Abstract
Background
Preterm infants suffer higher morbidity and mortality rates compared to full-term infants, but little is known about how changes to oral and respiratory tract microbiota may impact disease development.
Methods
Here, very preterm neonates (n = 50) were selected to study oral and respiratory microbiota development during the first few months post-birth, where 26 individuals were diagnosed with BPD and/or sepsis. These infants were compared to 14 healthy full-term infants and 16 adults. Microbiota diversity, composition, and species abundances were calculated from 16S ribosomal RNA gene sequences in buccal swabs and tracheal aspirates at two time points (within a week and 1–3 months post-birth).
Results
Collection time point was the biggest factor to significantly influence the preterm oral microbial diversity and composition. In addition, BPD and sepsis were linked to distinct preterm oral microbiota diversity and composition, and opportunistic pathogens previously associated with these diseases were identified in the initial sample for both healthy preterm neonates and those with the disease. Compared to the full-term infant and adult dataset, preterm infant diversity and composition was initially significantly different, but resembled full-term infant diversity and composition over time.
Conclusion
Overall, consequences of microbiota development need further examination in preterm infant infections and later development.
Impact
-
Non-gut microbiota research on preterm infants is limited.
-
At one week post-birth, preterm infants harbor distinct oral microbiota that are not shared with full-term children or adults, eventually becoming similar to full-term infants at 36 weeks postmenstrual age.
-
DNA from potential opportunistic pathogens was observed in the mouth and lungs of preterm infants within a week of birth, and microbes associated with BPD were identified in the lungs.
-
Oral microbiota in preterm infants over the first 2–3 months is unique and may be connected to short- and long-term health outcomes in these children.
Similar content being viewed by others
Introduction
Compared to full-term neonates, infants born prior to 37 weeks gestation (preterm) have a higher predisposition to chronic and infectious diseases, as many essential organs are underdeveloped at birth.1 For example, underdeveloped lungs often require oxygen, surfactants, and mechanical ventilation, which cause further damage to the lungs, and contribute to bronchopulmonary dysplasia (BPD).2 Pathogens, such as Ureaplasma,3 Streptococcus anginosus,4 and Streptococcus agalactiae (Group B Streptococcus),5 have been identified in the mouth and lungs of individuals with BPD6 and can cause infections that lead to sepsis or pneumonia,1 although the specific causes that underpin these diseases remain an active area of research.
Current research suggests that preterm health complications may be associated with the infant’s postnatal environment.7 Preterm infants live in a comparatively “unnatural” environment—within the neonatal intensive care unit—to most full-term infants. This altered environment, as well as delivery mode,8,9 feeding method,10,11 skin-to-skin contact (kangaroo care),12 and antibiotic use,13 can disturb or delay the commensal bacteria that colonize infants (microbiota) and are critical for health.14 Preterm infant microbiota research focused on non-gut sites is limited, although several studies are now investigating oral12,15,16,17 and skin16,18 microbiota in these children. Even fewer studies have examined preterm infant microbiota development over time, especially outside of the gut.19 In particular, oral microbiota development needs further attention, as oral microbiota can underpin both local and systemic diseases20,21 and may provide key insights into the development of short- and long-term health impacts for preterm infants, especially in the respiratory tract.
Early alterations to preterm infant microbiota could have downstream effects on development and health outcomes. Many alterations to preterm microbiota have been identified during the neonatal immunity window of opportunity—a non-redundant priming phase of the immune system.22,23 As preterm infants are highly susceptible to infections, it is possible that critical immune system processes, such as innate immune tolerization and reprogramming, are also altered in these infants.24 Indeed, Olin et al. (2018) demonstrated that preterm and full-term infant immune systems initially respond differently after birth, but profiles eventually converge after 3 months.25 Existing studies have identified a mirrored delay in gut microbiota development in preterm infants, suggesting that a linked delay in microbiota and immune system development during this critical window may contribute to both short- and long-term health consequences.25,26 Despite this, we know very little about how similar microbiota alterations outside of the gut in preterm infants may also play roles in immediate and life-long health.
Here, microbiota assessment was performed on a subset of infants born <29 weeks’ gestation (n = 50) who were participating in the n-3 Fatty Acids for Improvement in Respiratory Outcomes (N3RO) randomized controlled trial—a trial including 1273 extremely-to-very preterm infants, which investigated BPD development after docosahexaenoic acid (DHA) supplementation.27 Specifically, we characterized the development of oral and respiratory tract microbiota in healthy infants compared to those that developed the disease (i.e., BPD and sepsis). Oral swabs were collected within one week of birth and at 36 weeks’ postmenstrual age to compare oral microbiota diversity and compositional development over time with a published full-term infant and adult dataset.28 Overall, this study aims to investigate oral microbiota development in preterm infants during the first months of life and identify unique signatures in the oral microbiota that may be linked to disease development.
