtained from vehicle-treated and PIM-treated nude mice at each weekly measurement (n = 5). ***P < 0.001. Note that PIM significantly suppressed glioblastoma tumor growth. c The body weights of vehicle-treated and PIM-treated nude mice at each weekly measurement (n = 5). d Representative H&E staining results of subcutaneous xenograft tumor tissues excised from vehicle- and PIM-treated mice. e Representative images of immunohistochemical staining of DNA fragments (TUNEL), activated caspase-3 (red fluorescence) and NFATc3 in subcutaneous GBM xenograft tissues excised from vehicle- and PIM-treated mice. For activated caspase-3 staining, the nuclei of tumor cells were stained with DAPI. For NFATc3 staining, the red arrow indicates nuclear NFAT-positive cells, and the blue arrow indicates cytosolic NFAT-positive cells. f Representative bioluminescence images of orthotopic glioblastoma xenografts at each weekly assessment. U87-luc glioblastoma cells were implanted in the intracalvarium of nude mice, and the mice were then treated with 10 mg/kg PIM or an equal amount of saline solution (vehicle control) daily for 3 weeks via gavage. g The tumor size in vehicle-treated and PIM-treated nude mice was reflected by the bioluminescence intensity in orthotopic xenograft tumors measured weekly and plotted relative to the bioluminescence intensity at baseline (week 0).
Discussion
GBM is one of the most lethal types of human cancers, and 95% of patients die within 2 years of diagnosis. Although immunotherapy and targeted therapy, a popular research topic for curing tumors, have recently been used for GBM treatment [40], the overall survival improvement is still limited, with an increase of only 1 year. Therefore, therapy for GBM is still challenging. Since the downstream genes of NFATs, such as VEGFR and MMPs, are mostly associated with cancer cell invasion and migration, the NFAT signaling pathway is a promising therapeutic target for highly aggressive tumors, including GBM. In this study, we discovered that PIM was a potent NFAT signaling pathway inhibitor and exhibited effective antitumor activity against GBM both in vitro and in vivo. PIM has been used in the clinic, and from a safety perspective, it holds great promise for entering clinical trials and benefiting GBM patients. In addition, it is worth noting that the other seven potent NFAT signaling pathway inhibitors also showed high inhibitory activity against glioblastoma cell proliferation (Table S2). In particular, the antitumor effects of butoconazole, domperidone, sertaconazole, and mebhydrolin were reported for the first time. Many studies have revealed the antitumor activity of pimozide, raloxifene, and nefazodone [41, 42], and our data demonstrated that inhibition of the NFAT signaling pathway may constitute another novel mechanism underlying their tumor-suppressive effect.
Pimavanserin tartrate is an effective 5-HT2A receptor inverse agonist. In addition to affinity for the 5-HT2A receptor, PIM has also been reported to show high binding affinity for other receptors, including 5-HT2B/c, sigma1, adrenergic, dopamine, and histamine receptors. In this study, we identified that PIM significantly suppressed SOCE by inhibiting the CRAC channel (Fig. 4e), which then decreased the activity of its downstream signaling target NFAT. Therefore, PIM is a novel potent Ca2+-calcium-NFAT pathway inhibitor. The initiation of SOCE is a complex process. Although many CRAC channel inhibitors have been reported, few studies have elucidated how they might affect each critical node in CRAC channel activation. After a systematic study using a competitive binding assay and a novel optogenetic tool (Opto-CRAC channel), it was found that PIM effectively prevented STIM1 oligomerization, an essential event in CRAC channel activation. In addition, the process that initiates SOCE has been well elucidated, but the details of STIM1 puncta and ER-PM junction formation as well as the mechanism by which STIM1 translocates to ER-PM junctions are still not well explained. To date, several molecules have been reported to participate in modulating the processes controlled by these signaling nodes, such as EB1 [43] and STIMATE [44]. Whether other molecules are involved in these processes is still unclear. Considering that PIM inhibited TG-induced NFAT translocation and SOCE in a unique temperature-dependent manner that has not been reported before, PIM is anticipated to be a useful pharmaceutical tool to explore the regulatory mechanism of SOCE. It is well established that ER Ca2+ is important for many cellular processes. However, PIM can lower ER Ca2+ levels, which may be harmful to normal cells and should be given more attention in future clinical applications.
