Abstract
Background
Artificial intelligence (AI) is a promising tool in pathology, including cancer diagnosis, subtyping, grading, and prognostic prediction.
Methods
The aim of the study is to assess AI application in prostate cancer (PCa) histology. We carried out a systematic literature search in 3 databases. Primary outcome was AI accuracy in differentiating between PCa and benign hyperplasia. Secondary outcomes were AI accuracy in determining Gleason grade and agreement among AI and pathologists.
Results
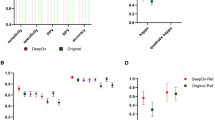
Our final sample consists of 24 studies conducted from 2007 to 2021. They aggregate data from roughly 8000 cases of prostate biopsy and 458 cases of radical prostatectomy (RP). Sensitivity for PCa diagnostic exceeded 90% and ranged from 87% to 100%, and specificity varied from 68% to 99%. Overall accuracy ranged from 83.7% to 98.3% with AUC reaching 0.99. The meta-analysis using the Mantel-Haenszel method showed pooled sensitivity of 0.96 with I2 = 80.7% and pooled specificity of 0.95 with I2 = 86.1%. Pooled positive likehood ratio was 15.3 with I2 = 87.3% and negative – was 0.04 with I2 = 78.6%. SROC (symmetric receiver operating characteristics) curve represents AUC = 0.99. For grading the accuracy of AI was lower: sensitivity for Gleason grading ranged from 77% to 87%, and specificity from 82% to 90%.
Conclusions
The accuracy of AI for PCa identification and grading is comparable to expert pathologists. This is a promising approach which has several possible clinical applications resulting in expedite and optimize pathology reports. AI introduction into common practice may be limited by difficult and time-consuming convolutional neural network training and tuning.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 4 print issues and online access
$259.00 per year
only $64.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
Data availability
The authors confirm that the data supporting the findings of this study are available within the article and its supplementary materials. Any additional data related to this study are available on request from the corresponding author Dmitry Enikeev.
References
Stamatiou K, Alevizos A, Agapitos E, Sofras F. Incidence of impalpable carcinoma of the prostate and of non-malignant and precarcinomatous lesions in Greek male population: An autopsy study. Prostate [Internet]. 2006;66:1319–28. https://onlinelibrary.wiley.com/doi/10.1002/pros.20339
Culp MB, Soerjomataram I, Efstathiou JA, Bray F, Jemal A. Recent global patterns in prostate cancer incidence and mortality rates. Eur Urol [Internet]. 2020;77:38–52. https://linkinghub.elsevier.com/retrieve/pii/S0302283819306190
Sun Y, Reynolds HM, Parameswaran B, Wraith D, Finnegan ME, Williams S, et al. Multiparametric MRI and radiomics in prostate cancer: A review. Australas Phys Eng Sci Med [Internet]. 2019;42:3–25. http://link.springer.com/10.1007/s13246-019-00730-z
Szentirmai E, Giannico GA. Intraductal carcinoma of the prostate. Pathologica 2020;112:17–24.