Methods
Study population and sample collection
All microbiota samples were collected under the Human Research Ethics Committee approval obtained for both preterm infants (Women and Children’s Hospital; HREC 2434/12/16; November 27, 2013) and adults (University of Adelaide; H2012-108). A subset of preterm infants enrolled in the N3RO trial, which investigated the effects of DHA, were selected for this study based on the opportunistic timing of their birth (2014–2015).27 Based on previous power calculations for human microbiota studies,29 we aimed to collect at least 20 healthy preterm infants and 20 with BPD. For all preterm (P) infants, patient metadata and population information were collected by medical record examination (Table 1). Preterm infant buccal swabs were collected at two time points—2–12 days post-birth (PT1) and 36 weeks’ postmenstrual age (PT2)—from 50 preterm infants delivered at the Women’s and Children’s Hospital in Adelaide, Australia (Fig. 1). Tracheal aspirate samples were collected from 14 preterm neonates who required intubation due to medical complications.
Oral samples were collected at similar time points from preterm and full-term infants (approximately during the first week post-birth and one month later). Adult oral samples were included in the analyses to further understand microbiota maturation of the infants. For preterm infants that were intubated (n = 14), tracheal aspirates were also collected to identify shared taxa between the mouth and lungs of these infants. Study ID refers to the QIITA study ID.
All samples (buccal swabs and tracheal aspirates) were frozen (–20 °C) immediately in empty tubes after collection to preserve the bacterial composition. To further explore infant oral maturation, published oral microbiota data were obtained from 13 mothers and 14 full-term (F) infants collected at two similar time points: 0–7 days post-birth (FT1) and 4th–5th week post-birth (FT2) (Study ID 2010, QIITA data repository; Fig. 1), but minimal, irrelevant metadata was available and therefore was not reported. To control for and track potential background signatures in our approach (i.e., comparing two datasets prepared in different labs with a different extraction method), three adult samples were also collected and processed following the same protocols as the preterm infant samples.
Sample DNA extraction, 16S ribosomal RNA library preparation, and DNA sequencing
All collected samples were prepared in a still-air room designed for low-biomass microbiota analysis using strict measures to reduce cross-contamination and the introduction of background DNA,30 as described in the Supplementary material. DNA was extracted from buccal swabs and tracheal aspirates using a previously published, in-house silica DNA extraction method designed to enhance DNA recovery;31 the method was modified to include mechanical lysis (see Supplementary material). DNA present within the laboratory and reagents was monitored using extraction blank controls (EBCs).
Bacterial DNA from each sample, including EBCs, was amplified in triplicate using primers that target the V4 region of the 16S ribosomal RNA (rRNA) gene,32 using previously described amplification conditions33 (also described in the Supplementary material). 16S rRNA libraries were prepared for sequencing (see Supplementary material).34 Libraries were sequenced with an Illumina MiSeq (2 × 150 bp) at the Australian Genomics Research Facility.
Pre-processing, ASV selection, and contaminant removal
Preterm infant sequences (PT1, PT2, and tracheal aspirates) and adult (n = 3) sequences were uploaded to QIITA data repository (Study ID 11832; https://qiita.ucsd.edu/study/description/11832). Demultiplexed sequences were trimmed to 150 bp and amplicon sequence variants (ASVs) were generated via Deblur.35 ASVs were merged with a full-term infant (FT1 and FT2) and maternal dataset (Study ID 2010) to create a SEPP insertion tree36 and the study dataset. To reduce bias from laboratory work being carried out in different laboratories, contaminant sequences in EBCs and NTCs from the preterm dataset were identified via Decontam37 and were subsequently removed from all biological samples (Supplementary Table S1) in QIIME2 (v2019.7).38 Following this, ASVs with less than 10 reads assigned were removed.
Diversity analyses, taxonomic classification, and statistical comparisons
In QIIME2 (v2019.7),38 ASVs were summarized into their taxonomic classification using the SILVA database (v132; 16S V4 515-806).39 Using a rarefied depth of 2000 sequences, (alpha) diversity was measured using observed species (OS) and Faith’s phylogenetic diversity (PD)40 metrics, and beta diversity (composition) was calculated using the unweighted UniFrac metric.41 In QIIME2, significant associations (q < 0.05; q = Benjamini–Hochberg FDR correction) between diversity and sample metadata were detected using pairwise Kruskal–Wallis tests,42 while the pairwise Fligner–Killeen test (R, stats package),43,44 was used to determine variability in diversity. Significant links between composition and metadata were examined using adonis45,46 (p < 0.05) and PERMANOVA (q < 0.05).45 For principal coordinates analysis (PCoA), 95% confidence intervals were determined using the stats eclipse function with ggplot2. For categories that had missing data (BPD (n = 2 missing) and probiotic, antibiotic, and antifungal administration (n = 10 missing), samples were removed prior to running analyses for those specific categories. Lastly, ANCOM47 was used to identify significant changes in an ASV abundance across samples, and only taxa that were significantly different in abundance for pairwise comparisons were displayed along with their corresponding W statistic value.