Pimavanserin tartrate is an antipsychotic used for the treatment of Parkinson’s disease-associated psychiatric symptoms. In fact, many antipsychotics, such as fluoxetine, chlorpromazine, olanzapine, risperidone, and quetiapine [45], have shown therapeutic efficacy in GBM. Moreover, epidemiologic studies have shown that the incidence of tumors in schizophrenia patients is significantly lower than that in the normal population [46]. These interesting results may partially be attributed to the ease with which antipsychotics can penetrate the BBB, which is important for drugs used for brain diseases. In addition, the targets of these drugs are GPCR receptors, and in the presence of defined amounts of stimuli in vivo, such as 5-HT and dopamine, the downstream signaling pathway of GPCRs is suppressed by these drugs, which may contribute to their antitumor activity. Inspired by the results of our study, especially the finding that most of the drugs we identified as potent NFAT signaling pathway inhibitors also targeted GPCRs, we speculated that there may be common pharmacophores in the structures of GPCR ligands and SOC channel (CRAC channel) inhibitors, such as hydrophobic groups and hydrogen receptors. Therefore, the SOC channel may be a novel target for some antipsychotics, a hypothesis that can be tested in future research.
To date, there are few studies revealing PIM’s antitumor activity. Only Sanjay K. Srivastava and Jinchun Qi recently reported that PIM showed antitumor effects in pancreatic [47] and bladder cancers, respectively. Inhibition of the Akt/Gli1 signaling axis and Wnt/β-catenin and Akt/mTOR pathways was highlighted to be responsible for PIM’s suppressive effect on tumor cell growth. By contrast, we performed comparative transcriptomic and proteomic analyses to globally evaluate the effect of PIM in glioblastoma U87 cells. It was found that PIM significantly downregulated the RB/E2F, MYC, ATR, FOXM1, and AuroraA/B signaling pathways (Fig. 6k), which impaired DNA replication, base metabolism, and mitosis and ultimately resulted in G1/S phase arrest and apoptosis. In addition to the changes described above, in PID pathway enrichment analysis, the calcineurin-NFAT pathway was significantly downregulated, an effect that may be attributed to the potent inhibitory activity of PIM against the NFAT signaling pathway. In addition, some favorable reciprocal changes, including downregulation of tumor-promoting genes (VCAN, SDC4, TNC, TAGLN, FBN1, LAMC1, etc.) and upregulation of tumor-suppressive genes (DAB2, FBLN5, TFPI2, IFNGR1, etc.), were observed, and all contributed to suppression of tumor cell invasion or migration. Certainly, some tumor-promoting genes, such as LUM, SAT1, SCG2, TIMP1, and IL1B, were upregulated, making tumor cells resistant to PIM treatment. In addition, it should be noted that PER2 had the most significantly decreased expression in the proteomic study. PER2 is also called Period Circadian Protein 2, and some studies have shown that it might be an independent risk factor for the occurrence of dyskinesia in patients with Parkinson’s disease (PD) [48], a finding that might reveal a new mechanism for PIM in PD treatment.
Interestingly, the upregulated genes after treatment with PIM were mainly associated with cholesterol homeostasis. Indeed, antipsychotic drugs are frequently reported to upregulate the expression of genes involved in cholesterol homeostasis [49]. However, the mechanism underlying this upregulation of cholesterol synthesis is not well known. Many studies have pointed toward their amphiphilic weak base features, which can disrupt the function of lysosomes. Our omics studies revealed that downregulation of the cholesterol transporter NPC2 and upregulation of mTORC1 activity may impede cholesterol trafficking from lysosomes to the ER, which then activates SREBPs and increases the expression of genes involved in cholesterol synthesis. Given that myelin plays an essential role in maintaining the normal function of the central nervous system and that cholesterol is its major component, an increase in cholesterol synthesis may contribute to the therapeutic efficacy of antipsychotics [50], including PIM. Conversely, increased cholesterol synthesis may also cause metabolic side effects, including body weight gain, hyperlipidemia, and cardiovascular diseases, which are commonly observed in patients with psychiatric disorders. In the case of PIM, upregulation of cholesterol synthesis may reduce its antitumor activity and result in side effects. Therefore, combination treatment with mTORC1 or cholesterol synthesis inhibitors may potentiate the antitumor activity of PIM.
