Abstract
KRAS is one of the most important proto-oncogenes. Its mutations occur in almost all tumor types, and KRAS mutant cancer is still lack of effective therapy. Prenyl-binding protein phosphodiesterase-δ (PDEδ) is required for the plasma membrane association and subsequent activation of KRAS oncogenic signaling. Recently, targeting PDEδ has provided new promise for KRAS mutant tumors. However, the therapeutic potential of PDEδ inhibition remains obscure. In this study, we explored how PDEδ inhibition was responded in KRAS mutant cancer cells, and identified KRAS mutant subset responsive to PDEδ inhibition. We first performed siRNA screen of KRAS growth dependency of a small panel of human cancer lines, and identified a subset of KRAS mutant cancer cells that were highly dependent on KRAS signaling. Among these cells, only a fraction of KRAS-dependent cells responded to PDEδ depletion, though KRAS plasma membrane association was effectively impaired. We revealed that the persistent RAF/MEK/ERK signaling seemed responsible for the lack of response to PDEδ depletion. A kinase array further identified that the feedback activation of EPH receptor A2 (EPHA2) accounted for the compensatory activation of RAF/MEK/ERK signaling in these cells. Simultaneous inhibition of EPHA2 and PDEδ led to the growth inhibition of KRAS mutant cancer cells. Together, this study gains a better understanding of PDEδ-targeted therapeutic strategy and suggests the combined inhibition of EPHA2 and PDEδ as a potential therapy for KRAS mutant cancer.
Similar content being viewed by others
Introduction
RAS proteins, encoded by an important family of proto-oncogenes, belong to a class of proteins called small GTPases and are involved in transmitting signals within cells [1, 2]. Alterations to RAS proto-oncogenes, in the form of point mutations, have been identified in almost all tumors. Oncogenic mutations to specific amino acids maintain RAS in a constitutively active GTP-bound state, resulting in the continuous activation of the RAS signal transduction pathway [3,4,5]. This signal is transmitted to downstream pathways, particularly the RAS/RAF/MEK/ERK and PI3K/AKT/mTOR signaling pathways, which further affect a plethora of cellular events, including the cell cycle progression, cell transformation, proliferation, apoptosis, migration, and differentiation [3, 6, 7]. The most commonly mutated RAS isoform is KRAS4B, often referred to as KRAS in human cells, which occurs in more than 90% of pancreatic tumors, 45% of colorectal tumors and 30% of lung tumors; these facts make the KRAS protein an attractive target in anticancer drug discovery [8]. However, decade-long efforts in exploiting this target have failed to yield clinically effective therapies, which is believed to be due to the special structure of the KRAS protein, the high affinity of KRAS for GTP and the complexity of the downstream signaling pathways of KRAS [2, 8,9,10].
Notably, recent studies have highlighted a new therapeutic strategy that targets a key step in the process of KRAS membrane localization [1]. The KRAS protein requires localization to the inner leaflet of the plasma membrane for oncogenic activity. The guanine nucleotide dissociation inhibitor (GDI)-like solubilization factor phosphodiesterase-δ (PDEδ) is required for the correct localization and signaling of farnesylated KRAS [11]. PDEδ belongs to a large family of phosphodiesterases (PDEs). It possesses a hydrophobic pocket and could interact with a farnesylated hydrophobic cysteine residue at the C-terminus of the KRAS protein [12]. This interaction sequesters KRAS from the cytosol, preventing KRAS from binding to endomembranes and thereby facilitating its diffusion in the cytoplasm. Interfering with the binding of mammalian PDEδ to KRAS has been shown to provide a novel way to suppress oncogenic RAS signaling. Zimmermann et al. discovered the first small molecule inhibitor, deltarasin, that bound the farnesyl-binding pocket of PDEδ and disrupted KRAS signaling [13].
