Abstract
Diffuse large B-cell lymphoma (DLBCL) is the most widespread type of non-Hodgkin lymphoma (NHL). As the most aggressive form of the DLBCL, the activated B-cell-like (ABC) subtype is often resistant to standard chemotherapies. Bruton’s tyrosine kinase (BTK) inhibitor ibrutinib provides a potential therapeutic approach for the DLBCL but fails to improve the outcome in the phase III trial. In the current study, we investigated the molecular mechanisms underlying ibrutinib resistance and explored new combination therapy with ibrutinib. We generated an ibrutinib-resistant ABC-DLBCL cell line (OCI-ly10-IR) through continuous exposure to ibrutinib. Transcriptome analysis of the parental and ibrutinib-resistant cell lines revealed that the ibrutinib-resistant cells had significantly lower expression of the unfolded protein response (UPR) marker genes. Overexpression of one UPR branch-XBP1s greatly potentiated ibrutinib-induced apoptosis in both sensitive and resistant cells. The UPR inhibitor tauroursodeoxycholic acid (TUDCA) partially reduced the apoptotic rate induced by the ibrutinib in sensitive cells. The UPR activator 2-deoxy-D-glucose (2-DG) in combination with the ibrutinib triggered even greater cell growth inhibition, apoptosis, and stronger calcium (Ca2+) flux inhibition than either of the agents alone. A combination treatment of ibrutinib (15 mg·kg−1·d−1, po.) and 2-DG (500 mg/kg, po, b.i.d.) synergistically retarded tumor growth in NOD/SCID mice bearing OCI-ly10-IR xenograft. In addition, ibrutinib induced the UPR in the sensitive cell lines but not in the resistant cell lines of the DLBCL. There was also a combined synergistic effect in the primary resistant DLBCL cell lines. Overall, our results suggest that targeting the UPR could be a potential combination strategy to overcome ibrutinib resistance in the DLBCL.
Similar content being viewed by others
Introduction
Diffuse large B-cell lymphoma (DLBCL) is the most common type of non-Hodgkin lymphoma (NHL), accounting for ~30%–40% of NHL worldwide [1]. Although DLBCL is classified into activated B-cell–like (ABC), germinal-center B-cell–like (GCB), and primary mediastinal B-cell lymphoma (PMBL), an additional 15%–30% of cases remain unclassified. These subgroups have distinct gene expression profiles, oncogenic aberrations, and clinical outcomes [2]. ABC-DLBCL is characterized by chronically active B-cell receptor (BCR) signaling that can be modulated by Bruton’s tyrosine kinase (BTK) activity. A phase I/II clinical trial of the single-agent ibrutinib, the first-in-class orally available covalent BTK inhibitor for relapsed and refractory DLBCL patients, led to a complete or partial response in 37% of ABC-DLBCL patients [3]. Despite these encouraging results, combination therapy of ibrutinib with rituximab plus cyclophosphamide, doxorubicin, vincristine, and prednisone (R-CHOP) used in a phase III clinical trial failed to meet the criteria by the primary endpoint to be considered a frontline treatment for patients with newly diagnosed DLBCL [4].
Ibrutinib has demonstrated clinical efficacy in several B-cell malignancies, including chronic lymphocytic leukemia (CLL), mantle cell lymphoma (MCL), and Waldenström’s macroglobulinemia (WM) [5, 6]. The overall response rate of ibrutinib in older patients was 90.5%, 68%, and 71% for WM, MCL, and CLL/small lymphocytic lymphoma (SLL), respectively [7]. Since ibrutinib was approved as a first-line treatment for patients with CLL in 2016, most patients have shown primary resistance or developed secondary resistance. Resistance mechanisms against ibrutinib were also identified in DLBCL. The different compositions of the CD79A/B and MYD88 mutation types led to different clinical responses to ibrutinib therapy [3]. Ibrutinib resistance was induced by CD79B overexpression via the activation of the AKT and/or MAPK pathway(s) [8] or downregulation of FOXO3a and PTEN levels and activation of AKT [9]. Higher BCL2 gene expression [10] or upregulated PIM1 expression [11] was also related to ibrutinib resistance in ABC DLBCL. Unfortunately, ibrutinib is not currently approved for the treatment of DLBCL. Moreover, the intrinsic mechanisms of ibrutinib resistance in the clinical treatment of DLBCL remain largely unknown.
To further explore ibrutinib resistance mechanisms in DLBCL, an ibrutinib-resistant ABC-DLBCL cell line was generated by prolonged exposure to ibrutinib. We compared the cDNA microarray profile of parental sensitive and derived resistant cells to identify the key gene signature relevant to ibrutinib resistance in DLBCL. These results provide, for the first time, a rationale for combining BTK inhibition and UPR activation in the treatment of DLBCL. Targeting the UPR-related pathway might be a novel strategy to kill malignant B cells and overcome ibrutinib resistance in DLBCL.
Materials and methods
Reagents
Ibrutinib (HY-10997), acalabrutinib (ACP-196, HY-17600), LBH589 (HY-10224), SAHA (HY-10221), AZD7762 (HY-10992), 2-deoxy-D-glucose (2-DG, HY-13966), tunicamycin (TM, HY-A0098), baricitinib (HY-15315), ruxolitinib (HY-50856), and MLN120B (HY-15473) were purchased from MedChemExpress (MCE.) The following antibodies were used: anti-CHOP (NB600-1335, Novus), anti-XBP1s (A1731, ABclonal), anti-BCL2 (3498s, CST), anti-BCLXL (2764, CST), anti-MCL1 (39224S, CST), anti-PARP (9542s, CST), anti-GAPDH (2118s, CST), anti-β-actin (AM1021b, ABGENT), goat anti-mouse secondary antibodies or goat anti-rabbit secondary antibodies (926–32211, Odyssey), goat anti-human IgM (109-006-129, Fc5µ fragment specific, Jackson ImmunoResearch Laboratories), and Fluo-4 NW Calcium Assay Kits (36206, Invitrogen).