Data availability
Additional experimental data are provided as Supplementary Information and are available from the corresponding author upon request. Raw RNA-Seq data are available in NCBI’s Sequence Read Archive (SRA) database under accession number PRJNA689001. The mass spectrometry proteomics data have been deposited to the ProteomeXchange Consortium (http://proteomecentral.proteomexchange.org) via the iProX partner repository with the dataset identifier PXD023207 [51].
References
Stylli SS. Novel treatment strategies for glioblastoma. Cancers. 2020;12:2883.
Neftel C, Laffy J, Filbin MG, Hara T, Shore ME, Rahme GJ, et al. An integrative model of cellular states, plasticity, and genetics for glioblastoma. Cell. 2019;178:835–49. e21.
Recurrent MET fusion genes represent a drug target in pediatric glioblastoma. Nat Med. 2016;22:1314–20.
Hu B, Wang Q, Wang YA, Hua S, Sauvé C-EG, Ong D, et al. Epigenetic activation of WNT5A drives glioblastoma stem cell differentiation and invasive growth. Cell. 2016;167:1281–95. e18.
Qin JJ, Nag S, Wang W, Zhou J, Zhang WD, Wang H, et al. NFAT as cancer target: mission possible? Biochim Biophys Acta. 2014;1846:297–311.
Muppidi JR. A role for NFAT signaling in ABC-DLBCL. Blood. 2020;135:81.
Kaunisto A, Henry WS, Montaser-Kouhsari L, Jaminet SC, Oh EY, Zhao L, et al. NFAT1 promotes intratumoral neutrophil infiltration by regulating IL8 expression in breast cancer. Mol Oncol. 2015;9:1140–54.
Tie X, Han S, Meng L, Wang Y, Wu A. NFAT1 is highly expressed in, and regulates the invasion of, glioblastoma multiforme cells. PLoS One. 2013;8:e66008.
Lee JV, Berry CT, Kim K, Sen P, Kim T, Carrer A, et al. Acetyl-CoA promotes glioblastoma cell adhesion and migration through Ca2+-NFAT signaling. Genes Dev. 2018;32:497–511.
Chigurupati S, Venkataraman R, Barrera D, Naganathan A, Madan M, Paul L, et al. Receptor channel TRPC6 is a key mediator of Notch-driven glioblastoma growth and invasiveness. Cancer Res. 2010;70:418–27.
Robbs BK, Cruz AL, Werneck MB, Mognol GP, Viola JP. Dual roles for NFAT transcription factor genes as oncogenes and tumor suppressors. Mol Cell Biol. 2008;28:7168–81.
Pushpakom S, Iorio F, Eyers PA, Escott KJ, Hopper S, Wells A, et al. Drug repurposing: progress, challenges and recommendations. Nat Rev Drug Discov. 2019;18:41–58.
Liu Z, Li H, He L, Xiang Y, Tian C, Li C, et al. Discovery of small-molecule inhibitors of the HSP90-calcineurin-NFAT pathway against glioblastoma. Cell Chem Biol. 2019;26:352–65. e7.
Abbas A, Roth BL. Pimavanserin tartrate: a 5-HT2A inverse agonist with potential for treating various neuropsychiatric disorders. Expert Opin Pharmacother. 2008;9:3251–9.
Yang Y, Liu N, He Y, Liu Y, Ge L, Zou L, et al. Improved calcium sensor GCaMP-X overcomes the calcium channel perturbations induced by the calmodulin in GCaMP. Nat Commun. 2018;9:1504.
Nguyen NT, Han W, Cao WM, Wang Y, Wen S, Huang Y, et al. Store-operated calcium entry mediated by ORAI and STIM. Compr Physiol. 2018;8:981–1002.
Liou J, Fivaz M, Inoue T, Meyer T. Live-cell imaging reveals sequential oligomerization and local plasma membrane targeting of stromal interaction molecule 1 after Ca2+ store depletion. Proc Natl Acad Sci USA. 2007;104:9301–6.