Results
Preterm infant cohort and assessment of comparative data
Preterm infants (n = 50; Table 1) in this study were enrolled in the N3RO trial, which investigated the effect of DHA supplementation (n = 26). These infants were born prior to 29 weeks’ gestation (26.4 ± 1.7 weeks) and had two collection time points—within the first week of birth (T1 = 4.2 ± 1.6 days) and at the equivalence of 36 weeks’ postmenstrual age (T2 = 65.7 ± 14.6). Twenty-six of these preterm infants developed BPD and/or sepsis (BPD only, n = 13; sepsis only, n = 6; and BPD and sepsis, n = 7). The impact of preterm infants that were twins or triplets (n = 15), infants that were given antibiotics (n = 39 of 40) as part of standard care, or DHA supplementation, were assessed in downstream analyses (Table 2).
As part of this study, we compare preterm infant oral microbiota development from this cohort to that of full-term infants and adults from a published study. To ensure that any biases due to preparation in different laboratories did not affect comparison and interpretation, multiple assessments were performed. Firstly, as there were no EBCs available for the full-term dataset, contaminant sequences observed in our preterm dataset (Supplementary Table S1) were removed from both datasets. These contaminant ASVs were typical of common laboratory contaminants.30,48,49 In addition, three adult samples processed identically to preterm infant samples were compared to the adult samples processed identically to the full-term infant samples to determine if the laboratory differences affected our assessment. Adult samples from both studies clustered together (Supplementary Fig. S1), indicating that minor differences in extraction processes minimally affected the samples and that these two studies are comparable. Following these assessments, in total 156 samples were retained and represented 8,738,663 sequences and 1553 ASVs.
Preterm oral microbiota stabilizes over time but is less affected by common maternal and environmental factors
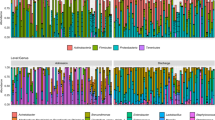
First, we wanted to investigate the diversity and composition of the preterm oral microbiota over the first few months post-birth. We observed high variability and diversity in PT1 samples (Fig. 2a, b) and a reduction in variability (Fligner–Killeen; OS: χ2 = 33.15, p = <0.0001; PD: χ2 = 29.91, p = <0.0001) and diversity (Kruskal–Wallis; OS: H = 10.47, q = 0.003; PD: H = 10.96, q = 0.002) in PT2 samples. This suggests that each preterm infant initially responds uniquely to their environment or inherits a different microbiota from their mother’s birth canal (or lack of passing through), but oral microbiota stabilize by 36 weeks’ postmenstrual age. We also observed that the composition of preterm infants separated according to time point (PERMANOVA; pseudo-F = 8.191, q = 0.001; Fig. 2c, d). To better understand compositional differences, we investigated microbial taxa abundance at the genus taxonomic level. While PT1 samples were highly variable between individuals (Supplementary Fig. S2), a clear distinction was observed between PT1 and PT2 samples. Specifically, Staphylococcus (average relative abundance; 61.8%) dominated PT1 samples (Fig. 3) and was significantly more abundant compared to PT2 (3.6%). Conversely, PT2 samples were dominated (relative abundance >1%) by Streptococcus (71.6%), but were also colonized by Gemella (6.1%), Rothia (5.7%), Veillonella (4.1%), Staphylococcus (3.6%), Prevotella (2.5%), and Haemophilus (1.2%). Overall, this suggests that oral microbiota variability in preterm infants decreases and stabilizes within two months.
Using 2000 sequences per sample, alpha diversity variation decreases over time in preterm infants using both (a) observed species and (b) Faith’s phylogenetic diversity metrics. The factor that had the most significant impact was the collection time point of the preterm samples, showing the differences in composition over a short period of time (c and d). Using unweighted UniFrac metric, composition of PT1 samples was more densely populated in the bottom section of PC2 but was spread across PC1, whereas PT2 and full-term infant samples had small density peaks across PC1 but overlapped each other in PC2 (e). Adults were significantly different from infant samples over Axis 2 and 3 (f). Samples were color-coded according to the time of collection and if they were a preterm or full-term infant, or adult. Pairwise significant differences are labeled with the corresponding q value.
Next, we wanted to investigate whether common maternal and environmental factors influenced the preterm oral microbiota. For the full preterm dataset (i.e., PT1 and PT2 samples combined), and the individual time points (i.e., PT1 and PT2 datasets individually), typical factors including feeding method and number of antibiotic doses did not significantly impact the preterm oral microbiota (Table 2). However, postnatal day had a significant influence on the microbial diversity and composition on the overall preterm and first time point datasets (Kruskal–Wallis q < 0.05; adonis p < 0.05; Table 2 and Supplementary Fig. S3). Birth mode, more specifically between caesarean section with and without labor induction, approached significance for microbial diversity within the overall preterm dataset (Kruskal–Wallis 0.05 < q < 1; Table 2), while the sex of the infant (Kruskal–Wallis; OS: H = 0.519 q = 0.023; PD: H = 7.782, q = 0.005; adonis p = 0.009; Table 2 and Supplementary Fig. S3) and number of antifungal doses significantly influenced the oral microbiota composition in the PT2 samples (adonis p = 0.006; Table 2 and Supplementary Fig. S3). We also determined that DHA supplementation did not influence the preterm oral microbiota overall and at each collection time points (Table 2).