Cell lines and culture conditions
OCI-ly10, an ABC-DLBCL cell line, was obtained from the American Type Culture Collection (ATCC) (Manassas, VA, USA) and maintained at 37 °C in Iscove’s modified Dulbecco’s medium supplemented with 20% fetal bovine serum (FBS) (GIBCO) and 100 U/mL penicillin and 100 µg/mL streptomycin. The SU-DHL-2, Toledo, SU-DHL-8, and Pfeiffer DLBCL cell lines were obtained from ATCC and cultured in RPMI-1640 medium supplemented with 10% FBS, 100 U/mL penicillin, and 100 µg/mL streptomycin (Invitrogen, USA). The TMD8 and HBL1 DLBCL cell lines were generously provided by Dr Lynn Wang (University of Chicago). In addition, all cell lines were frequently tested and determined to be mycoplasma-free (Shanghai Life iLab Biotechnology Co., Ltd.). All human cell lines were authenticated using STR profiling (Shanghai Genesky Biotechnology Co., Ltd.).
Induction of ibrutinib-resistant ABC-DLBCL cells
Ibrutinib-resistant (IR) cells were derived from an original parental cell line (OCI-ly10) by continuous exposure to gradually increased ibrutinib concentrations (1–400 nM). This development period was carried out for ~12 months. Vials of the treated cells were frozen after each increase in drug concentration. Before subsequent experiments, the cells were maintained in drug-free growth medium for at least 1 week. The cell line identities were confirmed using a short tandem repeat DNA analysis.
Cell viability assay
Briefly, 90 μL of cells (2 × 104 cells/well) were seeded into 96-well plates. After 2 h of incubation, the cells were treated with 10 μL of 0.01% DMSO or varying concentrations of the test compounds for 72 h. Cell viability was measured using the CellTiter 96® AQueous nonradioactive cell proliferation assay (MTS; Promega, Madison, WI). The combined solution of MTS/PMS (20 μL) was pipetted into each well of the 96-well plates and incubated at 37 °C for 2–4 h. Optical density was determined at 490 nm (background values measured at 690 nm were subtracted) using a SpectraMax 340 microplate reader (Molecular Devices, Sunnyvale, CA, USA). The growth inhibition ratio was calculated as follows:
Growth inhibition ratio = (ODcontrol − ODsample)/ODcontrol.
The IC50 values were derived from a nonlinear regression model (curve fit) based on a sigmoidal dose-response curve (variable slope) and computed using GraphPad software. The results are presented as the mean ± SEM from at least three separate assays performed in triplicate.
DNA sequencing and analysis
Qualified genomic DNA from the OCI-ly10 and OCI-ly10-IR cells was prepared for whole-exome sequencing (WES) by Novogene Co., LTD. Briefly, 0.6 μg of genomic DNA per sample was used as input material for the DNA sample preparation. Sequencing libraries were generated using the Agilent SureSelect Human All Exon kit (Agilent Technologies, CA, USA) following the manufacturer’s recommendations. They were sequenced on an Illumina HiSeq platform, and 150 bp paired-end reads were generated. SAMtools [12] mpileup and BCFtools were used for variant calling and identifying single-nucleotide polymorphism (SNPs) and insertion–deletion (Indels). Somatic single-nucleotide variant (SNVs) were detected with muTect [13] and somatic indels with Strelka [14]. The control-FREEC tool [15] was used to detect somatic copy-number variation (CNVs).
Microarray hybridization and analysis
Total RNA from the OCI-ly10 and OCI-ly10-IR cells was extracted using TRIzol reagent (TaKaRa, #9109) according to the manufacturer’s instructions. The RNA quality was assessed on an Agilent 2100 Bioanalyzer. To ensure biological confidence, three independent experiments were performed. Amplified RNA was prepared with an Affymetrix GeneChip 3′ IVT expression kit according to the kit protocol. Amplified RNA was purified, fragmented, and hybridized with a microprobe. After hybridization, the GeneChip PrimeView Human array (Affymetrix, 901838) was washed, dyed, and finally scanned to obtain pictures and original data. Gene sets that differed by age groups were considered significant at a false discovery rate (FDR) < 0.05. Pathway analysis of the selected genes was performed using a fold-change > 1.5 (or <−1.5). Pathway relationships were determined using Ingenuity Pathway Analysis (IPA) software (Ingenuity Systems). Upstream regulator analysis based on the IPA results was used to identify any molecule that affects the expression or function of the measured downstream target genes. Gene set enrichment analysis (GSEA) [16] was performed using the entire list of genes.
RNA isolation and reverse transcription reaction
Total cellular RNA was isolated from transfected cells using TRIzol reagent (Invitrogen) according to the manufacturer’s instructions. RNA was reverse transcribed by PrimeScript™ RT master mix (TaKaRa, #RR037A) according to the manufacturer’s protocol. The RNA concentration and integrity were tested using a NanoDrop 1000 UV/VIS spectrophotometer (Thermo Scientific, Wilmington, DE, USA). Total RNA (2.5 μg) was reverse transcribed using PrimeScript Reverse Transcriptase (TaKaRa, Dalian, China).
Real-time quantitative PCR
Real-time quantitative PCR was performed on an Mx3005P system (Agilent, CA, USA) following the manufacturer’s instructions. Amplification was initiated at 95 °C for 5 min followed by 40 cycles of denaturation at 95 °C for 30 s, annealing at 60 °C for 30 s, and extension at 72 °C for 30 s using AceQ® Universal SYBR quantitative PCR master mix (Vazyme, #Q511-02). Samples were analyzed in duplicate and normalized to actin expression using the 2−ΔΔCt method [17]. Primer sequences are listed in Supplementary Table 1.
Western blotting
Briefly, cells were washed with ice-cold PBS and lysed with RIPA buffer for 30 min. Protein-matched lysates were boiled in Laemmli sample buffer (Bio-Rad, #161-0747). Equal amounts of protein were separated using SDS-PAGE and subsequently transferred onto a nitrocellulose membrane. The membranes were blocked with TBST buffer with 5% nonfat dry milk and incubated with primary anti-human antibody followed by secondary antibody. The proteins of interest were visualized using the LI-COR infrared imaging system (Odyssey).