While this novel strategy appears to be promising for conquering this long-pursued target, it is also noted that in addition to KRAS, PDEδ is crucial for maintaining the membrane localization of other RAS family members, such as palmitoylated H- and N-RAS and solely farnesylated RHEB [12]. Moreover, PDEδ is reported to have an interactome composed of multiple proteins, such as RAB23, CDC42, and CNP proteins [14]. Inhibition of PDEδ is expected to have broad cellular effects, yielding uncertainties in KRAS-dependent cancer cells. Moreover, the anticancer activity of newly discovered small molecule inhibitors targeting PDEδ remains obscure. The mechanism of deltarasin, the first reported KRAS-PDEδ inhibitor, appears not to be limited to KRAS signaling [13]. The complexity of the role of PDEδ and the mechanism of action of PDEδ inhibition in KRAS-dependent cells require an in-depth understanding of PDEδ-targeted therapies.
This study aims to gain a better understanding of the effects of PDEδ inhibition in KRAS mutant cancer cells. Given the heterogeneous proprieties of KRAS mutant cancer, this study also seeks to identify the tumor subset responsive to PDEδ inhibition. By profiling a panel of KRAS mutant cancer cell lines, we discovered that PDEδ inhibition was effective in a small proportion of KRAS mutant cancer and was closely associated with the alteration of RAF/MEK/ERK signaling in these cells. We also discovered that the feedback activation of EPH receptor A2 (EPHA2) accounted for the compensatory activation of RAF/MEK/ERK signaling, leading to resistance to PDEδ inhibition. We therefore propose a new therapeutic strategy that concurrently inhibits PDEδ and EPHA2 for the treatment of KRAS mutant cancer.
Materials and methods
Reagents and antibodies
Deltarasin (S7224), BVD-523 (S7854), AZD6244 (S1008), and BMS-754807 (S1124) were obtained from Selleck (Houston, TX, USA). ALW-II-41-27 was obtained from AbMole (Houston, TX, USA). The antibodies used in this study were as follows: antibodies against phospho-c-RAF (Ser338) (9427s), c-RAF (53745s), phospho-p44/42 MAPK (ERK1/2) (Thr202/Tyr204) (4370L), p44/42 MAPK (ERK1/2) (4695s), phospho-AKT (Ser473) (4060s), AKT (4691s), phospho-p70 S6 kinase (Thr389) (9234s), p70 S6 kinase (2708s), phospho-EPHA2 (Tyr594) (3970s), phospho-EphA2 (Tyr588) (12677s), EPHA2 (6997s), IGF-I receptor β (3027s), phospho-IGF-I receptor β (Tyr1135) (3918s), phospho-WNK-1 (Thr60) (4946s), and phospho-GSK-3β (Ser9) (5558s) were all from Cell Signaling Technology (Cambridge, MA, USA). An antibody against KRAS (101667-T32) was purchased from Sino Biological, and an antibody against PDE6D (ab96825) was purchased from Abcam (Shanghai, China). GAPDH (60004-1-lg) was obtained from Proteintech (Rosemont, IL, USA).
Cell lines
NCI-A549, NCI-H23, NCI-H1792, NCI-H441, H460, and NCI-H358 cells were obtained from the American Type Culture Collection (ATCC, Manassas, VA, USA). NCI-SW1573, NCI-H2030, AsPC-1, Panc 02.03, Panc 10.05, and H1944 cells were obtained from Cobioer (Nanjing, China). All cell lines were routinely authenticated by short tandem repeat analysis (Genesky Biotechnologies, Shanghai, China) and were maintained in the appropriate culture medium suggested by the suppliers.