Assessment of the preterm oral microbiota in children with BPD and sepsis
As BPD and sepsis commonly affect preterm infants, we wanted to determine if BPD and sepsis are associated with shifts in the oral microbiota during the first few months post-birth. In infants at approximately four days old, the presence of BPD in preterm infants approached significance for oral microbiota diversity (OS: H = 2.935, q = 0.087; PD: H = 3.593, q = 0.058; Fig. 4) but not composition (adonis; R2 = 0.032, p = 0.112; Fig. 4). At this age, infants that presented with BPD during this study had a lower microbial diversity at T1 (median: 13 ASVs) compared to infants who did not present BPD (median: 35 ASVs). This was largely driven by the presence of Corynebacterium, which was found at 11.3% relative abundance in infants that developed BPD but only at 0.9% relative abundance in infants that did not develop BPD; however, this result was not significant using an ANCOM test of differential abundance. We also examined if the impacts of treating BPD with DHA or high dosages of antibiotics may have confounded this signal, and this was not the case (adonis; DHA = R2 = 0.034, p = 0.087; antibiotics dosage = R2 = 0.055, p = 0.330). At 36 weeks’ postmenstrual age, diversity and composition was not significantly different in infants with BPD.
Lower diversity is observed in PT1 samples for preterm infants that went on to develop BPD (pink) during this study (a, b); however, there was no compositional distinction between preterm infants that did or did not develop BPD (c). At the second time point, preterm infants who developed sepsis (purple) during this study had a significantly lower microbial richness (d), but not phylogenetic diversity (e). Significantly different oral microbiota composition was observed between infants that did or did not develop sepsis (f). Darker shades represent samples that developed either BPD and/or sepsis during this study.
For infants at approximately four days of age, sepsis had no effects on the diversity and composition of the oral microbiota. However, at 36 weeks postmenstrual age, infants that were diagnosed with sepsis during this study had a significantly lower microbial richness, but not PD (median of 7 ASVs for children with sepsis vs median of 10 ASVs for children without sepsis; OS: H = 3.949, q = 0.044; PD: H = 2.569, q = 0.109; Fig. 4) and significantly different composition (adonis: R2 = 0.046, p = 0.031; Fig. 4). While no species or genera were significantly more abundant in infants that presented sepsis, we observed Staphylococcus spp. at a higher relative abundance (13.6%) in infants with sepsis diagnosis, compared to infants that were not diagnosed with sepsis (0.6%; Supplementary Fig. S2). Interestingly, only four of thirteen infants diagnosed with sepsis were females. Furthermore, there were no significant associations for the interaction between BPD and sepsis, although PT1 approached significance (Supplementary Table S2). There were also no significant confounding effects between sepsis with DHA or high dosages of antibiotics (adonis; DHA: R2 = 0.028, p = 0.253; antibiotics dosage: R2 = 0.048, p = 0.271). As these results are difficult to disentangle, more samples are needed to determine if sepsis contributes to differences in the oral microbiota.
Oral microbiota are shared with lung microbiota
Unfortunately, some preterm infants enrolled in this study were intubated due to BPD complications. In these infants, we identified ASVs shared between PT1 oral samples and tracheal aspirate samples in the same individual (n = 12). Overall, 32 ASVs were shared between a PT1 oral sample and tracheal aspirate in at least one individual. Although lung samples contained few microbes and had stochastic diversity and composition, the most dominant shared ASVs (>1000 reads across individuals) were assigned as Paenibacillus, Staphylococcus, Erythrobacteraceae bacterium K-2-3, Streptococcus anginosus, Ureaplasma, Burkholderiaceae, and Streptococcus agalactiae sequences (Supplementary Fig. S4). Ureaplasma increases the risk of developing BPD4 and was also found as dominant taxa in PT1 samples of children suffering from BPD who did not receive intubation (Supplementary Figs. S2 and S4; >1000 sequences; healthy = 8, BPD = 2). In addition, Streptococcus anginosus and Streptococcus agalactiae have previously been associated with BPD and sepsis, respectively, and were also detected here; Streptococcus anginosus was found as a dominant species in one individual with BPD, while Streptococcus agalactiae was found as a dominant species in one individual with sepsis. Ureaplasma, Streptococcus anginosus, and Streptococcus agalactiae all reduced in abundance by T2 collection point. Together, this suggests that early oral colonizers in preterm infants can also spread to the lungs and may be linked to disease complications. However, these results also suggest that stabilization and maturation of the oral microbiota is linked to a reduction of opportunistic pathogens in these preterm children.