Zhang SL, Yu Y, Roos J, Kozak JA, Deerinck TJ, Ellisman MH, et al. STIM1 is a Ca2+ sensor that activates CRAC channels and migrates from the Ca2+ store to the plasma membrane. Nature. 2005;437:902–5.
He L, Zhang Y, Ma G, Tan P, Li Z, Zang S, et al. Near-infrared photoactivatable control of Ca2+ signaling and optogenetic immunomodulation. Elife. 2015;4:e10024.
Morgan AJ, Jacob R. Ionomycin enhances Ca2+ influx by stimulating store-regulated cation entry and not by a direct action at the plasma membrane. Biochem J. 1994;300:665–72.
Kang F, Zhou M, Huang X, Fan J, Wei L, Boulanger J, et al. E-syt1 re-arranges STIM1 clusters to stabilize ring-shaped ER-PM contact sites and accelerate Ca2+ store replenishment. Sci Rep. 2019;9:3975.
Woo JS, Sun Z, Srikanth S, Gwack Y. The short isoform of extended synaptotagmin-2 controls Ca2+ dynamics in T cells via interaction with STIM1. Sci Rep. 2020;10:14433.
Giordano F, Saheki Y, Idevall-Hagren O, Colombo SF, Pirruccello M, Milosevic I, et al. PI4,5P2-dependent and Ca2+-regulated ER-PM interactions mediated by the extended synaptotagmins. Cell. 2013;153:1494–509.
Schmidt JH, Pietkiewicz S, Naumann M, Lavrik IN. Quantification of CD95-induced apoptosis and NF-κB activation at the single cell level. J Immunol Methods. 2015;423:12–7.
Attwooll C, Lazzerini Denchi E, Helin K. The E2F family: specific functions and overlapping interests. EMBO J. 2004;23:4709–16.
Chen H, Liu H, Qing G. Targeting oncogenic Myc as a strategy for cancer treatment. Signal Transduct Target Ther. 2018;3:5.
Chen HZ, Tsai SY, Leone G. Emerging roles of E2Fs in cancer: an exit from cell cycle control. Nat Rev Cancer. 2009;9:785–97.
Mei L, Zhang J, He K, Zhang J. Ataxia telangiectasia and Rad3-related inhibitors and cancer therapy: where we stand. J Hematol Oncol. 2019;12:43.
Harbour JW, Luo RX, Santi AD, Postigo AA, Dean DC. Cdk phosphorylation triggers sequential intramolecular interactions that progressively block Rb functions as cells move through G1. Cell. 1999;98:859–69.
Laoukili J, Alvarez M, Meijer LA, Stahl M, Mohammed S, Kleij L, et al. Activation of FoxM1 during G2 requires cyclin A/Cdk-dependent relief of autorepression by the FoxM1 N-terminal domain. Mol Cell Biol. 2008;28:3076–87.
Carmena M, Wheelock M, Funabiki H, Earnshaw WC. The chromosomal passenger complex (CPC): from easy rider to the godfather of mitosis. Nat Rev Mol Cell Biol. 2012;13:789–803.
Bayliss R, Sardon T, Vernos I, Conti E. Structural basis of aurora-A activation by TPX2 at the mitotic spindle. Mol Cell. 2003;12:851–62.
Wood MA, McMahon SB, Cole MD. An ATPase/helicase complex is an essential cofactor for oncogenic transformation by c-Myc. Mol Cell. 2000;5:321–30.
Park J, Wood MA, Cole MD. BAF53 forms distinct nuclear complexes and functions as a critical c-Myc-interacting nuclear cofactor for oncogenic transformation. Mol Cell Biol. 2002;22:1307–16.
Luo J, Yang H, Song BL. Mechanisms and regulation of cholesterol homeostasis. Nat Rev Mol Cell Biol. 2020;21:225–45.
Rosenbaum AI, Maxfield FR. Niemann-Pick type C disease: molecular mechanisms and potential therapeutic approaches. J Neurochem. 2011;116:789–95.
Horton JD, Goldstein JL, Brown MS. SREBPs: activators of the complete program of cholesterol and fatty acid synthesis in the liver. J Clin Invest. 2002;109:1125–31.
