Abstract
Insights into symbiosis between eukaryotic hosts and their microbiomes have shifted paradigms on what determines host fitness, ecology, and behavior. Questions remain regarding the roles of host versus environment in shaping microbiomes, and how microbiome composition affects host fitness. Using a model system in ecology, phytoplankton, we tested whether microbiomes are host-specific, confer fitness benefits that are host-specific, and remain conserved in time in their composition and fitness effects. We used an experimental approach in which hosts were cleaned of bacteria and then exposed to bacterial communities from natural environments to permit recruitment of microbiomes. We found that phytoplankton microbiomes consisted of a subset of taxa recruited from these natural environments. Microbiome recruitment was host-specific, with host species explaining more variation in microbiome composition than environment. While microbiome composition shifted and then stabilized over time, host specificity remained for dozens of generations. Microbiomes increased host fitness, but these fitness effects were host-specific for only two of the five species. The shifts in microbiome composition over time amplified fitness benefits to the hosts. Overall, this work solidifies the importance of host factors in shaping microbiomes and elucidates the temporal dynamics of microbiome compositional and fitness effects.
Similar content being viewed by others
Introduction
It is becoming increasingly evident that many microbes that live in close association with eukaryotic hosts alter host fitness [1,2,3], host interactions with other species [4,5,6,7,8,9,10], and even host impacts on ecosystems [11, 12]. Current research efforts often focus on the relative importance of host genetics and environmental factors in shaping microbiome composition and how composition impacts microbiome effects on their hosts [13,14,15,16,17,18,19,20,21]. Here we test the extent to which host factors determine microbiome composition because this is important for predicting how changes in environment will or will not lead to changed microbiomes and, in turn, how these changes could alter host traits.
We use a classic model system in community ecology, eukaryotic phytoplankton, to investigate whether microbiomes are host-specific in composition and have host-specific fitness effects. Phytoplankton have been drawn on for the development of numerous ecological theories [22,23,24], are ecologically important [25, 26], and have external microbiomes that can be readily manipulated in experiments. Phytoplankton cells are surrounded by the phycosphere, a nutrient-rich diffusive boundary layer replete with heterotrophic bacteria that is analogous to the terrestrial plant rhizosphere and phyllosphere [27]. Some phycosphere bacteria attach directly to phytoplankton cells [28], while others rapidly enter and exit the phycosphere in response to gradients in algal exudates [29]. These interactions between phytoplankton and bacteria can range from mutually beneficial, such as an exchange of vitamins and nutrients, to parasitism and competition for limiting inorganic nutrients [27]. Experimental manipulation of these phycospheres has been shown to alter ecological interactions between species of phytoplankton [10]. By regulating phytoplankton population and community dynamics, these interactions between phytoplankton and their phycosphere can ultimately affect global cycling of carbon and nutrients [30].
A range of studies have detected host specificity of microbiomes using mostly comparative methods. Species specificity of microbiomes has been documented in plants [31], invertebrates [32], and vertebrates, including birds [33] and mammals where co-evolution has been detected between bacteria and their hosts [34, 35]. Even within-species genetic variation has been shown to explain a small but significant proportion of variation in the microbiomes of humans [36], maize [37, 38], Arabidopsis [39], and the cyanobacterium Microcystis [40]. Yet, there are notable counter examples that suggest limited genetic regulation of host microbiomes, either within or among species, for example in marine sponges [41]. Hosts may also modify the microbiota in their immediate environment, such that it may be challenging to separate effects of host versus environment [42]. However, when the relative effects of the environment versus host have been explicitly compared, studies have found stronger effects of environment [19, 43, 44]. In phytoplankton, some species-specific associations between bacteria and their algal host have been documented [45,46,47,48,49,50,51,52,53], but most of these studies are limited to one or a small number of phytoplankton species with a limited range of phylogenetic distances between taxa. Further, to our knowledge, only two experimental studies have compared the phycospheres of multiple species of phytoplankton and both have suggested host specificity [46, 47]. However, both studies were limited to a single source community of bacteria, and two host species. Further, when comparing two species, the Eigemann et al. study entailed incubating xenic algae, and so did not control for certain factors shaping community composition, such as priority effects. We aim to advance the field with experimental evidence that tracks how microbiomes assemble by using phytoplankton as an experimentally tractable system in which we can directly compare host and environmental drivers of microbiome composition.
Studies have also examined how a host is affected by the composition of its microbiome. Variation in composition of a host’s microbiome has been linked to variation in host fitness as a function of environmental change, including salt stress in algae [54], drought stress in plants [1], heat stress in corals [17], and temperature and desiccation stress in dung beetles [55]. Similarly, microbiome composition has been shown to affect host resistance to and survival from disease, including fungal pathogen exposure in amphibians and bats [15, 16], and colon cancer and obesity in humans [13, 14]. In addition, the presence of highly heritable bacterial taxa within the human gut microbiome has been shown to correlate with human host metabolism and cause metabolic changes when experimentally introduced into a mouse model [36]. Yet, we do not know the extent to which host specificity of microbiomes affects host fitness, or the persistence of these fitness effects long-term on the host. Our study aims to address these two questions to provide further insights into the fitness effects of host-specific microbiomes.
We used five green algal species varying in phylogenetic relatedness from summed branch lengths of 0.002–0.67 [56] to test whether phycospheres were distinct biomes compared to their surroundings, whether specific bacterial taxa associated with specific algal hosts, and if so, whether these associations were robust to environmentally based variation in bacterial community composition of the source communities. Second, using reciprocal transplant experiments, we tested whether phycosphere bacteria influenced algal growth of their host differently than that of non-hosts, hypothesizing that phycospheres would be most beneficial to their host species. Last, we tested whether phycosphere community composition changed under common garden laboratory conditions and how these changes affected host specificity of phycosphere composition and host fitness effects. These approaches move the field forward from observation inference to experimental evidence of the environmental and genetic drivers of microbiome assembly and host fitness effects.
Materials and methods
Species pool
Phytoplankton cultures were supplied from the University of Texas Culture Collection of Algae (UTEX, USA). We used five species of unicellular eukaryotic green algae: Chlorella sorokiniana, Coelastrum microporum, Monoraphidium minutum, Scenedesmus acuminatus, and Selenastrum capricornutum. Chlorella sorokiniana belongs to the order Chlorellales, while the remaining species belong to the Sphaeropleales. Elsewhere, we isolated heterotrophic bacteria from these long-established cultures (see Table S1 of ref. [10]). While we were only able to isolate 1–2 taxa per algal culture, these included taxa within three orders, Spingomonadales, Rhizobiales, and Actinomycetales, that also occurred in the experimentally assembled microbiomes described below.
Axenification
We rendered algae axenic using ultrasonication to detach bacterial cells and fluorescence-activated cell sorting (Synergy, University of Michigan Biomedical Research Flow Core, USA) onto solid growth media [57]. In brief, we sonicated 20-mL cultures maintained at 1 × 108 algal cells/mL on ice for 30 s at 20 W using a sonic dismembrator (Fisher model 100). We repeated sonication three times separated by 1 min rests. We centrifuged samples at 900 × g for 5 min, discarded supernatant, and resuspended pellets in 20 mL of COMBO [58]. We sorted single cells into wells of a 96-well plate containing COMBO agar with the flow cytometer located inside a biological safety cabinet to maintain sterility. We used a blue excitation laser and PerCP broad-pass emission filter (488-nm excitation, 665/30-nm emission) to sort algal cells. We covered plates with Breathe-Easy polyurethane films to permit gas exchange while maintaining sterility. We incubated plates for 7–12 days and confirmed that colonies were axenic via fluorescent microscopy and by attempted PCR amplification of the bacterial 16S rRNA gene [10].
Study site
We collected pond water on September 14, 2018 from a long-term experimental pond facility at the University of Michigan’s E. S. George Reserve (ESGR) in Pinckney, MI, USA. This facility includes nine ponds each 30 m in diameter and 2 m deep that were constructed in 1988. Naturalization of these ponds was accelerated by adding vegetation from a local pond at the ESGR [59]. These ponds currently harbor diverse communities of invertebrates, amphibians, and submerged and emergent vegetation.