Preterm oral microbiota at two months are similar to full-term infants
To understand preterm oral microbiota maturation, we compared all preterm and full-term infant oral microbiota at two analogous time points (PT1 and PT2 vs. FT1 and FT2), as well as adult oral samples. Interestingly, PT2 and FT1 samples both contained similar, yet low, microbial diversity and were not significantly different from one another (Kruskal–Wallis; OS: H = 0.041, q = 0.840; PD: H = 0.257, q = 0.612). Adult oral microbiota diversity was significantly different from both preterm and full-term infant samples (Fig. 2a, b and Supplementary Table S3; q < 0.05), suggesting that further maturation of infant microbiota may continue over time, as previously reported.50 Overall, this suggests that oral microbiota variability in preterm infants decreases and stabilizes within two months to resemble full-term neonates more closely.
Through the analysis of composition diversity, it was determined that preterm and full-term infant microbiota were significantly different from one another, as well as from adult samples (Fig. 2e, f and Supplementary Table S3; q < 0.05). While there was a minimal separation between sample groups in PC1 of the principal coordinate analysis, PT1, FT2, and adult samples clustered in their own groups in PC2 (Fig. 2e). However, PT2 and FT1 samples were not significantly different from each other (Fig. 2e, f and Supplementary Table S3; q > 0.05). We also noted that PT1, PT2 and FT1 were more dispersed in the PCoA for PC1 and PC2, and that FT2 and adult samples were more tightly clustered, potentially indicating that the oral microbiota stabilize and mature over time.
To further investigate the composition diversity, we compared the taxonomic composition of the infant and adult samples. As observed with the compositional diversity, there were minimal differences between PT2 and FT1 samples. Using ANCOM, the only taxa that significantly differed in abundance between PT2 and FT1 samples were Rothia, Paenibacillus, and Erythrobacteraceae bacterium K-2-3 ASVs, all of which were more abundant in PT2 samples (Supplementary Table S4) and could also be observed in PT1 samples. We also identified several genera present in all infant and adult mouths, including Streptococcus, Gemella, Rothia, Veillonella, Staphylococcus, Prevotella, and Haemophilus ASVs, suggesting that these microbes are introduced from a young age and maintained into adulthood.
Discussion
By comparing oral and lung microbiota of preterm and full-term infants over the first two months of life, we observed marked levels of interindividual variation and compositional differences in newborn preterm infants. However, these observations were ameliorated by 36 weeks postmenstrual age (i.e., children that are 2–3 months old), as their microbes were similar to full-term infants. Interestingly, differences in the overall and PT1 preterm datasets were driven by postnatal day, whereas shifts in PT2 samples were unique depending on the sex of the child, and number of antifungal doses. BPD and sepsis also were linked to shifts in oral microbiota diversity and composition at early and later time points, respectively. Lastly, potential pathogens identified in both the mouth and lungs of preterm infants with BPD or sepsis also provide a unique window into microbial colonization of the lungs linked to poor health outcomes. Overall, monitoring the oral microbiota may provide new insights into understanding the underlying mechanisms of BPD and sepsis in preterm infants compared to full-term counterparts.
Higher morbidity and mortality rates in preterm infants typically arise from respiratory and infectious diseases, including BPD and sepsis. While the initial goal of this study was to examine BPD, sepsis also unfortunately occurred in this cohort and was included in our downstream analyses. Using exact sequences (ASVs), we identified these taxa at the species level and they were assigned as key pathogens associated with BPD and sepsis (i.e., Ureaplasma,3 Streptococcus anginosus,4 and Streptococcus agalactiae (Group B Streptococcus).5 A single previous study surveying the cross-over between the oral and lung microbiota of preterm infants in relation to BPD also found dominant genera that were observed in our study.6 In this study, these pathogens were highly abundant in oral samples at the first time point but reduced in abundance by the second time point. Interestingly, we observed Corynebacterium at high abundance in the mouth of infants with BPD, which has also been observed at high abundance in the lungs of infants with severe BPD.51 We also detected Staphylococcus at a higher abundance in preterm infants with sepsis diagnosis. Indeed, it has been shown that Staphylococcus aureus can cause early- and late-onset sepsis (although more common in late-onset sepsis), as detected by cultures of blood or cerebrospinal fluid.52,53 While we do not yet know if these microbes are causative or present post-initiation of disease, further exploration into the use of these microbes as biomarkers for BPD and sepsis is warranted both in the context of early disease detection and assessment of potential severity. Furthermore, microbiota diversity and composition were distinct in preterm infants with sepsis compared to those without. While one other study has examined the link between sepsis and the oral microbiota, no significant differences in the mouths of infants diagnosed with sepsis were detected, likely due to a small sample size (total n = 7).19 Interestingly, more male infants in this study were affected by sepsis, and overall, we observed lower microbial diversity in male infants at ~65 days of age. In general, male infants more frequently suffer from diseases, such as early-onset sepsis and BPD.54,55 Overall, these results suggest that monitoring oral microbiota diversity in preterm infants may provide insights into respiratory disease development and outcomes, although further work needs to account for high levels of microbial diversity variation in newly born preterm infants.