Plasmid, shRNA, and siRNA transfection
A KRAS-GFP plasmid (HG12259-ANG) was obtained from Sino Biological (Beijing, China). For siRNA transfection, cells were plated at 30%–60% confluence in the OPTI-MEM serum-free medium and transfected with a specific siRNA duplex using Lipofectamine RNAiMAX (Invitrogen, CA, USA), according to the manufacturer’s instructions. siRNAs were ordered from Shanghai GenePharma (Shanghai, China). The sequences of the scrambled negative control (NC) and siRNAs targeting the indicated genes were as follows:
NC 5′-UUCUCCGAACGUGUCACGUTT-3′
KRAS #1 5′-GAGGAGUACAGUGCAAUGATT-3′
KRAS #2 5′-GGAUUCCUACAGGAAGCAATT-3′
PDE6D #1 5′-CUUCGACAGAACAAAUGGATT-3′
PDE6D #2 5′-AUGCCUAGAAGAAUGGUUCUTT-3′
siEPHA2 #1 5′-GCUCAAGUUUACUGUACGUTT-3′
siEPHA2 #2 5′-GUCCGUGUCUACUACAAGATT-3′
Cell viability assay
SRB assay
Cells were seeded in 96-well plates overnight. Sulforhodamine B assays (Thermo Fisher Scientific) were performed after treatment with the indicated reagents for 72 h. The absorbance (optical density [OD]) was read at a wavelength of 515–560 nm on an ELISA plate reader. The growth inhibition rate was calculated according to (OD treated/OD control) × 100%, in which untreated cells served as the control. The experiment was repeated for at least three times. The data represent the means of triplicate analyses of a representative experiment.
Crystal violet assay
Cells were seeded in 12-well plates overnight. A crystal violet assay was performed after treatment with the indicated reagents for 96 h. OD values were read at a wavelength of 515–560 nm on an ELISA plate reader. The growth inhibition rate was calculated according to (OD treated/OD control) × 100%, in which untreated cells served as the control. The experiment was repeated for at least three times. The data represent the mean from four replicate analyses of a representative experiment.
Immunoblotting analysis
Cells were lysed using preheated 2% SDS by vortexing vigorously for 10–15 s at maximum speed, followed by boiling for 30 min. The same amount of protein was subjected to SDS–PAGE, transferred to nitrocellulose membranes, and blocked for 1 h at room temperature with 3% BSA in 1 × Tris-buffered saline-Tween-20 (TBST; 25 mM Tris, 150 mM NaCl, and 2 mM KCl, pH 7.4, supplemented with 0.2% Tween-20). Blotting was performed with primary antibodies at 4 °C overnight. After washing the membranes with TBST three times for a total of 30 min, horseradish peroxidase-conjugated secondary antibodies were incubated at room temperature for 1 h. The membranes were washed three times with TBST for 30 min, and the proteins were visualized with an enhanced chemiluminescence assay (Thermo Fisher Scientific) or Femto chemiluminescence assay (Thermo Fisher Scientific).
Fluorescence microscopy
Cells were fixed with 4% paraformaldehyde and then treated with DAPI for 15–20 min. Images were captured using a laser scanning confocal microscope (Lavision Biotec TriM Scope). At least 50 cells were counted per sample for quantification. Cells with apparent KRAS plasma membrane localization were counted and then normalized to the total cell counts. The data represent the means of three independent experiments.
Phospho-kinase antibody arrays and phospho-RTK array kit
Cells (1 × 106) seeded in a 10-cm dish were treated with the indicated reagents for 48 h. Cells were lysed, and phosphorylation changes were detected by a Phospho-RTK Array Kit (#ARY001B, R&D Systems) and phospho-kinase antibody array (ARY003B, R&D Systems) following the manufacturer’s instructions.
Results
KRAS dependency varies in KRAS mutant cancer cell lines
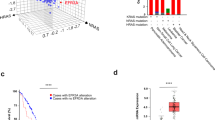
Previous studies have revealed that KRAS mutations, though they occur widely in human cancer, may not necessarily give rise to KRAS-dependent cell growth [15]. As such, we first screened the growth dependency of a small panel of human cancer cell lines bearing well-documented KRAS-activating mutations, mainly highly frequent mutations to G12 residuals (Fig. 1a). Indeed, knocking down KRAS expression using two independent siRNAs resulted in diverse effects on cell growth in these cells. Among the 12 tested cell lines, four cell lines, namely, Panc 02.03, H460, H1792, and H358, showed substantial growth suppression. Some cell lines, such as SW1573, H23, A549, AsPC-1, and Panc 10.05 cells, showed a moderate effect in response to KRAS depletion, while the remaining cells barely responded and were considered KRAS-independent cells (Fig. 1b). The knockdown efficiency was comparable among the tested cells.