Host-specificity incubation experiment
We first tested whether our experimental approach was effective in allowing for passage of bacteria into axenic algal cultures. We sealed jars of axenic algae with 12–14-kDa dialysis tubing (Spectrum Labs, New Brunswick, NJ, USA) impermeable to bacteria, or 3.0-μm mixed cellulose ester filters (MilliporeSigma) that were permeable to most bacteria. We attached jars of algae to aquarium tanks filled with pond water, submerging the jars so that jars were under at least 5 cm of water. After 7 days of incubation, we assessed growth differences of algae with and without bacterial exposure via chlorophyll-a fluorescence (435/680-nm excitation emission) as a proxy for algal growth with a Biotek plate reader. We also confirmed that 3.0-μm filters retained the inoculated algal species within jars and excluded other species of eukaryotic algae residing in pond water. See results in Fig. S1.
For our main experiment we used three of nine experimental ponds, selecting ponds distanced further apart to maximize environmental variation and their associated bacterial communities. During pond-water collection, we measured temperature, total chlorophyll, and the concentration of green algae, cyanobacteria, diatoms, and cryptophytes using a FluoroProbe (biological | biophysical | engineering Moldaenke). We transferred 40 L of pond water from each pond to a separate tank. Tanks were maintained at 20 °C under fluorescent lighting set to a 16:8-h light:dark cycle. Using stock cultures of axenic algae growing in COMBO, we inoculated algae into 100 mL of COMBO at 10,000,000 cells/mL in 4-oz glass jars sealed with 3.0-μm 90-mm mixed cellulose esters filters. Sealed jars were attached to the walls of each tank with ceramic magnets and epoxy to minimize pond-water debris from settling on and inhibiting passage through 3.0-μm filters. This orientation also minimized shading of algae through opaque 3.0-μm filters. Jars remained submerged in each of the three tanks of pond water for 72 h to permit colonization of the phycosphere.
At the end of 72 h, we characterized phycosphere bacterial communities via 16S rRNA sequencing. We passed 75 mL of algal cultures through an in-line sequential filtration setup of 3.0 and 0.22-μm 25-mm filters. These samples are referred to as either the “>3.0-μm” fraction representative of only particle-associated (i.e., algal-associated) microbes, and the “0.22–3.0-μm” fraction representative of free-living microbes. We transferred 5 mL of algal cultures to flasks containing 100 mL of COMBO. These transfers were likely ecological bottleneck events that sharply reduced population sizes and may have led to local extinctions. Flasks were incubated in a 20 °C Percival growth chamber with 81-μE lighting set to a 16:8-h light:dark cycle. See illustration of experimental design in Fig. S2.
Temporal stability experiment
We assessed temporal changes within algal phycospheres under controlled laboratory conditions for 4 weeks. Starting immediately after the 72-h incubation, we incubated communities in flasks for an additional 7 days. After this 10-day period, we collected particle-associated and free-living bacteria from 50 mL of cultures onto 0.22-μm filters using vacuum manifolds. These samples are referred to as “>0.22-μm” fraction, as they represent the whole microbial fraction. Concurrent with sampling for microbial composition, we started new cultures containing 5 mL of algal cultures into 100 mL of COMBO. Therefore, we surveyed phycosphere bacterial communities via 16S rRNA on days 3, 10, 17, 24, and 31 following the initial reintroduction of bacteria to axenic algae from the initial pond-water. See illustration of experimental design in Fig. S2.
Host specificity of fitness effects experiment
We aimed to test whether fitness benefits conferred by the phycosphere are host-specific. For each of 15 treatments (i.e., 5 algal species × 3 ponds), we diluted to 5000 algal cells/mL and obtained the microbial fraction by gently dislodging microbes from the mucilage of the phycosphere using a sonicator on a 10-mL aliquot of algal culture. We sonicated samples on ice for three runs of 30 s at 20 W with 1 min rests between runs (Fisher model 100). We centrifuged samples at 900 × g for 5 min, and then passed all microbial cells that remained suspended in the supernatant through a 3.0-μm filter to remove any remaining algal cells or larger cellular debris from the microbial filtrate.
We tested growth effects of these 15 bacterial filtrates on each of the five axenic algal species using chlorophyll-a fluorescence. We added axenic algae at 5000 cells/mL in COMBO in 48-well plates, as well as 10 μL of either a bacterial filtrate or sterile COMBO as an axenic control. We ran three replicates of these 75 combinations in a randomized design across six 48-well plates. We had at least one axenic control per species per plate to ensure sufficient controls. We measured fluorescence every second day for 14 days, at which time, most cultures appeared to be at carrying capacity as indicated by asymptotic growth curves.
Microbial sequencing
We thawed 0.22 and 3.0-μm 25-mm filters used to collect culture biomass and incubated them in 30-μL proteinase K, 100 μl of Qiagen ATL tissue lysis buffer, and 300-μL Qiagen AL lysis buffer for 1 h at 56 °C on a rotisserie. We lysed cells by vortexing for 10 min, homogenized lysates with a Qiashredder column, and purified DNA from filtrates using DNeasy Blood and Tissue kits (Qiagen, Hilden, Germany). We generated PCR amplicon of the V4 region of the 16S rRNA gene using 515f/806r primers [60]. DNA amplicon was sequenced on a 2 × 250 Illumina MiSeq v2 run at the University of Michigan Center for Microbial Systems.
Analyses and statistics
We analyzed gene survey data using the mothur v1.34.3 standard operating procedure [61, 62] and assigned taxonomy using the TaxAss pipeline, in which sequences were classified to a database of freshwater taxa [63] and then remaining unclassified sequences were assigned taxonomy using the larger SILVA database [64, 65]. We classified operational taxonomic units (OTUs) from Illumina reads at the 97% similarity level and removed OTUs classified through SILVA as chloroplast and 67 OTUs that were described as “Bacteria_unclassified” and shared close similarity to chloroplast via analyses with NCBI blastn. Removed OTUs contained the following keywords in the NCBI descriptions: eukaryote, chloroplast, mitochondrion, plastid, Scenedesmus, Actuodesmus, Tetradesmus, Mychonastes, Asterarcys, Hariotina, Pectinodesmus, Raphidocelis, Ourococcus, Kirchneriella, Chlorophyceae, Chlorophyta, Chlorella, Kumanoa, Chroomonas, Cryptomonas, Ceramium, Hydropuntia. Samples were rarefied to an even depth of 4900.
We first compared bacterial taxonomic richness, Shannon’s diversity, and Pileou’s evenness over time by comparing initial pond-water, day 3 algal cultures and day 31 algal cultures using the “phyloseq” [66] and “ggplot2” [67] packages in R. We ran an analysis of variance for each of these three metrics with time, filter, day, and host species as fixed effects.
We then evaluated temporal stability of bacterial community composition using a PCoA ordination, a betadispersion test to confirm homogeneity of dispersion among groups, an adonis test on a Bray–Curtis distance matrix with day, pond, and filter as fixed effects, and pairwise post hoc comparisons for day using the pairwise.perm.manova function in the RVAideMemoire package in R. We also analyzed microbial relative abundance data with the “EdgeR” package in R using standard procedures [46, 47, 50, 68]. Briefly, raw relative abundance counts were normalized using the calcNormFactors function, and read counts for OTUs were fit with quasi-likelihood negative binomial generalized log-linear models using the glmQLFit function. We used model contrasts of the initial pond-water bacterial community against algal phycospheres to generate log-fold change values of bacterial OTUs in cultures versus initial pond-water.