Several studies postulate that a disruption in the neonatal window of opportunity, which occurs during the first three years of life in layered phases (peri/postnatal, weaning, and post-weaning),24 could be responsible for diseases later in life. This window notably includes a critical period for interactions between the immune system and gut microbiota during the first 100 days,26 so examining how the microbiota develop over this period throughout the entire body is vital to understanding long-term immune system development. Here, we observed that the postnatal day and number of antifungal doses were significantly associated with oral microbial composition in preterm infants, which could have an impact on the initial development of the immune system.25 Furthermore, we see a convergence of microbial diversity and composition over time that resembles full-term infants; this effect has also been reported in the immune system.25 Future studies should examine these intertwined processes at a much finer time scale (i.e., each day during the peri/postnatal phase of immune system development), as well as how the health outcomes of these processes.
Our conclusions are limited by our sample collection times, overall sample size, and potential environmental or geographic signatures. First, variation in diversity within the PT1 samples could have been impacted by the post-birth collection day. If samples were collected on the same day post-birth (e.g., day 3) then this may reduce some of the variation associated with the first collection point. We also recognize that these results are limited to the mouth; collecting samples across the whole body would be required to understand how microbiomes of an individual work together.56 Second, an increased sample size would help to ensure that interactions between sepsis and oral microbiota can be fully elucidated. Thirdly, differences between preterm infants from a single hospital in Australia and a small number of full-term infants from a previously published study may be attributed to gestational age but may equally be confounded by the sampling location. Similarly, the difference between preterm infants with or without BPD could be confounded by gestational age. Lastly, pairing the whole-body approach with immune function, better environmental information (i.e., where the microbes originate from), social surveys, and more metadata would provide a holistic approach to the preterm infant microbiota development, could provide more information on the impacts on the neonatal window of opportunity and long-term health consequences.
Conclusions
Over the first few weeks of life, preterm infants are exposed to unnatural environments, which may influence microbiota colonization and the development of disease. Our study demonstrated that the oral microbiota of preterm infants is initially disturbed after birth but begins to resemble the oral microbiota of full-term infants over time. Furthermore, certain microbes detected in the oral microbiota were linked to the development of BPD and sepsis, suggesting that oral microbiota may play key roles in the development of lethal diseases in preterm infants.
Data availability
Preterm infant sequences and adult (n = 3) sequences were uploaded to QIITA data repository (Study ID 11832) and can be found at https://qiita.ucsd.edu/study/description/11832. Sequences downloaded for full-term infant and adult samples were obtained from the QIITA data repository (Study ID 2010) and can be found at https://qiita.ucsd.edu/study/description/2010.
References
Behrman, R. E., Butler, A. S. & Institute of Medicine (US) Committee on Understanding Premature Birth and Assuring Healthy Outcomes. Mortality and Acute Complications in Preterm Infants. Preterm Birth: Causes, Consequences, and Prevention (National Academies Press (US), 2007).
Thébaud, B. et al. Bronchopulmonary dysplasia. Nat. Rev. Dis. Prim. 5, 1–23 (2019).
Pammi, M. et al. Airway microbiome and development of bronchopulmonary dysplasia in preterm infants: a systematic review. J. Pediatr. 204, 126–133.e2 (2019).
Glaser, K. et al. Perinatal ureaplasma exposure is associated with increased risk of late onset sepsis and imbalanced inflammation in preterm infants and may add to lung injury. Front. Cell. Infect. Microbiol. 9, 68 (2019).
Doran, K. S. & Nizet, V. Molecular pathogenesis of neonatal group B streptococcal infection: no longer in its infancy. Mol. Microbiol. 54, 23–31 (2004).
Brewer, M. R. et al. Determinants of the lung microbiome in intubated premature infants at risk for bronchopulmonary dysplasia. J. Matern. Fetal Neonatal Med. 0, 1–7 (2019).
Santos, J., Pearce, S. E. & Stroustrup, A. Impact of hospital-based environmental exposures on neurodevelopmental outcomes of preterm infants. Curr. Opin. Pediatr. 27, 254–260 (2015).
Dominguez-Bello, M. G. et al. Delivery mode shapes the acquisition and structure of the initial microbiota across multiple body habitats in newborns. Proc. Natl Acad. Sci. USA 107, 11971–11975 (2010).
Chu, D. M. et al. Maturation of the infant microbiome community structure and function across multiple body sites and in relation to mode of delivery. Nat. Med. 23, 314–326 (2017).
Lif Holgerson, P. et al. Oral microbial profile discriminates breast-fed from formula-fed infants. J. Pediatr. Gastroenterol. Nutr. 56, 127–136 (2013).
Ho, N. T. et al. Meta-analysis of effects of exclusive breastfeeding on infant gut microbiota across populations. Nat. Commun. 9, 4169 (2018).