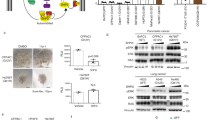
KRAS dependency varies in KRAS mutant cancer cell lines. a KRAS mutation status of cell lines used in this study. NSCLC, non-small cell lung cancer; PDAC, pancreatic ductal adenocarcinoma b KRAS mutant cell viability upon KRAS depletion. Cells were treated with KRAS siRNAs (siKRAS #1, #2) for 96 h. Cell viability was measured by crystal violet staining assay (upper panel). siRNA interference efficiency was measured by immunoblotting analysis (lower panel). The error bars represent the mean ± SD of four replicates. *P < 0.05; **P < 0.01; ***P < 0.001, n.s., not significant. c, d MAPK and AKT signaling changes upon KRAS depletion in KRAS-dependent (c) and KRAS-independent (d) cells. Cells were treated with KRAS siRNAs (siKRAS #1, #2) for 48 h before being subjected to immunoblotting analysis. e Cell sensitivity to ERK inhibitors. Cells were treated with the ERK inhibitor BVD-523 for 72 h, and cell viability was measured by SRB assay. The error bars represent the mean ± SD of triplicates. NC, scrambled siRNA used as a negative control
KRAS activation is known to drive oncogenic malignancy, mainly via two downstream pathways, the RAF/MEK/ERK pathway, also known as the MAPK/ERK pathway, and the phosphoinositide 3-kinase (PI3K) pathway [6]. We hence examined the key signaling molecules in these two downstream pathways. In both KRAS-dependent and KRAS-independent cancer cells, KRAS depletion led to the universal downregulation of the MAPK/ERK pathway, as indicated by the decreased phosphorylation levels of c-RAF and ERK, while the impact on the PI3K/AKT pathway was variable among the cell lines (Fig. 1c, d; S1a). Consistently, pharmacological inhibition of the MAPK/ERK pathway using the clinically tested ERK kinase inhibitor BVD-523 resulted in substantial cell growth inhibition in KRAS-dependent A549 and H358 cells, which was in great contrast to KRAS-independent cell lines, such as H441 and H2030 (Fig. 1e; S1b). These results identified a subset of KRAS mutant cancer cells whose growth was highly dependent on KRAS activation and established a linkage between KRAS oncogenic activation and MAPK/ERK signaling activation instead of the PI3K/AKT pathway.
PDEδ depletion is not necessarily associated with cell growth inhibition in KRAS-dependent cells
We then attempted to assess whether the disruption of PDEδ in KRAS-dependent cells could lead to growth inhibition. As the selectivity of the reported PDEδ inhibitors remains in question [13, 16], we chose to use two independent siRNAs to specifically knockdown PDE6D, which encodes PDEδ, and cell growth was then assessed using a crystal violet staining assay. In nine KRAS-dependent or partially dependent cells, only a small fraction of the cell lines, namely, A549, H460, and SW1573 cells, responded partially to PDEδ, as indicated by partially suppressed growth inhibition (Fig. 2a, upper panel); however, PDEδ was similarly depleted in all the tested cells (Fig. 2a, lower panel). In KRAS-independent cells, as expected, none of the cells responded to PDEδ depletion (Fig. 2a).