We then compared the effects of the pond environment versus algal host on bacterial community composition. To determine how these predictors varied over time, we ran adonis tests with pond and host species as fixed effects at each sampling day on Bray–Curtis distance matrices and pairwise post hoc comparisons for host species. For the pond water and day 3 sampling points for which we collected size-fractionated samples, we ran separate adonis tests on each size fraction (as reported in Table 1). We also report ordinations on day 3 using an abundance-weighted UniFrac distance metric to illustrate phylogenetic divergence of phycosphere communities. For the host specificity of fitness effects experiment, growth over time for each culture was measured as the area under the curve of the fitted logistic equation of chlorophyll-a fluorescence from Day 0 to 14 using the “growthcurver” package in R [69]. We ran this experiment using bacterial filtrates obtained after the 3-day incubation and after the 31-day temporal stability experiment. To compare the magnitude over time of fitness benefits conferred by the local phycosphere relative to axenic cultures, we used a linear mixed-effects model on the raw data points using the lmer function in the ‘lme4’ package in R with a composite term of axenic/xenic status by round as the fixed effect (i.e., (1) axenic in the initial phycosphere, (2) xenic in the initial phycosphere, (3) axenic in the final phycosphere, (4) xenic in the final phycosphere), algal species as a random effect, algal density determined as the area under the growth curve as the dependent variable, and the emmeans function in the “emmeans” packing in R for post hoc pairwise comparisons within the fixed effect. To evaluate effects of phycosphere origin on host fitness, we used linear mixed-effects models on the raw data points with bacterial filtrate (i.e., none, local, or nonlocal) as the fixed effect, algal species as a random effect, algal density determined as the area under the growth curve as the dependent variable, and post hoc pairwise comparisons within the fixed effect. We considered algal-host species as a random effect because we were interested in whether local vs. nonlocal phycospheres had significant fitness effects irrespective of host identity, however we reached the same biological conclusions with an analysis of variance treating both phycosphere origin and algal-host species as fixed effects.
Results
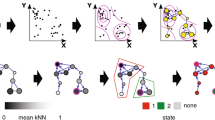
We observed broad trends in phycosphere communities that corresponded to time and algal host. These trends were evident across our dataset, despite each of the three initial source ponds containing distinct characteristics, including bacterial community composition (see Table S1 for pond metadata and Fig. S3 for dissimilarity among pond bacterial communities). By Day 3, a subset of the pond-water bacterial communities had moved across the 3.0-μm barriers and into the jars containing initially axenic algae cultures (Fig. 1A). While the number of OTUs we observed in our rarefied dataset from the ponds was 314 ± 25 OTUs (mean ± SE for the 0.22–3.0-μm fraction), the number of observed OTUs in the algal cultures immediately following the 72-h incubation (Day 3) was less than a third of these taxa (μ = 103 ± 3 SE for the 0.22–3.0-μm fraction; ANOVA: time—F2,136 = 260.6, p < 0.001, Fig. 1A). Observed taxonomic richness continued to decline over time within algal cultures as they were maintained under laboratory conditions (Day 10: μ = 103 ± 4 SE, Day 17: 76 ± 2, Day 24: 67 ± 2, Day 31: 63 ± 2 for the >0.2-μm fractions; Fig. 1A). The observed bacterial diversity within algal cultures was significantly lower compared to pond-water communities (Shannon’s diversity: pond water μ = 3.28 ± 0.10 SE and Day 3 = 2.68 ± 0.03 for the 0.22–3.0-μm fractions, and Day 31 = 2.76 ± 0.04 for the >0.22-μm fraction; Fig. 1B). Observed bacterial community evenness was not significantly different across time (including pond water: μ = 0.57 ± 0.01 SE; algal cultures on Day 3: 0.58 ± 0.005 SE; and algal cultures on Day 31: 0.67 ± 0.0076 SE, respectively; Fig. 1C). Bacterial community composition of the pond water differed sharply from the communities established in the algal cultures by Day 3 when using phylogenetic independent (Figs. 1D and 2) and dependent metrics (Fig. S4). Specifically, Proteobacteria in the 0.22–3.0-μm fractions increased from 32.0% (±1.7 SE) of the pond-water community to 58.6% (± 1.6 SE) on average in algal communities (Fig. 2C), while in the >3.0-μm fractions Proteobacteria increased from 19.2 (±1.5 SE) of the pond-water community to 68.4 (±2.3 SE) on average in the algal communities (Fig. 2D). This increase in phylum abundance was due in part to significant increases among 12 OTUs of α-, 11 OTUs of β-, and 2 OTUs of γ-Proteobacteria that increased relative to pond water in most algal cultures (Fig. 3, differential relative abundance compared to pond water of the 0.22–3.0-μm fraction, all false discovery rate-corrected p values < 0.05 and logFC > 1.0 in at least three of the five algal-host species, Tables S2 and S3). Among OTUs within the α-Proteobacteria that increased in relative abundance were those likely capable of fixing Nitrogen (Table S3: 4 OTUs in Rhizobiales and an OTU in Azospirillales). Bacteriodetes also increased in relative abundance in algal cultures relative to the source pond-water communities, but this result was algal-host-dependent (Fig. 2C, D). Phycosphere communities continued to change over time, with a notable transition between Days 3 and 10 (Fig. 1D), i.e., after the first week of culturing in laboratory media after the first 3 days of incubation in pond-water conditions. For example, selection against Actinobacteria in algal cultures was evident by Day 3 in the 0.22–3.0-μm fraction (logFCDay 3-pond water = −2.1 μ ± 0.17 SE, n = 13 OTUs) and intensified over time (logFCDay 31-pond water = −11.4 ± 0.33 SE, >0.22-μm fraction; Fig. 3). Verrucomicrobiae also declined significantly in relative abundance over time (logFCDay 3-pond water = −4.3 μ ± 0.43 SE, n = 12 OTUs, 0.22–3.0-μm fraction; logFCDay 31-pond water = −11.5 μ ± 0.60 SE, >0.22-μm fraction, Figs. 2C, D and S5). Further, most species of algae hosted a phycosphere community distinct from all other hosts (Fig. 2A: >3.0-μm fraction; Fig. 2B: 0.22–3.0-μm fraction). The only algal hosts that harbored statistically indistinguishable phycosphere communities were the closely related hosts M. minutum and S. capricornutum (adonis post hoc comparison, p > 0.05). The general signature of host specificity was evident despite significant variation attributable to three different bacterial communities across the source ponds (Fig. 2A, B). Host specificity was evident by Day 3 of the experiment (Table 1: 3.0 μm: adonis—F4,44 = 12.5, p < 0.001, R2 = 0.41; 0.22-μm fraction: adonis—F4,44 = 11.6, p < 0.001, R2 = 0.39), and remained a significant predictor of phycosphere composition through the end of the experiment (Table 1; adonis on Day 31: F4,41 = 3.4, p < 0.001, R2 = 0.23). Furthermore, host specificity explained 1.5–3× greater proportion of variation in phycosphere composition compared to the originating pond environment over the course of the experiment (Table 1). However, the magnitude of these host and environmental signatures decreased over the duration of the experiment, as indicated by the decline in R2 values over time and significant day*host and day*pond interactions in an adonis model (all p < 0.001). The host signature can be summarized by a higher relative abundance of Bacteroidetes within cultures of C. sorokiniana and C. microporum, while cultures of M. minutum, S. acuminatus, and S. capricornutum contained higher relative abundances of β-Proteobacteria. In addition, phylogenetic similarity between host pairs was positively correlated with similarity of the recruited phycosphere communities, where phylogenetic similarity is described in Table S4 and phycosphere similarity was calculated as the average Bray–Curtis distance among Day 3, >3.0-μm samples that are shown in Fig. 2A (R2 = 0.69, p = 0.003) (Fig. S6). The correlation between phylogenetic similarity and phycosphere similarity using the 0.22–3.0-μm samples shown in Fig. 2B was insignificant (R2 = 0.24, p = 0.15, slope = 0.10) (Fig. S6). Considering that phycosphere bacterial communities were a subset of whole-pond communities and that these phycosphere communities continued to change over time, we asked whether these communities were (1) beneficial to their algal host, and, if so, (2) whether the magnitude of this benefit changed over time. We separated phycosphere bacterial communities from Days 3 and 31 of each algal species and reintroduced the bacterial communities to axenic cultures of the same algal species. Phycosphere bacterial communities significantly increased algal population density relative to axenic algal cultures (Fig. 4; linear mixed-effects model p < 0.001, Table S5). Furthermore, while axenic algal controls reached similar population densities in the two sets of experiments using initial and final phycospheres, algae inoculated with final phycospheres reach significantly greater densities than algae inoculated with initial phycospheres (Fig. 4). Considering the significant host-specific signature, we tested whether the “local” phycosphere community benefited the host to a greater extent than the phycospheres of other algal species. We obtained phycospheres from Day 3 cultures, as well as Day 31 cultures, by which time community composition, richness, evenness, and magnitude of the host-specific signature had changed significantly (Fig. 1 and Table 1). These phycospheres associated with different algal species were notably distinct from one another: percent overlap in total bacterial OTUs between host pairs was 49.7–63.9% shared OTUs during Day 3 and 32.0–66.4% shared OTUs during Day 31. Despite these differences among phycosphere communities, we found consistent results at both time points: nonlocal phycospheres tended to have similar effects on algal population density relative to the phycospheres derived from the same host algal species (Fig. 5; linear mixed-effects models: Day 3-marginal R2 = 0.64, p < 0.0001, Day 31-marginal R2 = 0.68, p < 0.0001; Tukey’s post hoc comparisons significant with p < 0.05: axenic < local = nonlocal). While there were no significant trends, C. sorokiniana appeared to benefit more from nonlocal Day 3 phycospheres than local Day 3 phycospheres (post hoc p = 0.057), while S. acuminatus appeared to benefit more from local Day 31 phycosphere than nonlocal Day 31 phycospheres (post hoc p = 0.095). When testing for algal growth differences, we take into account the full growth trajectory by using the integrated area under the growth curve.