Truong M, Frye T, Messing E, Miyamoto H. Historical and contemporary perspectives on cribriform morphology in prostate cancer. Nat Rev Urol [Internet]. 2018;15:475–82. http://www.nature.com/articles/s41585-018-0013-1
Hamet P, Tremblay J. Artificial intelligence in medicine. Metab [Internet]. 2017;69:S36–40. https://linkinghub.elsevier.com/retrieve/pii/S002604951730015X
Ramesh A, Kambhampati C, Monson J, Drew P. Artificial intelligence in medicine. Ann R Coll Surg Engl [Internet]. 2004;86:334–8. http://www.ingentaselect.com/rpsv/cgi-bin/cgi?ini=xref&body=linker&reqdoi=10.1308/147870804290
Akkus Z, Cai J, Boonrod A, Zeinoddini A, Weston AD, Philbrick KA, et al. A Survey of Deep-Learning Applications in Ultrasound: Artificial Intelligence–Powered Ultrasound for Improving Clinical Workflow. J Am Coll Radio [Internet]. 2019;16:1318–28. https://linkinghub.elsevier.com/retrieve/pii/S1546144019307112
Mintz Y, Brodie R. Introduction to artificial intelligence in medicine. Minim Invasive Ther Allied Technol [Internet]. 2019;28:73–81. https://www.tandfonline.com/doi/full/10.1080/13645706.2019.1575882
Niel O, Bastard P. Artificial Intelligence in Nephrology: Core Concepts, Clinical Applications, and Perspectives. Am J Kidney Dis [Internet]. 2019;74:803–10. https://linkinghub.elsevier.com/retrieve/pii/S027263861930842X
Jiang Y, Yang M, Wang S, Li X, Sun Y. Emerging role of deep learning‐based artificial intelligence in tumor pathology. Cancer Commun [Internet]. 2020;40:154–66. https://onlinelibrary.wiley.com/doi/10.1002/cac2.12012
Niazi MKK, Parwani AV, Gurcan MN. Digital pathology and artificial intelligence. Lancet Oncol [Internet]. 2019;20:e253–61. https://linkinghub.elsevier.com/retrieve/pii/S1470204519301548
Acs B, Rantalainen M, Hartman J. Artificial intelligence as the next step towards precision pathology. J Intern Med [Internet]. 2020;288:62–81. https://onlinelibrary.wiley.com/doi/10.1111/joim.13030
Phillips B, Ball C, Sackett D, Badenoch D, Straus S, Haynes B, et al. Levels of Evidence [Internet]. Oxford Centre for Evidence-based Medicine. [cited 2020 Aug 20]. Available from: https://www.cebm.net/2009/06/oxford-centre-evidence-based-medicine-levels-evidence-march-2009/
Wittke C, Mayer J, Schweiggert F. On the classification of prostate carcinoma with methods from spatial statistics. IEEE Trans Inf Technol Biomed [Internet]. 2007;11:406–14. http://ieeexplore.ieee.org/document/4267694/
Gorelick L, Veksler O, Gaed M, Gomez JA, Moussa M, Bauman G, et al. Prostate Histopathology: Learning Tissue Component Histograms for Cancer Detection and Classification. IEEE Trans Med Imaging [Internet]. 2013;32:1804–18. http://ieeexplore.ieee.org/document/6522505/
Chen C, Huang Y, Fang P, Liang C, Chang R. A computer‐aided diagnosis system for differentiation and delineation of malignant regions on whole‐slide prostate histopathology image using spatial statistics and multidimensional DenseNet. Med Phys [Internet]. 2020;47:1021–33. https://onlinelibrary.wiley.com/doi/10.1002/mp.13964
Lucas M, Jansen I, Savci-Heijink CD, Meijer SL, de Boer OJ, van Leeuwen TG, et al. Deep learning for automatic Gleason pattern classification for grade group determination of prostate biopsies. Virchows Arch [Internet]. 2019;475:77–83. http://link.springer.com/10.1007/s00428-019-02577-x
Nir G, Karimi D, Goldenberg SL, Fazli L, Skinnider BF, Tavassoli P, et al. Comparison of Artificial Intelligence Techniques to Evaluate Performance of a Classifier for Automatic Grading of Prostate Cancer From Digitized Histopathologic Images. JAMA Netw Open [Internet]. 2019;2:e190442 http://jamanetworkopen.jamanetwork.com/article.aspx?doi=10.1001/jamanetworkopen.2019.0442