MAPK/ERK signaling alteration is associated with the response to PDEδ inhibition in KRAS-dependent cancer cells. a KRAS mutant cell viability upon PDEδ depletion. KRAS-dependent cells were treated with PDEδ siRNAs (siPDE6D #1, #2) for 96 h. Cell viability was measured by crystal violet staining assay (upper panel). siRNA interference efficiency was measured by immunoblotting (lower panel). The error bars represent the mean ± SD of four replicates. *P < 0.05; **P < 0.01; ***P < 0.001, n.s., not significant. b, c MAPK and AKT signaling changes upon PDEδ depletion in PDEδ-dependent (b) and PDEδ-independent cells (c). Cells were treated with siRNAs (siPDE6D #1, #2) for 48 h before being subjected to immunoblotting analysis. NC, scrambled siRNA used as a negative control
We asked whether the differential cell response upon PDEδ knockdown could possibly result from the differential impact on MAPK/ERK signaling. By simultaneously analyzing the MAPK/ERK pathway in PDEδ-responsive (red) and nonresponsive cells (blue), we discovered that siRNA depletion of PDEδ effectively diminished p-S6K, a surrogate marker for PDEδ function [12, 13, 16]. However, MAPK/ERK signaling was only decreased in PDEδ-responsive cells (Fig. 2b). For example, in A549, H460, and SW1573 cells, phosphorylation levels of c-RAF and ERK were effectively decreased. In contrast, in the rest of the cells in which PDEδ knockdown barely affected cell growth (Fig. 2c), alterations to RAF/MEK/ERK signaling were not observed. These results suggested that cell growth inhibition was associated with diminished MAPK/ERK signaling.
Together, these results suggest that PDEδ depletion is not necessarily associated with cell growth inhibition in KRAS-dependent cells. Effective MAPK/ERK signaling blockage is required for the growth inhibition.
PDEδ disruption effectively disrupts KRAS protein plasma membrane localization
We were then interested in understanding why PDEδ disruption in some cancer cells failed to diminish MAPK/ERK signaling, as KRAS impairment in these cells was apparently associated with MAPK/ERK signaling (Fig. 1c). Given that PDEδ is required for KRAS plasma membrane association prior to KRAS activation, we analyzed KRAS membrane localization in these cells. To this end, GFP-conjugated KRAS was transfected into the cells to visualize KRAS subcellular localization using fluorescence microscopy. In H460 cells, which are sensitive to PDEδ deficiency, knockdown of PDE6D gene expression efficiently impaired KRAS membrane association (Fig. 3a, left panel). Similar results were obtained using deltarasin, the first reported PDEδ inhibitor that has been shown to effectively disrupt KRAS plasma membrane localization [13] (Fig. 3b, left panel).
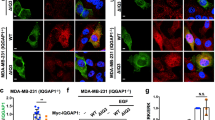
KRAS membrane localization upon PDEδ disruption in KRAS-dependent cancer cells. a Intracellular localization of KRAS upon PDEδ knockdown. Cells stably expressing GFP-KRAS were transfected with PDE6D siRNA for 48 h, and KRAS localization was visualized by GFP fluorescence. b Intracellular localization of KRAS upon PDEδ inhibition. Cells were treated with 5 μM deltarasin for the indicated time. KRAS membrane association was detected as described in (a). c Quantification of (a) and (b). Cells with evident KRAS membrane association were counted and normalized to the total cell counts. At least 50 cells were counted per sample. Scale bar, 10 μm. The error bars represent the mean ± SD of three independent analyses. **P < 0.01; ***P < 0.001. NC, scrambled siRNA used as a negative control
The assay was also conducted in AsPC-1 cells, in which PDE6D siRNA failed to inhibit cell growth or disrupt MAPK/ERK signaling. Interestingly, in PDEδ-nonresponsive cells, KRAS membrane localization was similarly impaired (Fig. 3a–c). These results indicated that KRAS plasma membrane association was impaired upon PDEδ depletion in PDEδ-nonresponsive cells, suggesting that the sustained MAPK/ERK pathway activation could possibly result from compensatory upstream signaling.
Feedback activation of EPHA2 activates the MAPK/ERK pathway upon PDEδ disruption
To explore the possible involvement of compensatory pathways, we used a phospho-RTK antibody and phospho-kinase antibody arrays, which covered 49 phosphorylated human receptor tyrosine kinases (RTKs) and 43 intracellular kinase phosphorylation sites, respectively. This approach allowed us to screen the kinases that were activated upon PDEδ disruption. At 48 h following PDEδ knockdown, we observed the feedback activation of several protein kinases, including IGF1R, EPHA2, GSK3α/β, and WNK-1, in H358 cells, which were shown to be nonresponsive to PDEδ knockdown (Fig. 4a, b). Among these altered kinases, the upregulation of IGF1R and EPHA2 induced by siPDE6D treatment was confirmed in an independent immunoblotting analysis (Fig. 4c).