Adenovirus transduction
OCI-ly10 and OCI-ly10-IR cells were seeded into a 24-well plate (1 × 106 cells/well) and transfected with a spliced XBP1 recombinant adenovirus vector (Ad-XBP1s) and negative green fluorescent protein (GFP) control adenovirus (Ad-GFP) in suspension. The adenovirus was a kindly gift from Professor Yong Liu (Wuhan University) [18]. The plates were sealed tightly and centrifuged at the low speed of 800 × g for 90 min in a flat fillet centrifuge. After centrifugation, the transduced cells were incubated at 37 °C for 24 h. GFP expression was examined 2 days after transduction using fluorescence microscopy. The transduced cells were used for cell growth inhibition or apoptosis assays.
Quantification of apoptosis
After drug treatment, the cells were washed twice with ice-cold PBS. Annexin V-APC reagent (Apoptosis Detection Kit; KeyGen) was diluted with Annexin V binding buffer at a ratio of 1:100 and added to each sample. After 5–15 min of incubation on ice in the dark, propidium iodide (PI) solution was added to each sample and immediately analyzed using flow cytometry (BD FACSCalibur). Data were analyzed using FlowJo software (Tree Star, Inc.).
Drug synergy analysis
Drug synergy was assessed online (https://synergyfinder.fimm.fi/synergy) using a Bliss independence model [19].
Calcium flux assay
Cells (1 × 105 cells/well) in a 384-well plate were incubated with the vehicle (dimethyl sulfoxide) or the indicated drug and 2 μM Fluo-4 NW solution for 30 min with 5% CO2 at 37 °C and for an additional 30 min at room temperature. While cells were incubated at room temperature, the agonist anti-IgM was diluted with assay buffer to 10 μg/mL. Assay buffer without anti-IgM was used for the control. Finally, the signal was captured using FDSS7000 (Hamamatsu A115) with excitation at 494 nm and emission at 516 nm wavelengths. After BCR stimulation with the anti-IgM treatment, fluorescence measurements were captured for an additional 4 min. The calcium (Ca2+) release signal was quantified using GraphPad software.
Xenograft study
Female NOD/SCID mice (17–19 g and 6–8 weeks old) were purchased (Jihui Laboratory Animal Care Co., Ltd, Shanghai, China) and used for tumor xenografts. The Institutional Animal Care and Use Committee (IACUC) of the Shanghai Institute of Materia Medica approved this project. The research was conducted in compliance with the Animal Welfare Act. The mice were maintained at the Shanghai Institute of Materia Medica Animal Facility. the OCI-ly10 cells (1 × 107) and OCI-ly10-IR cells (1 × 107) at a 1:1 PBS/Matrigel ratio were injected subcutaneously into the NOD-SCID mice. The mice were monitored closely for tumor growth through palpation and visual examination. When the tumor sizes were ~100 mm3, either the control vehicle (0.1% sodium dodecyl sulfate + 0.4% Kolliphor + 0.5% methylcellulose, q.d., p.o., no. = 6 mice), ibrutinib alone (5 or 15 mg/kg, q.d., p.o., no. = 6 mice), 2-DG alone (500 mg/kg, bid, p.o., no. = 6 mice), or a combination of ibrutinib and 2-DG was orally administered to the mice for 20 days. Xenograft tumor volume was measured as a function of tumor growth. Tumor volume was measured with a digital caliper and calculated as (length × width2) × 0.5. The animals were weighed every three days for 20 days.
Statistical analysis
The results were presented as means ± SEM. One-way analysis of variance (ANOVA) or the unpaired two-tailed Student’s t-test was used to determine the statistical significance for comparisons of two or more groups. Statistical analyses were performed using GraphPad Prism software (GraphPad Software, La Jolla, CA, USA). P < 0.05 was considered statistically significant (*P < 0.05, **P < 0.01, ***P < 0.001).
Results
Generation of ibrutinib-resistant ABC-DLBCL cells
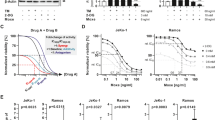
OCI-ly10 cells with ibrutinib resistance (OCI-ly10-IR cells) were derived from the OCI-ly10 cell line, an aggressive ABC subtype of DLBCL and sensitive to BTK inhibition by ibrutinib, by continuous exposure to ibrutinib [20] (Fig. 1a). After evaluating the proliferation rate, the OCI-ly10-IR cells showed a higher proliferation rate than shown by the OCI-ly10 cells (Fig. 1b). To verify the resistance of the OCI-ly10-IR cells to the BTK inhibitor, the cells were treated with ibrutinib and ACP-196 at a specified concentration. Both ibrutinib and ACP-196 showed lower efficacy in the OCI-ly10-IR cells than in the OCI-ly10 cells (Fig. 1c, d). In addition, other small molecular inhibitors that do not target BTK, such as baricitinib, MLN120B, LBH589, and AZD762, showed inhibition rates very similar to those of the OCI-ly10 or OCI-ly10-IR cells (Supplementary Fig. 1). This finding suggested that the OCI-ly10-IR cells were specifically resistant to BTK inhibitors. To further explore the cellular response to BCR downstream signaling, we performed real-time cytosolic calcium measurements to measure the Ca2+ flux in the OCI-ly10 and OCI-ly10-IR cells using a fluorescent calcium indicator (Fluo-4). We observed that the peak-rise or peak-decay time of Ca2+ flux was delayed in the OCI-ly10-IR cells compared with that in the OCI-ly10 cells, and the amplitude of Ca2+ flux in the OCI-ly10-IR cells was lower than that in the OCI-ly10 cells (Fig. 1e). We also noted that ibrutinib showed a weaker ability to inhibit Ca2+ flux in the OCI-ly10-IR cells than in the OCI-ly10 cells (Fig. 1f). Tumor growth and ibrutinib resistance were evaluated in vivo, and the results showed that the OCI-ly10-IR cells were resistant to 5 mg/kg ibrutinib treatment (Fig. 1g, h). We found that the OCI-ly10-IR cell line was specifically resistant to treatment with BTK inhibitors in vitro or in vivo.