Arvaniti E, Fricker KS, Moret M, Rupp N, Hermanns T, Fankhauser C, et al. Automated Gleason grading of prostate cancer tissue microarrays via deep learning. Sci Rep. [Internet]. 2018;8:12054 http://www.nature.com/articles/s41598-018-30535-1
Bulten W, Bándi P, Hoven J, Loo R, van de, Lotz J, Weiss N, et al. Epithelium segmentation using deep learning in H&E-stained prostate specimens with immunohistochemistry as reference standard. Sci Rep. [Internet]. 2019;9:864 http://www.nature.com/articles/s41598-018-37257-4
Ambrosini P, Hollemans E, Kweldam CF, van Leenders GJLH, Stallinga S, Vos F. Automated detection of cribriform growth patterns in prostate histology images. Sci Rep. [Internet]. 2020;10:1–13. https://doi.org/10.1038/s41598-020-71942-7
Bulten W, Pinckaers H, van Boven H, Vink R, de Bel T, van Ginneken B, et al. Automated deep-learning system for Gleason grading of prostate cancer using biopsies: a diagnostic study. Lancet Oncol [Internet]. 2020;21:233–41. https://doi.org/10.1016/S1470-2045(19)30739-9
Egevad L, Swanberg D, Delahunt B, Ström P, Kartasalo K, Olsson H, et al. Identification of areas of grading difficulties in prostate cancer and comparison with artificial intelligence assisted grading. Virchows Arch [Internet]. 2020;477:777–86. https://link.springer.com/10.1007/s00428-020-02858-w
Lokhande A, Bonthu S, Singhal N Carcino-Net: A Deep Learning Framework for Automated Gleason Grading of Prostate Biopsies. In: 2020 42nd Annual International Conference of the IEEE Engineering in Medicine & Biology Society (EMBC) [Internet]. IEEE; 2020. p. 1380–3. Available from: https://ieeexplore.ieee.org/document/9176235/
Marginean F, Arvidsson I, Simoulis A, Christian Overgaard N, Åström K, Heyden A, et al. An Artificial Intelligence–based Support Tool for Automation and Standardisation of Gleason Grading in Prostate Biopsies. Eur Urol. Focus [Internet]. 2020;46:5–11. https://linkinghub.elsevier.com/retrieve/pii/S2405456920302960
Nagpal K, Foote D, Tan F, Liu Y, Chen P-HC, Steiner DF, et al. Development and Validation of a Deep Learning Algorithm for Gleason Grading of Prostate Cancer From Biopsy Specimens. JAMA Oncol [Internet]. 2020;6:1372 https://jamanetwork.com/journals/jamaoncology/fullarticle/2768225
Pantanowitz L, Quiroga-Garza GM, Bien L, Heled R, Laifenfeld D, Linhart C, et al. An artificial intelligence algorithm for prostate cancer diagnosis in whole slide images of core needle biopsies: a blinded clinical validation and deployment study. Lancet Digit Heal [Internet] 2020;2:e407–16. https://doi.org/10.1016/S2589-7500(20)30159-X
Raciti P, Sue J, Ceballos R, Godrich R, Kunz JD, Kapur S, et al. Novel artificial intelligence system increases the detection of prostate cancer in whole slide images of core needle biopsies. Mod Pathol [Internet]. 2020;33:2058–66. https://doi.org/10.1038/s41379-020-0551-y
Silva-Rodríguez J, Colomer A, Sales MA, Molina R, Naranjo V. Going deeper through the Gleason scoring scale: An automatic end-to-end system for histology prostate grading and cribriform pattern detection. Comput Methods Prog Biomed [Internet]. 2020;195:105637. https://linkinghub.elsevier.com/retrieve/pii/S016926072031470X
Steiner DF, Nagpal K, Sayres R, Foote DJ, Wedin BD, Pearce A, et al. Evaluation of the Use of Combined Artificial Intelligence and Pathologist Assessment to Review and Grade Prostate Biopsies. JAMA Netw Open [Internet]. 2020;3:e2023267. https://jamanetwork.com/journals/jamanetworkopen/fullarticle/2772831