These bacterial communities now associated with algae were A lower in taxonomic richness, B lower in Shannon’s diversity, but C similar in Pielou’s evenness. As algal cultures were maintained under laboratory conditions, richness and diversity continued to decline whereas evenness remained similar. D Bacterial community composition described using a Bray–Curtis distance metric differed most between the community inhabiting pond water versus the subset of the community that migrated into the jars containing the initially axenic algal cultures, as well as between culture jars (i.e., Day 3) and culture flasks (i.e., Days 10–31). Bacterial composition differed significantly between all days, except Day 24, which did not differ significantly from Day 17 or Day 31 via pairwise post hoc tests where p < 0.05.
Host specificity was evident in A the >3.0-μm fraction and B the 0.22–3.0-μm fraction. Host specificity in the C >3.0-μm fraction and D 0.22–3.0-μm fraction could be explained in part by the prevalence of Bacteriodetes in cultures of C. microporum and C. sorokiniana versus Proteobacteria in cultures of the remaining three species. Bacterial composition associated with each algal host differed significantly for all host pairwise comparisons for both fractions, with the exception of M. minutum versus S. capricornutum, which did not harbor significantly different bacterial communities from each other for either fraction, via pairwise post hoc tests where p < 0.05. See Fig. S4 for analyses using the phylogenetic-dependent UniFrac distance metric.
Among OTUs comprising at least 0.1% of all algal culture reads, OTUs of Proteobacteria and Bacteroidetes showed variable results depending on taxon, whereas OTUs of Actinobacteria showed consistent patterns of underrepresentation. See Fig. S5 for these three phyla with no relative abundance threshold. Included in analyses were 0.22–3.0-μm samples for pond water and Day 3, and >0.22-μm samples for Days 10–31. OTU lines with fewer than six data points were absent from the normalized datasets at those time points.
Phycospheres were reintroduced to the algal species of origin to generate “xenic” strains. A Initial phycospheres originated from algal cultures obtained on Day 3, which was 72 h following exposure of axenic algae to pond water, and B final phycospheres originated from Day 31 of this experiment following 4 weeks of maintenance under controlled laboratory conditions. See Figs. 1 and 3 for further description of bacterial community composition changes over this timespan. Total area under the curve of fluorescence-based estimates of algal population density was compared across the first 14 days of growth. Tests of initial and final phycospheres inherently required two rounds of experiments completed at different times. Linear mixed-effects model was significant (p < 0.0001) with shared lettering indicating those treatments that do not differ significantly according to post hoc comparisons (p > 0.05).
Phycospheres reintroduced to the algal species of origin were referred to as “local,” and all others were referred to as “nonlocal.” A When phycospheres were reintroduced immediately after the assembly event, algal densities reached significantly higher levels when xenic. B When phycospheres were isolated from cultures 4 weeks after the assembly event and reintroduced to axenic algae, algal densities still reached significantly higher levels when xenic than axenic. For both initial and final phycosphere: linear mixed-effects models p < 0.0001 with post hoc comparisons indicating: axenic < local = nonlocal.
Discussion
Our results demonstrate that algal phycospheres are microenvironments that select for and against specific bacterial taxa, shaping communities of associated bacteria that are appreciably distinct from surrounding freshwater environments. With increasing evidence regarding the role of host microbiomes on host fitness and ecological interactions, identifying the relative importance of host versus environment on microbiome composition and temporal stability is an important factor to predicting variability in fitness traits and ecological outcomes. Among the five phytoplankton species we included in our study, host identity was the main driver of phycosphere composition, which is in agreement with and expands upon previous studies [46, 47, 50]. While the host-specific signature remained significant throughout the course of the experiment, effects of phycospheres on host fitness were not significantly improved based on host-specific composition. Yet, effects on host fitness were improved as phycosphere communities changed over time, presumably by ecological selection favoring more beneficial populations. Our results further our understanding of the forces shaping community assembly and stability of host microbiomes and the link between microbiome composition and host fitness.
Our study shows that the algal phycosphere is recruited in a host-specific manner that persists across environmental variation in the source communities of bacteria. The four processes that could contribute to community assembly of the phycosphere include dispersal, diversification, drift, and selection [70]. Our experimental approach was designed to facilitate equal opportunity for the dispersal of bacterial taxa to any of the algal-host species. Further, the 4-week time frame of our experiment minimizes community differences due to diversification. Drift likely contributed to loss of low-abundance bacterial taxa [71], in part through the weekly bottleneck effect due to transferring a subset of each culture to fresh media. However, ecological selection likely predominated through (1) chemotaxis of beneficial, commensal, and pathogenic bacteria in response to species-specific algal exudates [72, 73], and (2) subsequent selection by the algal host in promoting beneficial bacteria and warding off harmful bacteria, such as through mechanisms that have been documented in plants [74, 75]. Bacteria may also play an active role in selection because while bacterial growth is dependent on the substrates made available by the host, bacteria could also stimulate the host to alter algal physiology in ways that optimize for bacterial growth. Our experimental design precludes us from differentiating the role of algae versus bacteria in driving these host-specific associations. Other studies have seen similar signals of specificity, but these studies were all limited in their approach, including: comparison of only distantly related, xenic hosts [46], using only two hosts species [47], or drawing comparisons from surveys of associated microbiomes rather than experimental tests [45, 48, 51,52,53, 76, 77]. In addition, among our limited dataset of five phytoplankton hosts, we found that host species that were more closely related phylogenetically harbored significantly more similar phycospheres relative to distantly related hosts. An expanded study using more species of phytoplankton could further clarify whether this host-specific signal is taxonomically conserved, such as what has been shown in hominids [34]. Further, we found that the host-specific signature was detectable after a duration spanning an estimated 30–80 host generations (based on typical doubling times of 9–24 h for the algal species used) [78, 79]. However, the proportion of variation in microbial community composition explained by host species declined significantly over the course of the experiment, which is similar to results from a study that found genotypic signal in the tomato plant microbiome persisted but declined in relative importance over time [80].