a A schematic diagram of DLBCL ibrutinib-resistant cell line generation. b The OCI-ly10-IR cell growth rate was determined by proliferation assay; cells were counted for 7 days and the growth rates were compared to those of the parental OCI-ly10 cell line. c, d Parental and resistant cell lines were treated with the recommended doses of ibrutinib and acalabrutinib for 72 h and then assessed by MTS assay. e Differences in the Ca2+ kinetic traces of the OCI-ly10 and OCI-ly10-IR cells. f Ca2+ flux inhibition rate of ibrutinib in the OCI-ly10 and OCI-ly10-IR cells. Data are presented as the means ± SEM of the fluorescence ratios. g, h Growth curves of the xenograft tumors derived from OCI-ly10 cells and OCI-ly10-IR cells and treated with vehicle or ibrutinib alone (n = 6). *P < 0.05, **P < 0.01, ***P < 0.001 compared with the vehicle group. Standard error means (SEM) are indicated as error bars.
UPR suppression correlated closely with acquired ibrutinib resistance of DLBCL cell lines
The WES sequencing data showed that BTK and its related pathway-specific genes were not mutated in resistant cells (Supplementary Fig. 2). To analyze the underlying mechanism of drug resistance that might be involved in the established ibrutinib-resistant cells, a transcriptome microarray analysis was performed. The upstream regulation analysis showed that, in the OCI-ly10-IR cells, 523 genes were significantly upregulated with 691 genes significantly downregulated compared with the levels in the parental cells (Fig. 2a). The GSEA analysis with expression-altered genes showed that the UPR gene signature was downregulated (Fig. 2b). The three major mediators of the UPR pathways, ATF4, ATF6, and XBP1, were analyzed. While the ATF6 or ATF4 gene signatures were not markedly changed (Fig. 2c, d), the XBP1 signature was remarkably reduced (Fig. 2e). XBP1, an upstream transcription factor, and its 18 downstream genes were significantly downregulated according to the IPA analysis (Fig. 2f). To verify the transcriptome data, quantitative PCR validation was performed. The results showed that the expression of BIP, XBP1s, CHOP, DNAJB9, and other UPR marker genes was significantly downregulated (Fig. 2g), a finding similar to that of the XBP1 gene and its typical downstream genes S1PR1, MCFP2, FAS, and IL24 (Fig. 2h).
a Hierarchical clustering analysis of normalized log2-transformed total gene expression profiles in the OCI-ly10-IR cells compared with those of the OCI-ly10 parental cells (red, high; green, low). GSEA of the UPR gene signature (b), ATF6 gene signature (c), ATF4 gene signature (d), and XBP1 gene signature (e) in the OCI-ly10-IR cells compared with those of the OCI-ly10 cells; the false discovery rate (FDR) and P-values are indicated. f Ontological analysis of differentially expressed probes using IPA. g UPR marker genes BIP, IRE1, XBP1s, PERK, ATF4, ATF6, CHOP, and DNAJB9 were examined using qRT-PCR. h qRT-PCR validation of the differentially modified mRNAs involved in XBP1 downstream target genes. Data are represented as the means ± SEM of three separate experiments. *P < 0.05, **P < 0.01 compared with the DMSO-treated group.
Overexpression of XBP1s potentiated ibrutinib-induced cell growth inhibition and apoptosis
The UPR gene signature, especially the XBP1s gene signature, was downregulated in the ibrutinib-resistant DLBCL cells. To investigate the role of XBP1s in the ibrutinib-resistance mechanism of ABC-DLBCL cells, the XBP1s gene was overexpressed by transducing OCI-ly10-IR cells with spliced XBP1-GFP adenovirus (Ad-XBP1s). Quantitative PCR (Fig. 3a) and Western blot (Fig. 3b) analyses demonstrated that the mRNA and protein level of XBP1s was markedly increased after Ad-XBP1s transduction.
a Gene expression validation of the OCI-ly10-IR genes after transduced with XBP1s-overexpressing adenovirus. b Protein expression validation of the OCI-ly10-IR cells transduced with XBP1s-overexpressing adenovirus. Apoptosis rate was determined by propidium iodide (PI) and Annexin V-APC staining of the transduced OCI-ly10 (c) and OCI-ly10-IR (d) cells following treatment with 1 μM ibrutinib or DMSO. e OCI-ly10 cells were pretreated with TUDCA for 4 h and incubated at 37 °C in 200 nM ibrutinib. After 48 h, the cells were collected for Annexin V-PI analysis using flow cytometry. f Data are reported as ratios of the mean percentage of Annexin V-positive cells in three different experiments. Data are presented as the means ± SEM of three separate experiments. *P < 0.05 compared with the DMSO-treated group.
The overexpressed XBP1 OCI-ly10-IR cells were treated with the recommended concentration of ibrutinib. After ibrutinib treatment for 48 h, cell apoptosis was detected using flow cytometry. With 1 μM ibrutinib treatment, the cell apoptosis rate was increased in the XBP1s overexpression group compared with that of the control group (Fig. 3d). Similarly, the same function attributed to overexpressed XBP1s was found in the OCI-ly10 cells (Fig. 3c). Additionally, TUDCA, an inhibitor of the UPR, significantly reduced the apoptosis rate induced by ibrutinib in the OCI-ly10 cells. The percentage of apoptotic cells was 7.03%, 6.58%, 21.23%, and 14.61% in the DMSO, 100 μM TUDCA, 200 nM ibrutinib, and 200 nM ibrutinib + 100 μM TUDCA groups (Fig. 3e, f), respectively, which showed that the induction of apoptosis by ibrutinib can be reversed by TUDCA.