Feedback activation of EPHA2 results in compensatory MAPK activation in PDEδ-depleted cells. a, b Phospho-RTK antibody and phospho-kinase antibody arrays. Cells were treated with siPDE6D (#1) for 48 h, and the cell lysates were subjected to phospho-RTK antibody array (a) or phospho-kinase antibody array (b). c Immunoblotting analysis confirmed the feedback activation of EPHA2 and IGF1R in PDEδ-depleted cells. H358 cells were treated with the indicated siRNA for 48 h before being subjected to immunoblotting. d MAPK signaling upon concurrent inhibition of PDEδ and EPHA2. Cells were transfected with the indicated siRNAs for 48 h before exposure to the EPHA2 inhibitor ALW-II-41-27 for 4 h. e MAPK signaling upon concurrent inhibition of PDEδ and IGF1R. Cells were transfected with the indicated siRNAs for 48 h before exposure to the IGF1R inhibitor BMS-754807 for 5 h. f, g Cell viability upon concurrent inhibition of PDEδ and MEK or EPHA2. KRAS-dependent Panc 10.05 cells were treated with PDEδ siRNA in combination with the MEK inhibitor AZD6244 or the EPHA2 inhibitor ALW-II-41-27 for 96 h. Cell viability was measured by crystal violet staining assay. NC, scrambled siRNA used as a negative control. h A schematic model
We next tested whether the feedback activation of IGF1R and EPHA was responsible for persistent MAPK/ERK activation upon PDE6D knockdown. To this end, cells were treated simultaneously with IGF1R or EPHA2 inhibitors [17, 18] and PDE6D siRNA. MAPK/ERK signaling was examined using an immunoblotting analysis. It was observed that the EPHA2 inhibitor ALW-II-41-27 could diminish the MAPK/ERK pathway signaling in a panel of PDEδ-nonresponsive cell lines, including H358, Panc 02.03, Panc 10.05, and AsPC-1 cells, in which PDE6D siRNA alone failed to reduce KRAS-induced MAPK/ERK signaling (Fig. 4d; S2a). In contrast, treatment with the IGF1R inhibitor BMS-754807 barely affected the phosphorylation of c-RAF and ERK upon PDEδ knockdown (Fig. 4e; S2b).
These results suggested that the feedback activation of EPHA2 activated the MAPK/ERK pathway upon PDEδ disruption. We further applied the EPHA2 inhibitor ALW-II-41-27 or EPHA2 siRNA to test whether the cells could be re-sensitized to PDEδ inhibition. Indeed, we discovered that while PDEδ depletion alone barely affected cell viability, combined treatment with ALW-II-41-27 (Fig. 4g) and EPHA2 siRNA (Fig. S2c) largely inhibited the growth of Panc 10.05 cells. Similar results were obtained by combining a MEK inhibitor and PDE6D siRNA (Fig. 4f).
Discussion
Targeting KRAS plasma membrane association has been recently proposed for exploration as a new therapeutic opportunity for KRAS mutant cancer, which highlights PDEδ as an attractive target to impede KRAS oncogenic signaling. However, the current understanding of this new strategy is very limited, especially considering that the complex signaling network of PDEδ is known to be involved. In addition, given the heterogeneous properties of KRAS mutant cancer, it is conceivable that PDEδ inhibition may not yield therapeutic benefits in broad KRAS mutant contexts. It will be important to stratify the subset responsive to PDEδ inhibition. In this study, we discovered that only a fraction of KRAS-dependent cells responds to PDEδ inhibition, although KRAS plasma membrane association is efficiently impaired. The feedback activation of EPHA2 accounts for the limited response to PDEδ inhibition via a mechanism that involves the compensatory activation of the MAPK/ERK signaling (Fig. 4h). These insights help gain a better understanding of the PDEδ-targeted therapeutic strategy and suggest the combined inhibition of EPHA2 and PDEδ as a potential therapy for KRAS mutant cancer.