Relative to the surrounding pond-water environments, algal cultures in our experiments were enriched with Bacteroidetes and Proteobacteria. A previous experimental recruitment of bacteria from lake water into xenic cultures of two phytoplankton species, the green alga Desmodesmus armatus and diatom Stephanodiscus minutulus, found that Proteobacteria were most common [46]. Both phyla frequently dominate the bacterial communities inhabiting freshwater particles, which are often composed of algal aggregates and colonies [81, 82]. These phyla have also been found residing in abundance within the phycospheres of green algal cultures derived from freshwater environments [83]. Proteobacteria, particularly β-Proteobacteria, have been shown to thrive in environments rich in algal exudates and in culture-based experiments that have used algal exudates as the sole source of carbon [84, 85]. Further, apparent selection against Actinobacteria is in-line with their known nature as free-living bacteria that do not form strong associations with algae [86,87,88]. We also observed ongoing selection against Verrucomicrobia, which contrasts with existing literature that suggests this phylum is generally associated with nutrient-rich environments such as algal phycospheres [86, 87], but is in-line with a recent study indicating that the Opitutae class within Verrucomicrobia tends to be free-living [89].
Observed bacterial richness in algal cultures declined over the course of the experiment, suggesting either loss of rare species through ecological drift and bottlenecks imposed through weekly culture transfers, or selection against certain bacterial taxa either due to inadequate abiotic laboratory conditions (i.e., light, media, temperature), reduced environmental variation due to controlled laboratory conditions, or continued selection by their algal hosts. In the context of this general decline in species richness, certain increases in the relative abundance of taxa are notable. Specifically, our results suggest that the β-Proteobacteria clade Bet1-A plays an important role in algal phycospheres. The Bet1-A-classified OTU was one of the most abundant OTUs within the final phycospheres of each of the five algal species. The best characterized taxon in Bet1-A is the ubiquitous and abundant genus, Limnohabitans. Limnohabitans appear to benefit from association with algae based on its enrichment during phytoplankton blooms [90, 91], contributing 7–45% of all bacteria in cultures of algal Cryptophyta and Chlorophyta [92], and its enrichment when grown on algal extracellular exudates [93].
Changes in phycosphere composition over the duration of the study help clarify how composition affects host fitness. Phycosphere shifts over the course of the experiment were in a direction that proved beneficial to algal hosts. The increased relative abundance from Day 3 to Day 31 of several taxa in the Rhizobiales, likely nitrogen-fixers [94], could in part explain this result (see Table S3; especially OTU #10 which increased on average from 2.1% of the pond-water community to as much as an average of 11.1% of the final phycosphere communities of M. minutum). Although algae were cultured in a nitrate-rich medium, ammonium provided by N-fixers could further promote algal growth. Further, considering that these final phycospheres were also lower in bacterial richness and showed a significantly weaker signature of host specificity, a sizable proportion of the host-specific signature may be due to host-specific pathogenic bacteria rather than beneficial taxa. Loss of these pathogenic bacteria over time could explain improved algal growth when inoculated with the final phycosphere communities. While this remains speculative, some support for suppression of pathogens on Day 31 relative to Day 3 can be seen in the tendency for higher population densities for C. sorokiniana when grown with the phycosphere communities of any other algal species than with the reintroduction of their own phycosphere on Day 3 but not on Day 31. For example, the high incidence in initial cultures of C. sorokiniana of the common plant pathogen, Burkholderiales, compared to other initial cultures and the final cultures could explain this pattern (see OUT #4, Table S2) [95]. In addition, the lack of clear evidence of host-specific fitness effects could have been dependent on environmental conditions. For example, host-specific gut microbiomes in beetles have been shown to be particularly beneficial when ecologically relevant stressors were applied, including temperature and drought stress [55]. Further studies in phytoplankton could investigate whether any host-specific benefits of phycospheres become more evident under environmental stress, such as depleted nutrient conditions instead of the nutrient-rich media that we used in our study. A second explanation for the lack of host-specific fitness effects is that significant host specificity of algal phycospheres in terms of taxonomic bacterial community composition may not translate into functional differences. Functional convergence despite taxonomic divergence of microbiomes has been noted in other systems, including phycospheres of cyanobacteria [40]. In addition, although our microbiome transplants introduced quite different microbial taxa compared to the native community, we do not know whether these communities survived in their entirety. Comparing the inoculated community with that which becomes established with the non-native host, as well as multiple sequential transfers of a microbiome (i.e., from the native to non-native host, and then back to native host), would be valuable future directions for clarifying the importance of host identity in maintaining a phycosphere that confers fitness benefits. In addition, we do not know whether bacterial taxa established themselves within the algal phycosphere of non-native hosts. Co-occurrence, rather than a true microbiome, may still benefit the host, such as through degradation of toxic byproducts of photosynthesis, but spatial proximity within the phycosphere might be necessary for certain functions. Regardless of whether these transplanted microbiomes fully integrated into the microbiomes of non-native hosts, we see that these non-native microbiomes significantly increased host fitness relative to axenic hosts.
Our study has several important limitations. First, with only five species of phytoplankton hosts and no genetic variation within host species, we are limited from drawing strong quantitative conclusions regarding whether genetic distance among hosts is correlated with phycosphere community dissimilarity. Our study suggests a positive correlation, in-line with other studies that have detected distinct microbiomes associated with different host species [46, 76], as well as with genotypes within species [37, 39, 40]. Second, our experimental design has important limitations that differ from environmental conditions. For example, we used one exposure event rather than continued exposure to environmental bacteria as would occur in nature. Therefore, phycosphere communities could not gain taxa beyond those obtained during the 72-h incubation event. Community composition could continue to evolve in terms of relative abundance and local extinction, but taxa that might have been advantageous during late-stage succession could not be recruited to the community. Our use of closed experimental systems and the fact that most environmental bacteria have generation times under 1 day, might explain why temporal shifts, although significant, are of lesser magnitude after Day 10 (Fig. 1D). Despite this limitation of a closed system, it is important to note that the microbiome shifts that did occur were sufficient to confer significantly increased host fitness. Considering the decline in the host-specific signature over time, and yet fitness increase, these results suggest that closed systems facilitated the selection of a more beneficial phycosphere than might have been attainable with an influx of additional, perhaps pathogenic bacteria. A modified experimental design that tracks phycosphere changes in parallel between open and closed systems that either do/do not maintain exposure to the environment might clarify the relative roles of ecological drift versus selection. An additional limitation of the experimental design is that sealing axenic algal cultures with filters might also select against larger bacterial taxa and those exclusively attached to larger particles. For example, in the Great Lakes and inland lakes in Michigan, the Cyanobacteria, Planctomycetes, Bacteroidetes, and Verrucomicrobia phyla have been found more commonly associated with particles (>3.0-µm fraction), while the β-Proteobacteria and Actinobacteria have been found more commonly associated with the free-living lake water fraction (0.22–3.0-µm fraction) [88, 89, 96]. Any buildup of microbial biofilm on the filters could have further reduced passage of bacteria into the algal cultures. Further, we cannot conclude from our experimental design what mechanism lead to host-specific composition. For example, we cannot disentangle whether bacteria were selectively attracted via chemotaxis in response to algal metabolites into the different species of algal cultures, or whether similar bacterial communities entered all cultures and then, through differential selection, resulted in divergent communities. Furthermore, although we chose common host species, we did not confirm that each species occurred within each pond, thus an appropriate seed community of host-associated bacteria may not have existed for each species in each pond. Last, our study was limited in geographic and environmental range to those bacterial communities occurring concurrently within ponds of a 0.013-km2 area. Two of the three ponds used within this area harbored quite similar bacterial communities, and so the total variance among pools may have started off quite low. Future experiments rectifying these issues by using field-based experiments spanning a wider range of temperature and nutrients could help elucidate whether the trends we see in controlled laboratory conditions hold under more variable conditions in situ.