Chemical activation of the UPR with 2-DG synergistically enhanced ibrutinib-induced anticancer effects in vitro and in vivo
To improve ibrutinib efficacy through combination therapy, we combined the UPR activator 2-DG with ibrutinib. The combination enhanced the gene expression of XBP1, XBP1s, PERK, and CHOP (Fig. 4a). A combination of the two drugs resulted in a mean Bliss score of 31.384 (Fig. 4b). The addition of 0.5 mM 2-DG resulted in a nearly 5-fold decrease in the IC50 of ibrutinib (Fig. 4c). The cell viability percentage was 83.57%, 96.35%, and 51.60% for the ibrutinib-alone group, 2-DG-alone group, and ibrutinib + 2-DG combination group (Fig. 4d), respectively. The apoptotic rate was 6.9%, 11.93%, 8.79%, and 23.56% for the DMSO control group, 2-DG alone group, ibrutinib alone group, and ibrutinib + 2-DG combined group, respectively (Fig. 4e). The drug combination increased CHOP protein expression, downregulated MCL1, and enhanced PARP cleavage in vitro (Fig. 4f), and it significantly inhibited Ca2+ flux in the OCI-ly10-IR cells (Fig. 4g). To explore the combined effect of the ibrutinib + 2-DG combination and 2-DG groups in vivo, we established a xenograft model by transferring OCI-ly10-IR cells into NOD/SCID mice. A combination treatment of ibrutinib and 2-DG caused significant tumor regression in the OCI-ly10-IR-cell-bearing mice (Fig. 4h). The tumor weight in the mice of the combined treatment group was significantly reduced compared with that in the single treatment groups (P < 0.001) (Fig. 4i). All the mice with <20% body weight loss in each group tolerated both the single and combined treatments (P > 0.05). None of the mice died prematurely, suggesting that the lack of the combination correlated with systemic toxicity (Fig. 4j).
a Fold changes of the XBP1, XBP1s, PERK, ATF6, and CHOP genes are shown as indicated by qRT-PCRs performed 8 h after the treatment of the OCI-ly10-IR cells with 1 μM ibrutinib or 2-DG. b–d Percent cytotoxicity was measured by MTS assay, and percent of synergy was assessed using the Bliss independence method. Bliss scores denote the means of triplicate concentration matrices. Growth inhibition curves were derived from these data. e Percentages of early and late apoptotic cells 48 h after the OCI-ly10-IR cells were treated with IC50 concentrations of ibrutinib and 2-DG as measured by flow cytometry with Annexin V-FITC and PI staining. f Protein expression of CHOP, BCL2, BCLXL, MCL1, and PARP 48 h after the OCI-ly10-IR cells were treated with IC50 concentrations of ibrutinib and 2-DG, with β-actin used as the loading control. g Combined effect of ibrutinib and 2-DG on Ca2+ flux in the OCI-ly10-IR cells. h Growth curves of the xenograft tumors derived from the OCI-ly10-IR cells and treated with vehicle, ibrutinib alone (15 mg/kg, q.d.), 2-DG alone (500 mg/kg, bid), or a combination of ibrutinib and 2-DG (n = 6). i Tumor weight at the end of the 20-day period of different treatments. j Body weight measurements for every group. SEM is indicated as an error bar. *P < 0.05, **P < 0.01, ***P < 0.001. ##P < 0.01.
Discussion
The responses of various B-cell malignancies to ibrutinib are often limited by the ability of the cells to bypass the target via alternative pathways or the acquired mutations in BTK or a component in its pathway, such as the alternative activation of the NF-κB pathway or mutation of CD79A/B or MYD88 [21]. Although ibrutinib is critically more effective in ABC DLBCL than GCB DLBCL, it has limited efficacy as a single agent. The underlying primary or acquired resistance mechanisms of DLBCL to ibrutinib remain largely unknown.
Here, we generated the ibrutinib-resistant OCI-ly10-IR cell line, which was also resistant to the second-generation BTK inhibitor ACP-196 but was not resistant to inhibitors against other targets, such as JAK, HDAC, NF-κB, or CHK1 inhibitors (Supplementary Fig. 1). The OCI-ly10-IR cells showed a higher proliferation rate in vitro and in vivo (Fig. 1) and did not have BTK Cys481 or other BTK pathway-related reported mutations (Supplementary Fig. 2). Moreover, the transcriptional differences between the parental and resistant cells were explored. The UPR, especially the products of XBP1s-related genes, were downregulated significantly in the resistant cells. Previously, several analyses of specific ibrutinib-resistance mechanisms in CLL, MCL, and WM were studied [21,22,23]. It was shown that resistance to ibrutinib in some CLL or MCL patients was mostly due to mutations in BTK and PLCγ2 following prolonged treatment [24, 25]. BTK and PLCγ2 mutations can be drivers of ibrutinib resistance but are not necessary for acquired resistance because nearly 10%–20% of patients relapse without any detectable mutations in these genes [26]. After the introduction of the BTK C418S mutation into DLBCL cells [10, 27], either the subsequent increase in BCL2 expression or the reactivation of BTK-PLCγ2-ERK1/2 signaling led to resistance to ibrutinib. However, no remarkable loss of UPR- or XBP1s-related genes were found among the genes expressed in these cell lines, which were carrying the BTK C418S mutation.
Spliced XBP1 mRNA encodes the active 54 kDa form of the XBP1s basic leucine zipper transcription factor, which is involved in multiple signaling pathways, including MHC II gene regulation, plasma cell differentiation, angiogenesis, and endoplasmic reticulum (ER) stress [28]. Previously, it was reported that XBP1 splicing mediated a major adaptive pathway and was upregulated in various cancers, sustained the growth and survival of tumor cells [29] as a synthetic lethal partner of Myc, promoted tumorigenesis and progression in Myc-driven cancers [30], and induced immunosuppressive activity [31] related to tamoxifen-treated drug resistance in breast cancer [32, 33]. Thus, pharmacologically targeting the IRE1/XBP1 pathway may be a strategy against cancer, such as multiple myeloma (MM) [34, 35], CLL [36], Ewing’s sarcoma [37], and breast cancer [30].