Ström P, Kartasalo K, Olsson H, Solorzano L, Delahunt B, Berney DM, et al. Artificial intelligence for diagnosis and grading of prostate cancer in biopsies: a population-based, diagnostic study. Lancet Oncol [Internet]. 2020;21:222–32. https://linkinghub.elsevier.com/retrieve/pii/S1470204519307387
Yang Q, Xu Z, Liao C, Cai J, Huang Y, Chen H, et al. Epithelium segmentation and automated Gleason grading of prostate cancer via deep learning in label‐free multiphoton microscopic images. J Biophotonics [Internet]. 2020;13:1–11. https://onlinelibrary.wiley.com/doi/10.1002/jbio.201900203
Bulten W, Balkenhol M, Belinga J-JA, Brilhante A, Çakır A, Egevad L, et al. Artificial intelligence assistance significantly improves Gleason grading of prostate biopsies by pathologists. Mod Pathol [Internet]. 2021;34:660–71. https://www.nature.com/articles/s41379-020-0640-y
Kudo MS, de Souza VMG, de Souza Amaral G, de Souza Melo PA, Estivallet CLN, Santos ER, et al. The potential of convolutional neural network diagnosing prostate cancer. Res Biomed Eng [Internet]. 2021;37:25–31. http://link.springer.com/10.1007/s42600-020-00095-3
Mun Y, Paik I, Shin S-J, Kwak T-Y, Chang H. Yet Another Automated Gleason Grading System (YAAGGS) by weakly supervised deep learning. npj Digit Med [Internet] 2021;4:99. https://doi.org/10.1038/s41746-021-00469-6
Perincheri S, Levi AW, Celli R, Gershkovich P, Rimm D, Morrow JS, et al. An independent assessment of an artificial intelligence system for prostate cancer detection shows strong diagnostic accuracy. Mod Pathol [Internet]. 2021;34:1588–95. https://doi.org/10.1038/s41379-021-00794-x
Silva LM, Pereira EM, Salles PG, Godrich R, Ceballos R, Kunz JD, et al. Independent real‐world application of a clinical‐grade automated prostate cancer detection system. J Pathol [Internet]. 2021;254:147–58. https://onlinelibrary.wiley.com/doi/10.1002/path.5662
Rigby AS. Statistical methods in epidemiology. v. Towards an understanding of the kappa coefficient. Disabil Rehabil [Internet]. 2000;22:339–44. http://www.tandfonline.com/doi/full/10.1080/096382800296575
Swets J. Measuring the accuracy of diagnostic systems. Science [Internet] 1988;240:1285–93. https://www.sciencemag.org/lookup/doi/10.1126/science.3287615
Rivero Belenchón I, Checcucci E, Gómez Rivas J, Puliatti S, Taratkin M, Kowalewski KF, et al. Comment on “Artificial intelligence to predict oncological outcome directly from hematoxylin and eosin-stained slides in urology: a systematic review.”. Minerva Urol Nephrol. 2022;74:810–2.
Checcucci E, Rosati S, De Cillis S, Vagni M, Giordano N, Piana A, et al. Artificial intelligence for target prostate biopsy outcomes prediction the potential application of fuzzy logic. Prostate Cancer Prostatic Dis. 2022;25:359–62.
Checcucci E, De Cillis S, Granato S, Chang P, Afyouni AS, Okhunov Z. Applications of neural networks in urology: a systematic review. Curr Opin Urol. 2020;30:788–807.
Kumar N, Gupta R, Gupta S. Whole Slide Imaging (WSI) in Pathology: Current Perspectives and Future Directions. J Digit Imaging. 2020;33:1034–40.
Spratt DE, Sun Y, Van der Wal D, Huang S-C, Mohamad O, Armstrong AJ, et al. An AI-derived digital pathology-based biomarker to predict the benefit of androgen deprivation therapy in localized prostate cancer with validation in NRG/RTOG 9408. J Clin Oncol [Internet]. 2022;40:223–223. https://doi.org/10.1200/JCO.2022.40.6_suppl.223. 6_suppl
Beck AH, Sangoi AR, Leung S, Marinelli RJ, Nielsen TO, van de Vijver MJ, et al. Systematic analysis of breast cancer morphology uncovers stromal features associated with survival. Sci Transl Med. 2011;3:108ra113.