Despite these limitations, we can draw several conclusions from this work. Host microbiomes located external to their phytoplankton hosts are indeed composed of distinct biomes relative to their surroundings, in-line with extensive previous research on the phycosphere [27]. Certain aspects of phycosphere composition were consistent across multiple host species and multiple originating environments, including enrichment of α, β, and γ-Proteobacteria and depletion of Actinobacteria and Verrucomicrobia. Further, our results indicate that the phytoplankton phycosphere is determined more so by host biology than variation in environment. In turn, although a wider range of environmental conditions needs to be tested, these results indicate that traits conferred to a host by its phycosphere might be moderately deterministic in that they broadly benefit host fitness. Yet, phycospheres are not host-specific in these fitness benefits, at least under the scenarios tested in our study. Nonetheless, as phycospheres further changed and then stabilized in composition during laboratory culture, fitness effects on their hosts increased, indicating a dependence of fitness effects on phycosphere community composition. This indicates the value of microbiome characterization in understanding observed variation in host fitness across space and time.
Expanding our work beyond phytoplankton to include implications more broadly for host microbiome systems, this work adds to a range of studies suggesting host taxonomy filters microbial communities from the surrounding environment, resulting in microbiomes that are species- and genotype-specific [31,32,33, 35, 80]. However, it is also important to note that several studies directly comparing environment versus host have found stronger effects of environment, particularly in the vertebrate guts of carnivores, herbivores, and omnivores (including humans) [19, 43, 44]. Differing conclusions across this range of studies may in part be due to a focus on autotrophic versus heterotrophic hosts. Autotrophs recruit a microbiome by exudates that are produced as a byproduct of the host’s physiology (i.e., photosynthesis), while the most commonly studied microbiome of heterotrophs, the gut microbiome, is recruited primarily based on host diet, rather than host-dependent metabolites. Selection imposed by autotrophic hosts on their microbiome may be more persistent and therefore a host signature more evident. This could explain why the environment seems to be a more important driver than host taxonomy among many heterotrophic hosts. Experimental studies incorporating autotrophic and heterotrophic model organisms could clarify the role of environment versus host in shaping host microbiome composition and microbiome-based fitness effects.
References
Lau JA, Lennon JT. Rapid responses of soil microorganisms improve plant fitness in novel environments. Proc Natl Acad Sci. 2012;109:14058.
Rosshart SP, Vassallo BG, Angeletti D, Hutchinson DS, Morgan AP, Takeda K, et al. Wild mouse gut microbiota promotes host fitness and improves disease resistance. Cell. 2017;171:1015–1028.e13.
Lynch J, Hsiao E. Microbiomes as sources of emergent host phenotypes. Science. 2019;365:1405–9.
Holland MA, Polacco JC. PPFMs and other covert contaminants: Is there more to plant physiology than just plant? Annu Rev Plant Physiol Plant Mol Biol. 1994;45:197–209.
Cho I, Blaser MJ. The human microbiome: at the interface of health and disease. Nat Rev Genet. 2012;13:260.
Coleman-Derr D, Tringe SG. Building the crops of tomorrow: advantages of symbiont-based approaches to improving abiotic stress tolerance. Front Microbiol. 2014;5:283.
Sampson TR, Mazmanian SK. Control of brain development, funtion, and behavior by the microbiome. Cell Host Microbe. 2015;17:565–76.
Siefert A, Zillig KW, Friesen ML, Strauss SY. Soil microbial communities alter conspecific and congeneric competition consistent with patterns of field coexistence in three Trifolium congeners. J Ecol. 2018;106:1876–91.
Siefert A, Zillig KW, Friesen ML, Strauss SY. Mutualists stabilize the coexistence of congeneric legumes. Am Naturalist. 2019;193:200–12.
Jackrel SL, Schmidt KC, Cardinale BJ, Denef VJ. Microbiomes reduce their host’s sensitivity to interspecific interactions. mBio. 2020;11:1–11.
Paustian K, Lehmann J, Ogle S, Reay D, Robertson GP, Smith P. Climate-smart soils. Nature. 2016;532:49.
Shi W, Moon CD, Leahy SC, Kang D, Froula J, Kittelmann S, et al. Methane yield phenotypes linked to differential gene expression in the sheep rumen microbiome. Genome Res. 2014;24:1517–25.
Ridaura VK, Faith JJ, Rey FE, Cheng J, Duncan AE, Kau AL, et al. Gut microbiota from twins discordant for obesity modulate metabolism in mice. Science. 2013;341:1241214.
Zackular JP, Baxter NT, Iverson KD, Sadler WD, Petrosino JF, Chen GY, et al. The gut microbiome modulates colon tumorigenesis. mBio. 2013;4:e00692–13.
Kueneman JG, Woodhams DC, Harris R, Archer HM, Knight R, McKenzie VJ. Probiotic treatment restores protection against lethal fungal infection lost during amphibian captivity. Proc R Soc B: Biol Sci. 2016;283:20161553.
Cheng TL, Mayberry H, McGuire LP, Hoyt JR, Langwig KE, Nguyen H, et al. Efficacy of a probiotic bacterium to treat bats affected by the disease white‐nose syndrome. J Appl Ecol. 2017;54:701–8.
Ziegler M, Seneca FO, Yum LK, Palumbi SR, Voolstra CR. Bacterial community dynamics are linked to patterns of coral heat tolerance. Nat Commun. 2017;8:14213.
Björk JR, Hui FK, O’Hara RB, Montoya JM. Uncovering the drivers of host‐associated microbiota with joint species distribution modelling. Mol Ecol. 2018;27:2714–24.
Youngblut ND, Reischer GH, Walters W, Schuster N, Walzer C, Stalder G, et al. Host diet and evolutionary history explain different aspects of gut microbiome diversity among vertebrate clades. Nat Commun. 2019;10:2200.
David AS, Quintana-Ascencio PF, Menges ES, Thapa-Magar KB, Afkhami ME, Searcy CA. Soil microbiomes underlie population persistence of an endangered plant species. Am Naturalist. 2019;194:488–94.
David AS, Thapa‐Magar KB, Afkhami ME. Microbial mitigation–exacerbation continuum: a novel framework for microbiome effects on hosts in the face of stress. Ecology. 2018;99:517–23.
Chesson P. Mechanisms of maintenance of species diversity. Annu Rev Ecol Syst. 2000;31:343–66.
Hutchinson GE. The paradox of the plankton. Am Naturalist. 1961;95:137–45.
Litchman E, Klausmeier CA, Schofield OM, Falkowski PG. The role of functional traits and trade-offs in structuring phytoplankton communities: scaling from cellular to ecosystem level. Ecol Lett. 2007;10:1170–81.
Falkowski PG, Katz ME, Knoll AH, Quigg A, Raven JA, Schofield O, et al. The evolution of modern eukaryotic phytoplankton. Science. 2004;305:354.
Field CB, Behrenfeld MJ, Randerson JT, Falkowski P. Primary production of the biosphere: Integrating terrestrial and oceanic components. Science. 1998;281:237.
Seymour JR, Amin SA, Raina J-B, Stocker R. Zooming in on the phycosphere: the ecological interface for phytoplankton–bacteria relationships. Nat Microbiol. 2017;2:17065.
Kaczmarska I, Ehrman JM, Bates SS, Green DH, Léger C, Harris J. Diversity and distribution of epibiotic bacteria on Pseudo-nitzschia multiseries (Bacillariophyceae) in culture, and comparison with those on diatoms in native seawater. Harmful Algae. 2005;4:725–41.
Smriga S, Fernandez VI, Mitchell JG, Stocker R. Chemotaxis toward phytoplankton drives organic matter partitioning among marine bacteria. Proc Natl Acad Sci. 2016;113:1576–81.
Cirri E, Pohnert G. Algae–bacteria interactions that balance the planktonic microbiome. N Phytologist. 2019;223:100–6.
Kembel SW, O’Connor TK, Arnold HK, Hubbell SP, Wright SJ, Green JL. Relationships between phyllosphere bacterial communities and plant functional traits in a neotropical forest. Proc Natl Acad Sci. 2014;111:13715–20.