However, it was also reported that lower XBP1 levels predicted a poorer response to the proteasome inhibitor bortezomib both in vitro and in MM patients [38], and decreased XBP1 activity may be associated with bortezomib resistance [39]. Similarly, XBP1s activity was suppressed in bortezomib-refractory primary MM, which may have induced bortezomib resistance by blocking commitment to plasma cell differentiation and lowering ER stress [40]. Previous studies showed that GCB-DLBCL tumors had significantly lower XBP1 gene signature expression than ABC-DLBCL tumors taken from clinical samples [41]. The overexpression of XBP1s in a GCB-DLBCL cell xenograft model significantly suppressed tumor growth [42]. We confirmed a decrease in the expression of XBP1s and its regulated genes in the BTK inhibitor-induced resistant cells, and the overexpression of XBP1s potentiated ibrutinib-induced cell growth inhibition and apoptosis (Fig. 3c–e). Furthermore, TUDCA, the chemical chaperone hampering UPR activation [43], significantly alleviated ibrutinib-induced apoptosis of sensitive ABC-DLBCL cells (Fig. 3f). Ibrutinib-induced UPR activation in sensitive cell lines but was not acquired for activation of the primary resistant DLBCL cell lines (Fig. 4a, b). Overall, these results suggested that UPR activation might be involved in ibrutinib-induced anti-DLBCL activity. Ibrutinib attenuated the development of acute lymphoblastic leukemia by driving ER stress-induced cell death [44]. However, the activation of the ER stress response is associated with the malignant progression of CLL. XBP1 deficiency decelerates the malignant progression of CLL-associated disease [36]. Furthermore, partial UPR activation promotes CLL survival, and the antitumor activity of ibrutinib might be partially due to inhibition of UPR activation [45]. Taken together, the function and role of the UPR or XBP1s might markedly differ in different hematological malignancies.
Published studies together with our current findings demonstrated that activating the UPR might enhance ibrutinib-induced anti-DLBCL activity. The UPR activator 2-DG [46] has shown sufficient safety in clinical trials but has limited efficacy. Its combination with berberine [47], a pan-UPR inhibitor [48], and a BCL2 antagonist [49] was shown to significantly reduce tumor growth, but nothing has been reported on the combination efficacy of ibrutinib and 2-DG. According to our results, a combination of ibrutinib and 2-DG synergistically inhibited cell proliferation and Ca2+ flux, induced cell apoptosis in vitro, and impeded tumor growth in the xenograft model. The primary mechanism showed that CHOP, a trigger of ER stress-induced apoptosis [50], was increased, the anti-apoptotic MCL1 protein was decreased, and PARP cleavage was increased upon administration of the combination treatment (Fig. 4f). Moreover, we found that the anticancer effect of the BTK inhibitor in primary resistant DLBCL cells was enhanced by 2-DG (Supplementary Fig. 3c–e). Although we initially found the loss of UPR in only one resistant cell line, the combined potential of ibrutinib and UPR activation in additional primary resistant cell lines suggests that a loss of UPR can be the general mechanism underlying ibrutinib resistance in diffuse large B-cell lymphoma, especially in the presence of wild-type BTK.
Chronic active BCR signaling is required for cell survival in ABC-DLBCL [20]. PLCγ2, downstream of BTK, influenced the Ca2+ flux in the ER via IP3 binding to the Ca2+ channel IP3 receptor, releasing Ca2+ from the ER [51]. Thus, ibrutinib inhibited Ca2+ flux through BTK inhibition. We demonstrated that the ibrutinib-induced Ca2+ flux change was significantly less profound in the resistant cells than it was in the sensitive cells (Fig. 1f). Previous studies indicated that the disruption of Ca2+ signaling can trigger ER stress [52]. An unknown mechanism is suggested to be critical for Ca2+ flux homeostasis tolerance in OCI-ly10-IR cells, which might contribute to the UPR signature in resistant cells.
To date, no evidence has revealed whether targeting of UPR-related pathways can potentiate the sensitivity of DLBCL to ibrutinib. In our study, the downregulation of UPR-related gene expression was linked to resistance to single-agent ibrutinib therapy, suggesting that UPR activation can drive cells to undergo apoptosis. Although the effect of other UPR pathways, such as ATF4 and ATF6, on ABC-DLBCL cell sensitivity to ibrutinib is unclear, we provide evidence herein that activation of the UPR via 2-DG can potentiate the activity of ibrutinib in both sensitive and resistant DLBCL cells. Based on findings from both the in vitro and in vivo models, a combination of ibrutinib and 2-DG is promising as a treatment for DLBCL and should be applied in further clinical trials.
References
Alizadeh AA, Eisen MB, Davis RE, Ma C, Lossos IS, Rosenwald A, et al. Distinct types of diffuse large B-cell lymphoma identified by gene expression profiling. Nature. 2000;403:503–11.
Pasqualucci L, Dalla-Favera R. The genetic landscape of diffuse large B-cell lymphoma. Semin Hematol. 2015;52:67–76.
Wilson WH, Young RM, Schmitz R, Yang Y, Pittaluga S, Wright G, et al. Targeting B cell receptor signaling with ibrutinib in diffuse large B cell lymphoma. Nat Med. 2015;21:922–6.
Younes A, Sehn LH, Johnson P, Zinzani PL, Hong X, Zhu J, et al. Randomized phase III trial of ibrutinib and plus cyclophosphamide, doxorubicin, vincristine, and prednisone in non-germinal center B-cell diffuse large B-Cell lymphoma. J Clin Oncol. 2019;37:1285–95.
IMBRUVICA (ibrutinib). Prescribing information. Horsham, PA: Janssen Biotech; Sunnyvale, CA: Pharmacyclics; 2018.
IMBRUVICA (ibrutinib). Summary of product characteristics. Beerse, Belgium: Janssen Pharmaceutical NV; 2018.
Hendriks RW, Yuvaraj S, Kil LP. Targeting Bruton’s tyrosine kinase in B cell malignancies. Nat Rev Cancer. 2014;14:219–32.
Kim JH, Kim WS, Ryu K, Kim SJ, Park C. CD79B limits response of diffuse large B cell lymphoma to ibrutinib. Leuk Lymphoma. 2016;57:1413–22.
Kapoor I, Li Y, Sharma A, Zhu H, Bodo J, Xu W, et al. Resistance to BTK inhibition by ibrutinib can be overcome by preventing FOXO3a nuclear export and PI3K/AKT activation in B-cell lymphoid malignancies. Cell Death Dis. 2019;10:924.