Doyle S, Feldman MD, Shih N, Tomaszewski J, Madabhushi A. Cascaded discrimination of normal, abnormal, and confounder classes in histopathology: Gleason grading of prostate cancer. BMC Bioinforma. 2012;13:282.
LeCun Y, Bengio Y, Hinton G. Deep learning. Nature 2015;521:436–44.
Angermueller C, Pärnamaa T, Parts L, Stegle O. Deep learning for computational biology. Mol Syst Biol. 2016;12:878.
El Naqa I, Boone JM, Benedict SH, Goodsitt MM, Chan H-P, Drukker K, et al. AI in medical physics: guidelines for publication. Vol. 48, Medical physics. United States; 2021. p. 4711–4.
Ruamviboonsuk P, Chantra S, Seresirikachorn K, Ruamviboonsuk V, Sangroongruangsri S. Economic Evaluations of Artificial Intelligence in Ophthalmology. Asia-Pac J Ophthalmol [Internet]. 2021;10:307–16. https://journals.lww.com/10.1097/APO.0000000000000403
Mori Y, Kudo S, East JE, Rastogi A, Bretthauer M, Misawa M, et al. Cost savings in colonoscopy with artificial intelligence-aided polyp diagnosis: An add-on analysis of a clinical trial (with video). Gastrointest Endosc [Internet]. 2020;92:905–911.e1. https://linkinghub.elsevier.com/retrieve/pii/S0016510720340347
Mayo RC, Leung JWT. Impact of Artificial Intelligence on Women’s Imaging: Cost-Benefit Analysis. Am J Roentgenol [Internet]. 2019;212:1172–3. https://www.ajronline.org/doi/10.2214/AJR.18.20419
Salcedo J, Rosales M, Kim JS, Nuno D, Suen S-C, Chang AH. Cost-effectiveness of artificial intelligence monitoring for active tuberculosis treatment: A modeling study. Durand-Zaleski I, editor. PLoS One [Internet]. 2021 Jul;16:e0254950. Available from: https://dx.plos.org/10.1371/journal.pone.0254950
Schwendicke F, Rossi JG, Göstemeyer G, Elhennawy K, Cantu AG, Gaudin R, et al. Cost-effectiveness of Artificial Intelligence for Proximal Caries Detection. J Dent Res [Internet]. 2021;100:369–76. http://journals.sagepub.com/doi/10.1177/0022034520972335
Loeb S, Bjurlin MA, Nicholson J, Tammela TL, Penson DF, Carter HB, et al. Overdiagnosis and Overtreatment of Prostate Cancer. Eur Urol [Internet]. 2014;65:1046–55. https://linkinghub.elsevier.com/retrieve/pii/S0302283813014905
Author information
Authors and Affiliations
Consortia
Contributions
AM: conception; interpretation; writing. MT: conception; writing. AB: interpretation; writing. JGR: conception; editing. SP: conception; editing. EC: conception; editing. IRB: conception; editing. K-FK: conception; editing. AS: conception; writing. NS: conception; editing. JYCT: conception; editing. VK: interpretation; statistical analysis. ALA: conception; editing. GEC: conception; editing. DE: conception; interpretation; writing.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Morozov, A., Taratkin, M., Bazarkin, A. et al. A systematic review and meta-analysis of artificial intelligence diagnostic accuracy in prostate cancer histology identification and grading. Prostate Cancer Prostatic Dis 26, 681–692 (2023). https://doi.org/10.1038/s41391-023-00673-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41391-023-00673-3
This article is cited by
-
Developers-Doctor-patients: the artificial intelligence’s trifecta
Prostate Cancer and Prostatic Diseases (2024)
-
Quality of information and appropriateness of Open AI outputs for prostate cancer
Prostate Cancer and Prostatic Diseases (2024)
-
Artificial intelligence and urology: ethical considerations for urologists and patients
Nature Reviews Urology (2024)