It remains unclear how EPHA2 is specifically activated upon PDEδ inhibition in KRAS-dependent cancer. Currently, there is no evidence suggesting a direct link between PDEδ and EPHA2. Limited evidence thus far may suggest the involvement of CDC42, which possibly mediates the association between PDEδ and EPHA2. EPHA2 is known to regulate RHO/RAC/CDC42 [19, 20]. It has also been shown that PDEδ could interact with CDC42 [14]. One possibility is that the impairment of PDEδ could induce the feedback response of CDC42, which further provokes EPHA2 via a feedback loop. This hypothesis will be interesting to explore in the future.
In addition, a large number of studies have shown that EPHA2 interacts with KRAS and downstream signaling pathways [21] EPHA2 belongs to receptor tyrosine kinase families that regulate multiple downstream signaling pathways, including the MAPK/ERK and PI3K/AKT/mTOR pathways [19, 22, 23]. It is possible that impaired PDEδ causes KRAS to lose control over its direct downstream MAPK/ERK signaling pathway. In this scenario, a higher level of EPHA2 or its active status is more likely to allow the feedback activation of the MAPK/ERK pathway. Indeed, we compared the EPHA2 expression in KRAS-dependent cells using Cell Line Encyclopedia (CCLE) data sets and found that EPHA2 expression was relatively higher in cells that were nonresponsive to PDEδ inhibition than in those that were responsive to PDEδ inhibition (Fig. S3a). In parallel, the expression level of IGF-IR did not differ between PDEδ-responsive and nonresponsive cell lines (Fig. S3b). We further examined the phosphorylation level of EPHA2 in these cells. EPHA2 was often activated, as indicated by the basal EPHA2 phosphorylation level, in PDEδ-nonresponsive cells (Fig. S3c). Certainly, the two aforementioned possibilities could exist in parallel. A more in-depth study will be required to carefully dissect how EPHA2 is specifically provoked upon PDEδ depletion.
Finally, in this study, we discovered that EPHA2-induced MAPK/ERK signaling is essential for determining KRAS mutant cancer survival. Due to the complexity of signaling pathways involving KRAS and the heterogeneous properties of KRAS mutants, it remains unclear whether other pathways beyond EPHA2-MAPK signaling are involved in determining the survival of KRAS mutant cancer. It is possible that the novel strategy of combining PDE6D and EPHA2 inhibition discovered in this study may be applicable to only a subset of KRAS mutant cancer.
Change history
16 January 2024
A Correction to this paper has been published: https://doi.org/10.1038/s41401-023-01208-1
References
Cox AD, Der CJ. Ras history: the saga continues. Small GTPases. 2010;1:2–27.
Cox AD, Fesik SW, Kimmelman AC, Luo J, Der CJ. Drugging the undruggable RAS: mission possible? Nat Rev Drug Discov. 2014;13:828–51.
Papke B, Der CJ. Drugging RAS: know the enemy. Science. 2017;355:1158–63.
Karnoub AE, Weinberg RA. Ras oncogenes: split personalities. Nat Rev Mol Cell Biol. 2008;9:517–31.
Fehrenbacher N, Bar-Sagi D, Philips M. Ras/MAPK signaling from endomembranes. Mol Oncol. 2009;3:297–307.
Hubbard PA, Moody CL, Murali R. Allosteric modulation of Ras and the PI3K/AKT/mTOR pathway: emerging therapeutic opportunities. Front Physiol. 2014;5:478.
Martini M, De Santis MC, Braccini L, Gulluni F, Hirsch E. PI3K/AKT signaling pathway and cancer: an updated review. Ann Med. 2014;46:372–83.
Pylayeva-Gupta Y, Grabocka E, Bar-Sagi D. RAS oncogenes: weaving a tumorigenic web. Nat Rev Cancer. 2011;11:761–74.
Thompson H. US National Cancer Institute’s new Ras project targets an old foe. Nat Med. 2013;19:949–50.