Hendricks-Muñoz, K. D. et al. Skin-to-skin care and the development of the preterm infant oral microbiome. Am. J. Perinatol. 32, 1205–1216 (2015).
Gasparrini, A. J. et al. Persistent metagenomic signatures of early-life hospitalization and antibiotic treatment in the infant gut microbiota and resistome. Nat. Microbiol. 4, 2285–2297 (2019).
Tanaka, M. & Nakayama, J. Development of the gut microbiota in infancy and its impact on health in later life. Allergol. Int. 66, 515–522 (2017).
Sohn, K., Kalanetra, K. M., Mills, D. A. & Underwood, M. A. Buccal administration of human colostrum: impact on the oral microbiota of premature infants. J. Perinatol. 36, 106–111 (2016).
Younge, N. E., Araújo-Pérez, F., Brandon, D. & Seed, P. C. Early-life skin microbiota in hospitalized preterm and full-term infants. Microbiome 6, 98 (2018).
Li, H. et al. Impacts of delivery mode on very low birth weight infants’ oral microbiome. Pediatr. Neonatol. 61, 201–209 (2020).
Pammi, M. et al. Development of the cutaneous microbiome in the preterm infant: a prospective longitudinal study. PLoS One 12, e0176669 (2017).
Young, G. R. et al. Acquisition and development of the extremely preterm infant microbiota across multiple anatomical sites. J. Pediatr. Gastroenterol. Nutr. 70, 12–19 (2020).
Lu, M., Xuan, S. & Wang, Z. Oral microbiota: a new view of body health. Food Sci. Hum. Wellness 8, 8–15 (2019).
Graves, D. T., Corrêa, J. D. & Silva, T. A. The oral microbiota is modified by systemic diseases. J. Dent. Res. 98, 148–156 (2019).
Torow, N. & Hornef, M. W. The neonatal window of opportunity: setting the stage for life-long host-microbial interaction and immune homeostasis. J. Immunol. 198, 557–563 (2017).
Renz, H. et al. The neonatal window of opportunity—early priming for life. J. Allergy Clin. Immunol. 141, 1212–1214 (2018).
Hornef, M. W. & Torow, N. ‘Layered immunity’ and the ‘neonatal window of opportunity’ – timed succession of non-redundant phases to establish mucosal host–microbial homeostasis after birth. Immunology 159, 15–25 (2020).
Olin, A. et al. Stereotypic immune system development in newborn children. Cell 174, 1277–1292.e14 (2018).
Arrieta, M.-C. et al. Early infancy microbial and metabolic alterations affect risk of childhood asthma. Sci. Transl. Med. 7, 307ra152 (2015).
Collins, C. T. et al. Docosahexaenoic acid and bronchopulmonary dysplasia in preterm infants. N. Engl. J. Med. 376, 1245–1255 (2017).
Dominguez-Bello, M. G. et al. Partial restoration of the microbiota of cesarean-born infants via vaginal microbial transfer. Nat. Med. 22, 250–253 (2016).
La Rosa, P. S. et al. Hypothesis testing and power calculations for taxonomic-based human microbiome data. PLoS One 7, e52078 (2012).
Weyrich, L. S. et al. Laboratory contamination over time during low-biomass sample analysis. Mol. Ecol. Resour. 19, 982–996 (2019).
Rohland, N. & Hofreiter, M. Comparison and optimization of ancient DNA extraction. BioTechniques 42, 343–352 (2007).
Caporaso, J. G. et al. Ultra-high-throughput microbial community analysis on the Illumina HiSeq and MiSeq platforms. ISME J. 6, 1621–1624 (2012).
Selway, C. A. et al. Transfer of environmental microbes to the skin and respiratory tract of humans after urban green space exposure. Environ. Int. 145, 106084 (2020).
Rohland, N. & Reich, D. Cost-effective, high-throughput DNA sequencing libraries for multiplexed target capture. Genome Res. 22, 939–946 (2012).
Amir, A. et al. Deblur rapidly resolves single-nucleotide community sequence patterns. mSystems 2, e00191–16 (2017).
Janssen, S. et al. Phylogenetic placement of exact amplicon sequences improves associations with clinical information. mSystems 3, e00021–18 (2018).
Davis, N. M., Proctor, D. M., Holmes, S. P., Relman, D. A. & Callahan, B. J. Simple statistical identification and removal of contaminant sequences in marker-gene and metagenomics data. Microbiome 6, 226 (2018).
Bolyen, E. et al. Reproducible, interactive, scalable and extensible microbiome data science using QIIME 2. Nat. Biotechnol. 37, 852–857 (2019).
Quast, C. et al. The SILVA ribosomal RNA gene database project: improved data processing and web-based tools. Nucleic Acids Res. 41, D590–D596 (2013).
Faith, D. P. Conservation evaluation and phylogenetic diversity. Biol. Conserv. 61, 1–10 (1992).