Kuo H-P, Ezell SA, Schweighofer KJ, Cheung LWK, Hsieh S, Apatira M, et al. Combination of ibrutinib and ABT-199 in diffuse large B-cell lymphoma and follicular lymphoma. Mol Cancer Ther. 2017;16:1246–56.
Hsu-Ping K, Scott AE, Sidney H, Karl JS, Leo WKC, Shiquan W. et al. The role of PIM1 in the ibrutinib-resistant ABC subtype of diffuse large B-cell lymphoma. Am J Cancer Res. 2016;6:2489–501.
Li H, Handsaker B, Wysoker A, Fennell T, Ruan J, Homer N, et al. The Sequence Alignment/Map format and SAMtools. Bioinformation. 2009;25:2078–9.
Cibulskis K, Lawrence MS, Carter SL, Sivachenko A, Jaffe D, Sougnez C, et al. Sensitive detection of somatic point mutations in impure and heterogeneous cancer samples. Nat Biotechnol. 2013;31:213–9.
Saunders CT, Wong WS, Swamy S, Becq J, Murray LJ, Cheetham RK. Strelka: accurate somatic small-variant calling from sequenced tumor-normal sample pairs. Bioinformatics. 2012;28:1811–7.
Boeva V, Popova T, Bleakley K, Chiche P, Cappo J, Schleiermacher G, et al. Control-FREEC: a tool for assessing copy number and allelic content using next-generation sequencing data. Bioinformatics. 2012;28:423–5.
Subramanian A, Tamayo P, Mootha VK, Mukherjee S, Ebert BL, Gillette MA, et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc Natl Acad Sci U S A. 2005;102:15545–50.
Schmittgen TD, Livak KJ. Analyzing real-time PCR data by the comparative C(T) method. Nat Protoc. 2008;3:1101–8.
Shao M, Shan B, Liu Y, Deng Y, Yan C, Wu Y, et al. Hepatic IRE1alpha regulates fasting-induced metabolic adaptive programs through the XBP1s-PPARalpha axis signalling. Nat Commun. 2014;5:3528. https://doi.org/10.1038/ncomms4528.
Ianevski A, He L, Aittokallio T, Tang J. Synergy-Finder: a web application for analyzing drug combination dose-response matrix data. Bioinformatics. 2017;33:2413–5.
Davis RE, Ngo VN, Lenz G, Tolar P, Young RM, Romesser PB, et al. Chronic active B-cell-receptor signalling in diffuse large B-cell lymphoma. Nature. 2010;463:88–92.
Zhang SQ, Smith SM, Zhang SY, Lynn Wang Y. Mechanisms of ibrutinib resistance in chronic lymphocytic leukaemia and non-Hodgkin lymphoma. Br J Haematol. 2015;170:445–56.
Liu TM, Woyach JA, Zhong Y, Lozanski A, Lozanski G, Dong S, et al. Hypermorphic mutation of phospholipase C, gamma2 acquired in ibrutinib-resistant CLL confers BTK independency upon B-cell receptor activation. Blood. 2015;126:61–8.
Jimmy L, Zhang LL, Wu W, Guo H, Li Y, Sukhanova M, et al. Activation of MYC, a bona fide client of HSP90, contributes to intrinsic ibrutinib resistance in MCL. Blood Adv. 2018;2:2039–51. https://doi.org/10.1182/bloodadvances.2018016048.
Furman RR, Cheng S, Lu P, Setty M, Perez AR, Guo A, et al. Ibrutinib resistance in chronic lymphocytic leukemia. N Engl J Med. 2014;370:2352–4.
Chiron D, Di Liberto M, Martin P, Huang X, Sharman J, Blecua P, et al. Cell-cycle reprogramming for PI3K inhibition overrides a relapse-specific C481S BTK mutation revealed by longitudinal functional genomics in mantle cell lymphoma. Cancer Discov. 2014;4:1022–35.
Lampson BL, Brown JR. Are BTK and PLCG2 mutations necessary and sufficient for ibrutinib resistance in chronic lymphocytic leukemia? Expert Rev Hematol. 2018;11:185–94.
Chen JG, Liu X, Munshi M, Xu L, Tsakmaklis N, Demos MG, et al. BTK (Cys481Ser) drives ibrutinib resistance via ERK1/2 and protects BTK (wild-type) MYD88-mutated cells by a paracrine mechanism. Blood. 2018;131:2047–59.
Yoshida H, Matsui T, Yamamoto A, Okada T, Mori K. XBP1 mRNA is induced by ATF6 and spliced by IRE1 in response to ER stress to produce a highly active transcription factor. Cell. 2001;107:881–91.
Xie H, Tang CH, Song JH, Mancuso A, Del Valle JR, Cao J, et al. IRE1alpha RNase-dependent lipid homeostasis promotes survival in Myc-transformed cancers. J Clin Invest. 2018;128:1300–16.
Zhao N, Cao J, Xu L, Tang Q, Dobrolecki LE, Lv X, et al. Pharmacological targeting of MYC-regulated IRE1/XBP1 pathway suppresses MYC-driven breast cancer. J Clin Invest. 2018;128:1283–99.
Cubillos-Ruiz JR, Bettigole SE, Glimcher LH. Molecular pathways: immunosuppressive roles of IRE1alpha-XBP1 signaling in dendritic cells of the tumor microenvironment. Clin Cancer Res. 2016;22:2121–6.
Ming J, Ruan S, Wang M, Ye D, Fan N, Meng Q, et al. A novel chemical, STF-083010, reverses tamoxifen-related drug resistance in breast cancer by inhibiting IRE1/XBP1. Oncotarget. 2015;6:40692–703.
Gomez BP, Riggins RB, Shajahan AN, Klimach U, Wang A, Crawford AC, et al. Human X-box binding protein-1 confers both estrogen independence and antiestrogen resistance in breast cancer cell lines. FASEB J. 2007;21:4013–27.