Mansi L, Viel E, Curtit E, Medioni J, Le Tourneau C. Targeting the RAS signalling pathway in cancer. Bull Cancer. 2011;98:1019–28.
Baehr W. Membrane protein transport in photoreceptors: the function of PDEdelta: the Proctor lecture. Invest Ophthalmol Vis Sci. 2014;55:8653–66.
Chandra A, Grecco HE, Pisupati V, Perera D, Cassidy L, Skoulidis F, et al. The GDI-like solubilizing factor PDEdelta sustains the spatial organization and signalling of Ras family proteins. Nat Cell Biol. 2011;14:148–58.
Zimmermann G, Papke B, Ismail S, Vartak N, Chandra A, Hoffmann M, et al. Small molecule inhibition of the KRAS-PDEdelta interaction impairs oncogenic KRAS signalling. Nature. 2013;497:638–42.
Kuchler P, Zimmermann G, Winzker M, Janning P, Waldmann H, Ziegler S. Identification of novel PDEdelta interacting proteins. Bioorg Med Chem. 2018;26:1426–34.
Singh A, Greninger P, Rhodes D, Koopman L, Violette S, Bardeesy N, et al. A gene expression signature associated with “K-Ras addiction” reveals regulators of EMT and tumor cell survival. Cancer Cell. 2009;15:489–500.
Papke B, Murarka S, Vogel HA, Martin-Gago P, Kovacevic M, Truxius DC, et al. Identification of pyrazolopyridazinones as PDEdelta inhibitors. Nat Commun. 2016;7:11360.
Bid HK, London CA, Gao J, Zhong H, Hollingsworth RE, Fernandez S, et al. Dual targeting of the type 1 insulin-like growth factor receptor and its ligands as an effective antiangiogenic strategy. Clin Cancer Res. 2013;19:2984–94.
Duxbury MS, Ito H, Zinner MJ, Ashley SW, Whang EE. EphA2: a determinant of malignant cellular behavior and a potential therapeutic target in pancreatic adenocarcinoma. Oncogene. 2004;23:1448–56.
Pasquale EB. Eph-ephrin bidirectional signaling in physiology and disease. Cell. 2008;133:38–52.
Fang WB, Ireton RC, Zhuang G, Takahashi T, Reynolds A, Chen J. Overexpression of EPHA2 receptor destabilizes adherens junctions via a RhoA-dependent mechanism. J Cell Sci. 2008;121:358–68.
Zhou Y, Sakurai H. Emerging and diverse functions of the EphA2 noncanonical pathway in cancer progression. Biol Pharm Bull. 2017;40:1616–24.
Amato KR, Wang S, Hastings AK, Youngblood VM, Santapuram PR, Chen H, et al. Genetic and pharmacologic inhibition of EPHA2 promotes apoptosis in NSCLC. J Clin Invest. 2014;124:2037–49.
Amato KR, Wang S, Tan L, Hastings AK, Song W, Lovly CM, et al. EPHA2 blockade overcomes acquired resistance to EGFR kinase inhibitors in lung cancer. Cancer Res. 2016;76:305–18.
Acknowledgements
This work was supported by the National Natural Science Foundation of China (No. 81821005), and the Strategic Priority Research Program of the Chinese Academy of Sciences (Nos XDA12020345 and XDA12020230).
Author information
Authors and Affiliations
Contributions
MH conceived the whole project; BX and MH initiated the project; MYG and JD provided supervision; YHC performed the research and analyzed the data; HL performed the experiments in the initial stage of the project; XMW, NS, and ST provided technique assistance; MH and YHC wrote the paper. All authors approved the final version of the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Supplementary information
Rights and permissions
About this article
Cite this article
Chen, Yh., Lv, H., Shen, N. et al. EPHA2 feedback activation limits the response to PDEδ inhibition in KRAS-dependent cancer cells. Acta Pharmacol Sin 41, 270–277 (2020). https://doi.org/10.1038/s41401-019-0268-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41401-019-0268-y