Reveillaud J, Maignien L, Eren AM, Huber JA, Apprill A, Sogin ML, et al. Host-specificity among abundant and rare taxa in the sponge microbiome. ISME J. 2014;8:1198–209.
Hird SM, Sánchez C, Carstens BC, Brumfield RT. Comparative gut microbiota of 59 neotropical bird species. Front Microbiol. 2015;6:1403.
Ochman H, Worobey M, Kuo C-H, Ndjango J-BN, Peeters M, Hahn BH, et al. Evolutionary relationships of wild hominids recapitulated by gut microbial communities. PLoS Biol. 2010;8:e1000546.
Moeller AH, Caro-Quintero A, Mjungu D, Georgiev AV, Lonsdorf EV, Muller MN, et al. Cospeciation of gut microbiota with hominids. Science. 2016;353:380.
Goodrich JK, Waters JL, Poole AC, Sutter JL, Koren O, Blekhman R, et al. Human genetics shape the gut microbiome. Cell. 2014;159:789–99.
Peiffer JA, Spor A, Koren O, Jin Z, Tringe SG, Dangl JL, et al. Diversity and heritability of the maize rhizosphere microbiome under field conditions. Proc Natl Acad Sci. 2013;110:6548.
Walters WA, Jin Z, Youngblut N, Wallace JG, Sutter J, Zhang W, et al. Large-scale replicated field study of maize rhizosphere identifies heritable microbes. Proc Natl Acad Sci. 2018;115:7368–73.
Lundberg DS, Lebeis SL, Paredes SH, Yourstone S, Gehring J, Malfatti S, et al. Defining the core Arabidopsis thaliana root microbiome. Nature. 2012;488:86.
Jackrel SL, White JD, Evans JT, Buffin K, Hayden K, Sarnelle O, et al. Genome evolution and host‐microbiome shifts correspond with intraspecific niche divergence within harmful algal bloom‐forming Microcystis aeruginosa. Mol Ecol. 2019;28:3994–4011.
Björk JR, Diéz-Vives C, Astudillo-García C, Archie EA, Montoya JM. Vertical transmission of sponge microbiota is inconsistent and unfaithful. Nat Ecol Evol. 2019;3:1172–83.
Wong AC-N, Luo Y, Jing X, Franzenburg S, Bost A, Douglas AE. The host as the driver of the microbiota in the gut and external environment of Drosophila melanogaster. Appl Environ Microbiol. 2015;81:6232–40.
Hird SM, Carstens BC, Cardiff SW, Dittmann DL, Brumfield RT. Sampling locality is more detectable than taxonomy or ecology in the gut microbiota of the brood-parasitic Brown-headed Cowbird (Molothrus ater). PeerJ. 2014;2:e321.
Rothschild D, Weissbrod O, Barkan E, Kurilshikov A, Korem T, Zeevi D, et al. Environment dominates over host genetics in shaping human gut microbiota. Nature. 2018;555:210–5.
Behringer G, Ochsenkühn MA, Fei C, Fanning J, Koester JA, Amin SA. Bacterial communities of diatoms display strong conservation across strains and time. Front Microbiol. 2018;9:659.
Eigemann F, Hilt S, Salka I, Grossart H-P. Bacterial community composition associated with freshwater algae: Species specificity vs. dependency on environmental conditions and source community. FEMS Microbiol Ecol. 2013;83:650–63.
Grossart HP, Levold F, Allgaier M, Simon M, Brinkhoff T. Marine diatom species harbour distinct bacterial communities. Environ Microbiol. 2005;7:860–73.
Hold GL, Smith EA, Rappë MS, Maas EW, Moore ER, Stroempl C, et al. Characterisation of bacterial communities associated with toxic and non-toxic dinoflagellates: Alexandrium spp. and Scrippsiella trochoidea. FEMS Microbiol Ecol. 2001;37:161–73.
Krohn-Molt I, Alawi M, Förstner KU, Wiegandt A, Burkhardt L, Indenbirken D, et al. Insights into microalga and bacteria interactions of selected phycosphere biofilms using metagenomic, transcriptomic, and proteomic approaches. Front Microbiol. 2017;8:1941.
Mönnich J, Tebben J, Bergemann J, Case R, Wohlrab S, Harder T. Niche-based assembly of bacterial consortia on the diatom Thalassiosira rotula is stable and reproducible. The ISME Journal. 2020.
Sapp M, Schwaderer AS, Wiltshire KH, Hoppe H-G, Gerdts G, Wichels A. Species-specific bacterial communities in the phycosphere of microalgae? Microb Ecol. 2007;53:683–99.
Schäfer H, Abbas B, Witte H, Muyzer G. Genetic diversity of ‘satellite’ bacteria present in cultures of marine diatoms. FEMS Microbiol Ecol. 2002;42:25–35.
Sison-Mangus MP, Jiang S, Tran KN, Kudela RM. Host-specific adaptation governs the interaction of the marine diatom, Pseudo-nitzschia and their microbiota. ISME J. 2014;8:63.
Dittami SM, Duboscq-Bidot L, Perennou M, Gobet A, Corre E, Boyen C, et al. Host–microbe interactions as a driver of acclimation to salinity gradients in brown algal cultures. ISME J. 2016;10:51–63.
Schwab DB, Riggs HE, Newton ILG, Moczek AP. Developmental and ecological benefits of the maternally transmitted microbiota in a dung beetle. Am Naturalist. 2016;188:679–92.
Alexandrou MA, Cardinale BJ, Hall JD, Delwiche CF, Fritschie K, Narwani A, et al. Evolutionary relatedness does not predict competition and co-occurrence in natural or experimental communities of green algae. Proc R Soc B: Biol Sci. 2015;282:20141745.
Cho DH, Ramanan R, Kim BH, Lee J, Kim S, Yoo C, et al. Novel approach for the development of axenic microalgal cultures from environmental samples. J Phycol. 2013;49:802–10.
Kilham SS, Kreeger DA, Lynn SG, Goulden CE, Herrera L. COMBO: a defined freshwater culture medium for algae and zooplankton. Hydrobiologia. 1998;377:147–59.
Werner EE, McPeek MA. Direct and indirect effects of predators on two anuran species along an environmental gradient. Ecology. 1994;75:1368–82.
Bergmann GT, Bates ST, Eilers KG, Lauber CL, Caporaso JG, Walters WA, et al. The under-recognized dominance of Verrucomicrobia in soil bacterial communities. Soil Biol Biochem. 2011;43:1450–5.
Kozich JJ, Westcott SL, Baxter NT, Highlander SK, Schloss PD. Development of a dual-index sequencing strategy and curation pipeline for analyzing amplicon sequence data on the MiSeq Illumina sequencing platform. Appl Environ Microbiol. 2013;79:5112–20.
Schloss PD, Westcott SL, Ryabin T, Hall JR, Hartmann M, Hollister EB, et al. Introducing mothur: open-source, platform-independent, community-supported software for describing and comparing microbial communities. Appl Environ Microbiol. 2009;75:7537–41.
Newton RJ, Jones SE, Eiler A, McMahon KD, Bertilsson S. A guide to the natural history of freshwater lake bacteria. Microbiol Mol Biol Rev. 2011;75:14–49.
Quast C, Pruesse E, Yilmaz P, Gerken J, Schweer T, Yarza P, et al. The SILVA ribosomal RNA gene database project: improved data processing and web-based tools. Nucleic Acids Res. 2012;41:D590–6.
Wang Q, Garrity GM, Tiedje JM, Cole JR. Naïve bayesian classifier for rapid assignment of rRNA sequences into the new bacterial taxonomy. Appl Environ Microbiol. 2007;73:5261–7.
McMurdie PJ, Holmes S. phyloseq: an R package for reproducible interactive analysis and graphics of microbiome census data. PloS ONE. 2013;8:e61217.
Wickham H. ggplot2: elegant graphics for data analysis. Springer-Verlag New York; 2016.
Robinson MD, McCarthy DJ, Smyth GK. edgeR: a bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics. 2009;26:139–40.