Lozupone, C. & Knight, R. UniFrac: a new phylogenetic method for comparing microbial communities. Appl. Environ. Microbiol. 71, 8228–8235 (2005).
Kruskal, W. H. & Wallis, W. A. Use of ranks in one-criterion variance analysis. J. Am. Stat. Assoc. 47, 583–621 (1952).
Fligner, M. A. & Killeen, T. J. Distribution-free two-sample tests for scale. J. Am. Stat. Assoc. 71, 210–213 (1976).
Conover, W. J., Johnson, M. E. & Johnson, M. M. A comparative study of tests for homogeneity of variances, with applications to the outer continental shelf bidding data. Technometrics 23, 351–361 (1981).
Anderson, M. J. A new method for non-parametric multivariate analysis of variance. Austral Ecol. 26, 32–46 (2001).
Oksanen, J. et al. vegan: Community Ecology Package (2018).
Mandal, S. et al. Analysis of composition of microbiomes: a novel method for studying microbial composition. Microb. Ecol. Health Dis. 26, 27663 (2015).
Salter, S. J. et al. Reagent and laboratory contamination can critically impact sequence-based microbiome analyses. BMC Biol. 12, 87 (2014).
Eisenhofer, R. et al. Contamination in low microbial biomass microbiome studies: issues and recommendations. Trends Microbiol. 27, 105–117 (2019).
Lif Holgerson, P., Esberg, A., Sjödin, A., West, C. E. & Johansson, I. A longitudinal study of the development of the saliva microbiome in infants 2 days to 5 years compared to the microbiome in adolescents. Sci. Rep. 10, 9629 (2020).
Imamura, T. et al. The microbiome of the lower respiratory tract in premature infants with and without severe bronchopulmonary dysplasia. Am. J. Perinatol. 34, 80–87 (2017).
Poonam Sharma, P. K. Staphylococcus Aureus – the predominant pathogen in the neonatal ICU of a tertiary care hospital in Amritsar, India. J. Clin. Diagn. Res. 7, 66–69 (2013).
Isaacs, D., Fraser, S., Hogg, G. & Li, H. Y. Staphylococcus aureus infections in Australasian neonatal nurseries. Arch. Dis. Child. Fetal Neonatal Ed. 89, F331–F335 (2004).
Neubauer, V., Griesmaier, E., Ralser, E. & Kiechl-Kohlendorfer, U. The effect of sex on outcome of preterm infants – a population-based survey. Acta Paediatr. 101, 906–911 (2012).
Trembath, A. & Laughon, M. Predictors of bronchopulmonary dysplasia. Clin. Perinatol. 39, 585–601 (2012).
Chu, D. M. et al. Maturation of the infant microbiome community structure and function across multiple body sites and in relation to mode of delivery. Nat. Med. 23, 314–326 (2017).
Acknowledgements
We appreciate the participants and their families for providing samples. We would like to thank members of the N3RO steering committee, including Carmel T. Collins, Maria Makrides, Irmeli Penttila, Thomas R. Sullivan, Robert A. Gibson from the University of Adelaide, and Andrew McPhee from the Women’s and Children’s hospital for their contribution to this study. Furthermore, we would like to thank Irmeli Penttila and Naomi Fink for the help with study design (IP) and sample collection (IP and NF). In addition, we would like to thank Jennifer Young for laboratory troubleshooting, Paul Gooding at the Australian Genome Research Facility for providing sequencing, and Salva Herrando-Perez for statistical support.
Funding
Supported by Australian Government Research Training Program Scholarship (C.A.S.); Women’s and Children’s Hospital Foundation MS McLeod Research Fund Postdoctoral Fellowship (C.T.C.); National Health and Medical Research Council (NHMRC) Fellowships 1132596 (C.T.C.), 1061704 and 1154912 (M.M.), and 1173576 (T.S.); Australian Research Council Future Fellowship FT180100407 (L.S.W.). The N3RO trial was supported by a grant (1022112) from the NHMRC. Open Access funding enabled and organized by CAUL and its Member Institutions.
Author information
Authors and Affiliations
Consortia
Contributions
C.T.C., L.S.W., M.M.: conceptualization. C.A.S.: laboratory and bioinformatic analyses. C.A.S. drafted the initial manuscript. All authors (C.A.S., C.T.C., M.M., T.R.S., L.S.W.) interpreted the data and critically reviewed and revised the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The treatment and control emulsion for the trial were donated by Clover Corporation, Australia. The company had no role in the study design or conduct; in the data collection, management, analysis, or interpretation; or in the preparation, review, or approval of any manuscripts arising from this study.
Consent to participate
Participant consent for the study was described in a previously published study (Collins et al.27).
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Selway, C.A., Collins, C.T., Makrides, M. et al. Variable preterm oral microbiome stabilizes and reflects a full-term infant profile within three months. Pediatr Res (2023). https://doi.org/10.1038/s41390-023-02517-1
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41390-023-02517-1