Laplante M, Sabatini DM. An emerging role of mTOR in lipid biosynthesis. Curr Biol. 2009;19:R1046–52.
Eid W, Dauner K, Courtney KC, Gagnon A, Parks RJ, Sorisky A, et al. mTORC1 activates SREBP-2 by suppressing cholesterol trafficking to lysosomes in mammalian cells. Proc Natl Acad Sci USA. 2017;114:7999–8004.
Lim M, Xia Y, Bettegowda C, Weller M. Current state of immunotherapy for glioblastoma. Nat Rev Clin Oncol. 2018;15:422–42.
Varalda M, Antona A, Bettio V, Roy K, Vachamaram A, Yellenki V, et al. Psychotropic drugs show anticancer activity by disrupting mitochondrial and lysosomal function. Front Oncol. 2020;10:562196.
Pappas SG, Jordan VC. Raloxifene for the treatment and prevention of breast cancer? Expert Rev Anticancer Ther. 2001;1:334–40.
Chang CL, Chen YJ, Quintanilla CG, Hsieh TS, Liou J. EB1 binding restricts STIM1 translocation to ER-PM junctions and regulates store-operated Ca2+ entry. J Cell Biol. 2018;217:2047–58.
Jing J, He L, Sun A, Quintana A, Ding Y, Ma G, et al. Proteomic mapping of ER–PM junctions identifies STIMATE as a regulator of Ca2+ influx. Nat Cell Biol. 2015;17:1339–47.
Wiklund ED, Catts VS, Catts SV, Ng TF, Whitaker NJ, Brown AJ, et al. Cytotoxic effects of antipsychotic drugs implicate cholesterol homeostasis as a novel chemotherapeutic target. Int J Cancer. 2010;126:28–40.
Barak Y, Achiron A, Mandel M, Mirecki I, Aizenberg D. Reduced cancer incidence among patients with schizophrenia. Cancer. 2005;104:2817–21.
Ramachandran S, Srivastava SK. Repurposing pimavanserin, an anti-Parkinson drug for pancreatic cancer therapy. Mol Ther Oncolytics. 2020;19:19–32.
Breen DP, Vuono R, Nawarathna U, Fisher K, Shneerson JM, Reddy AB, et al. Sleep and circadian rhythm regulation in early Parkinson disease. JAMA Neurol. 2014;71:589–95.
Canfrán-Duque A, Casado ME, Pastor O, Sánchez-Wandelmer J, de la Peña G, Lerma M, et al. Atypical antipsychotics alter cholesterol and fatty acid metabolism in vitro. J Lipid Res. 2013;54:310–24.
Fernø J, Skrede S, Vik-Mo AO, Håvik B, Steen VM. Drug-induced activation of SREBP-controlled lipogenic gene expression in CNS-related cell lines: marked differences between various antipsychotic drugs. BMC Neurosci. 2006;7:69.
Ma J, Chen T, Wu S, Yang C, Bai M, Shu K, et al. iProX: an integrated proteome resource. Nucleic Acids Res. 2019;47:D1211–D7.
Acknowledgements
This work was supported by the National Natural Science Foundation of China (21927811, 91753111, 21907061, 62006144), the Key Research and Development Program of Shandong Province (2018YFJH0502), the Postdoctoral Research Foundation of China (2017M622225), Postdoctoral Innovation Foundation of Shandong Province (201703009) and Jinan Innovative Team Project (2019GXRC039).
Author information
Authors and Affiliations
Contributions
ZZL, XNL, PL and BT designed the research. ZZL, XNL, HLL, YPJ, RCF, LZJ, and YQZ conducted experiments and acquired the data. ZZL, XNL, HLL, YPJ, RCF, QKZ, LZJ, XQG, YQW, and MQY performed data analysis. ZZL, XNL, HLL, PL, and BT wrote and revised the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Supplementary information
Rights and permissions
About this article
Cite this article
Liu, Zz., Liu, Xn., Fan, Rc. et al. Identification of pimavanserin tartrate as a potent Ca2+-calcineurin-NFAT pathway inhibitor for glioblastoma therapy. Acta Pharmacol Sin 42, 1860–1874 (2021). https://doi.org/10.1038/s41401-021-00724-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41401-021-00724-2