Sprouffske K, Wagner A. Growthcurver: an R package for obtaining interpretable metrics from microbial growth curves. BMC Bioinform. 2016;17:172.
Vellend M The theory of ecological communities. Princeton University Press. 2016.
Vellend BM. Conceptual synthesis in community ecology. Q Rev Biol. 2010;85:183–206.
Seyedsayamdost MR, Case RJ, Kolter R, Clardy J. The Jekyll-and-Hyde chemistry of Phaeobacter gallaeciensis. Nat Chem. 2011;3:331–5.
Stocker R, Seymour JR. Ecology and physics of bacterial chemotaxis in the ocean. Microbiol Mol Biol Rev. 2012;76:792–812.
Amin SA, Parker MS, Armbrust EV. Interactions between diatoms and bacteria. Microbiol Mol Biol Rev. 2012;76:667–84.
Mathesius U, Mulders S, Gao M, Teplitski M, Caetano-Anollés G, Rolfe BG, et al. Extensive and specific responses of a eukaryote to bacterial quorum-sensing signals. Proc Natl Acad Sci. 2003;100:1444–9.
Jasti S, Sieracki ME, Poulton NJ, Giewat MW, Rooney-Varga JN. Phylogenetic diversity and specificity of bacteria closely associated with Alexandrium spp. and other phytoplankton. Appl Environ Microbiol. 2005;71:3483–94.
Sapp M, Wichels A, Gerdts G. Impacts of cultivation of marine diatoms on the associated bacterial community. Appl Environ Microbiol. 2007;73:3117–20.
Chisti Y. Biodiesel from microalgae. Biotechnol Adv. 2007;25:294–306.
Rosenberg JN, Kobayashi N, Barnes A, Noel EA, Betenbaugh MJ, Oyler GA. Comparative analyses of three Chlorella species in response to light and sugar reveal distinctive lipid accumulation patterns in the microalga C. sorokiniana. PloS ONE. 2014;9:e92460.
Morella NM, Weng FC-H, Joubert PM, Metcalf CJE, Lindow S, Koskella B. Successive passaging of a plant-associated microbiome reveals robust habitat and host genotype-dependent selection. Proc Natl Acad Sci. 2020;117:1148–59.
Weiss P, Schweitzer B, Amann R, Simon M. Identification in situ and dynamics of bacteria on limnetic organic aggregates (lake snow). Appl Environ Microbiol. 1996;62:1998–2005.
Lemarchand C, Jardillier L, Carrias J-F, Richardot M, Debroas D, Sime-Ngando T, et al. Community composition and activity of prokaryotes associated to detrital particles in two contrasting lake ecosystems. FEMS Microbiol Ecol. 2006;57:442–51.
Ramanan R, Kang Z, Kim B-H, Cho D-H, Jin L, Oh H-M, et al. Phycosphere bacterial diversity in green algae reveals an apparent similarity across habitats. Algal Res. 2015;8:140–4.
Pernthaler J, Posch T, S̆imek K, Vrba J, Pernthaler A, Glöckner FO, et al. Predator-specific enrichment of Actinobacteria from a cosmopolitan freshwater clade in mixed continuous culture. Appl Environ Microbiol. 2001;67:2145–55.
Šimek K, Kasalický V, Jezbera J, Jezberová J, Hejzlar J, Hahn MW. Broad habitat range of the phylogenetically narrow R-BT065 cluster, representing a core group of the Betaproteobacterial genus Limnohabitans. Appl Environ Microbiol. 2010;76:631–9.
Haukka K, Kolmonen E, Hyder R, Hietala J, Vakkilainen K, Kairesalo T, et al. Effect of nutrient loading on bacterioplankton community composition in lake mesocosms. Microb Ecol. 2006;51:137–46.
Kolmonen E, Sivonen K, Rapala J, Haukka K. Diversity of cyanobacteria and heterotrophic bacteria in cyanobacterial blooms in Lake Joutikas, Finland. Aquat Microb Ecol. 2004;36:201–11.
Schmidt ML, White JD, Denef VJ. Phylogenetic conservation of freshwater lake habitat preference varies between abundant bacterioplankton phyla. Environ Microbiol. 2016;18:1212–26.
Chiang E, Schmidt ML, Berry MA, Biddanda BA, Burtner A, Johengen TH, et al. Verrucomicrobia are prevalent in north-temperate freshwater lakes and display class-level preferences between lake habitats. PloS ONE. 2018;13:1–20.
Simek K, Hornak K, Jezbera J, Nedoma J, Znachor P, Hejzlar J, et al. Spatio-temporal patterns of bacterioplankton production and community composition related to phytoplankton composition and protistan bacterivory in a dam reservoir. Aquat Microb Ecol. 2008;51:249–62.
Šimek K, Nedoma J, Znachor P, Kasalický V, Jezbera J, Hornňák K, et al. A finely tuned symphony of factors modulates the microbial food web of a freshwater reservoir in spring. Limnol Oceanogr. 2014;59:1477–92.
Šimek K, Kasalický V, Zapomělová E, Horňák K. Alga-derived substrates select for distinct Betaproteobacterial lineages and contribute to niche separation in Limnohabitans strains. Appl Environ Microbiol. 2011;77:7307–15.
Horňák K, Kasalický V, Šimek K, Grossart HP. Strain‐specific consumption and transformation of alga‐derived dissolved organic matter by members of the Limnohabitans‐C and Polynucleobacter‐B clusters of Betaproteobacteria. Environ Microbiol. 2017;19:4519–35.
Spaink HP. Root nodulation and infection factors produced by Rhizobial bacteria. Annu Rev Microbiol. 2000;54:257–88.
Pérez‐Pantoja D, Donoso R, Agulló L, Córdova M, Seeger M, Pieper DH, et al. Genomic analysis of the potential for aromatic compounds biodegradation in Burkholderiales. Environ Microbiol. 2012;14:1091–117.
Denef VJ, Carrick HJ, Cavaletto J, Chiang E, Johengen TH, Vanderploeg HA. Lake bacterial assemblage composition is sensitive to biological disturbance caused by an invasive filter feeder. mSphere. 2017;2:e00189-17.
Acknowledgements
We thank Dylan Baker, James Lauer, Anna Ortega, and Nate Arringdale for assistance with processing samples. We thank Prof. Brad J. Cardinale (BJC) for contributing algal cultures and facilities, Ruben Props for optimization of the TaxAss/mothur SOP, and Jacob Evans for assistance with data analysis. We also thank participating undergraduate students of the 2018 M-Sci summer class cohort for assisting with baseline experiments. This work was funded by an NSF-EAGER #1737680 to VJD, EFRI-PRSB #1332342 to VJD and BJC, and a Dow Sustainability Postdoctoral Fellowship to SLJ.
Author information
Authors and Affiliations
Contributions
VJD and SLJ conceived of project design, SLJ carried out phycosphere assembly experiment and phycosphere swap experiment, JWY extracted 16S rRNA DNA and analyzed these data, KCS rendered algae axenic and contributed to project design, SLJ and VJD drafted the paper with contributions from all other authors.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Jackrel, S.L., Yang, J.W., Schmidt, K.C. et al. Host specificity of microbiome assembly and its fitness effects in phytoplankton. ISME J 15, 774–788 (2021). https://doi.org/10.1038/s41396-020-00812-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41396-020-00812-x
This article is cited by
-
Microbiome-based enrichment pattern mining has enabled a deeper understanding of the biome–species–function relationship
Communications Biology (2023)
-
Abundant Sulfitobacter marine bacteria protect Emiliania huxleyi algae from pathogenic bacteria
ISME Communications (2023)
-
Phage Infection Benefits Marine Diatom Phaeodactylum tricornutum by Regulating the Associated Bacterial Community
Microbial Ecology (2023)
-
Selection constrains lottery assembly in the microbiomes of closely related diatom species
ISME Communications (2022)
-
Disentangling Diet- and Medium-Associated Microbes in Shaping Daphnia Gut Microbiome
Microbial Ecology (2022)