Harnoss JM, Le Thomas A, Shemorry A, Marsters SA, Lawrence DA, Lu M, et al. Disruption of IRE1alpha through its kinase domain attenuates multiple myeloma. Proc Natl Acad Sci USA. 2019;116:16420–9.
Mimura N, Fulciniti M, Gorgun G, Tai YT, Cirstea D, Santo L, et al. Blockade of XBP1 splicing by inhibition of IRE1alpha is a promising therapeutic option in multiple myeloma. Blood. 2012;119:5772–81.
Tang CH, Ranatunga S, Kriss CL, Cubitt CL, Tao J, Pinilla-Ibarz JA, et al. Inhibition of ER stress-associated IRE-1/XBP-1 pathway reduces leukemic cell survival. J Clin Invest. 2014;124:2585–98.
Tanabe Y, Suehara Y, Kohsaka S, Hayashi T, Akaike K, Mukaihara K, et al. IRE1alpha-XBP1 inhibitors exerted anti-tumor activities in Ewing’s sarcoma. Oncotarget. 2018;9:14428–43.
Ling SC, Lau EK, Al-Shabeeb A, Nikolic A, Catalano A, Iland H, et al. Response of myeloma to the proteasome inhibitor bortezomib is correlated with the unfolded protein response regulator XBP-1. Haematologica. 2012;97:64–72.
Nikesitch N, Tao C, Lai K, Killingsworth M, Bae S, Wang M, et al. Predicting the response of multiple myeloma to the proteasome inhibitor Bortezomib by evaluation of the unfolded protein response. Blood Cancer J. 2016;6:e432.
Leung-Hagesteijn C, Erdmann N, Cheung G, Keats JJ, Stewart AK, Reece DE, et al. Xbp1s-negative tumor B cells and pre-plasmablasts mediate therapeutic proteasome inhibitor resistance in multiple myeloma. Cancer Cell. 2013;24:289–304.
Shaffer AL, Wright G, Yang L, Powell J, Ngo V, Lamy L, et al. A library of gene expression signatures to illuminate normal and pathological lymphoid biology. Immunol Rev. 2006;210:67–85.
Bujisic B, De Gassart A, Tallant R, Demaria O, Zaffalon L, Chelbi S, et al. Impairment of both IRE1 expression and XBP1 activation is a hallmark of GCB DLBCL and contributes to tumor growth. Blood. 2017;129:2420–8.
de Almeida SF, Picarote G, Fleming JV, Carmo-Fonseca M, Azevedo JE, de Sousa M. Chemical chaperones reduce endoplasmic reticulum stress and prevent mutant HFE aggregate formation. J Biol Chem. 2007;282:27905–12.
Li ZWJ, Sheng L. Ibrutinib improves the development of acute lymphoblastic leukemia by activating endoplasmic reticulum stress-induced cell death. Pharmazie. 2018;73:294–9.
Krysov S, Steele AJ, Coelho V, Linley A, Sanchez Hidalgo M, Carter M, et al. Stimulation of surface IgM of chronic lymphocytic leukemia cells induces an unfolded protein response dependent on BTK and SYK. Blood. 2014;124:3101–9.
Xi HKM, Liu H, Wangpaichitr M, You M, Liu X, Savaraj N, et al. 2-Deoxy-D-glucose activates autophagy via endoplasmic reticulum stress rather than ATP depletion. Cancer Chemother Pharmacol. 2011;67:899–910.
Huang H, Liu H, Liu C, Fan L, Zhang X, Gao A, et al. Disruption of the unfolded protein response (UPR) by lead compound selectively suppresses cancer cell growth. Cancer Lett. 2015;360:257–68.
Fan LX, Liu CM, Gao AH, Zhou YB, Li J. Berberine combined with 2-deoxy-D-glucose synergistically enhances cancer cell proliferation inhibition via energy depletion and unfolded protein response disruption. Biochim Biophys Acta. 2013;1830:5175–83.
Lucantoni F, Dussmann H, Llorente-Folch I, Prehn JHM. BCL2 and BCL(X)L selective inhibitors decrease mitochondrial ATP production in breast cancer cells and are synthetically lethal when combined with 2-deoxy-D-glucose. Oncotarget. 2018;9:26046–63.
Yang J, Wang Z, Fang Y, Jiang J, Zhao F, Wong H. Pharmacokinetics, pharmacodynamics, metabolism, distribution, and excretion of carfilzomib in rats. Drug Metab Dispos. 2011;39:1873–82.
Kurosaki T, Maeda A, Ishiai M, Hashimoto A, Inabe K, Takata M. Regulation of the phospholipase C-gamma2 pathway in B cells. Immunol Rev. 2000;176:19–29.
Carreras-Sureda A, Pihan P, Hetz C. Calcium signaling at the endoplasmic reticulum: fine-tuning stress responses. Cell Calcium. 2018;70:24–31.
Acknowledgements
This work was supported by the Strategic Priority Research Program of the Chinese Academy of Sciences (CAS, XDA12020208), National Major Scientific and Technological Special Project for “Significant New Drugs Development” (2018ZX09711002-007-001 and 2018ZX09711002-011-011), the National Natural Science Foundation of China for Innovation Research Group (81821005), Shanghai municipal commission of science and technology (biomedicine, 19431900700, and 18431907100), and Ministry of Science and Technology of the People’s Republic of China.
Author information
Authors and Affiliations
Contributions
YBZ and JL developed hypotheses. YBZ, XTZ, and JL conceived the project, provided supervision, and are responsible for all data, figures, and text; XTZ performed the research and analyzed the data; HLW and BF conducted some of the experiments and data analysis; LX, ZJW, YQX, WBW, JNL, and AHG assisted with part of data analysis; XBH, WJK, TCD, and CMX assisted with the animal studies; XTZ and YBZ wrote the manuscript. All authors approved the final version of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Supplementary information
Rights and permissions
About this article
Cite this article
Zhang, Xt., Hu, Xb., Wang, Hl. et al. Activation of unfolded protein response overcomes Ibrutinib resistance in diffuse large B-cell lymphoma. Acta Pharmacol Sin 42, 814–823 (2021). https://doi.org/10.1038/s41401-020-00505-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41401-020-00505-3