Abstract
Background
Patients following repair of an esophageal atresia (EA) or tracheoesophageal fistula (TEF) carry an increased risk of long-term cardiopulmonary malaise. The role of the airway microbiome in EA/TEF patients remains unclear.
Methods
All EA/TEF patients treated between 1980 and 2010 were invited to a prospective clinical examination, spirometry, and spiroergometry. The airway microbiome was determined from deep induced sputum by 16 S rRNA gene sequencing. The results were compared to a healthy age- and sex-matched control group.
Results
Nineteen EA/TEF patients with a mean age of 24.7 ± 7 years and 19 age- and sex-matched controls were included. EA/TEF patients showed a significantly lower muscle mass, lower maximum vital capacity (VCmax), and higher rates of restrictive ventilation disorders. Spiroergometry revealed a significantly lower relative performance capacity and lower peak VO2 in EA/TEF patients. Alpha- and beta-diversity of the airway microbiome did not differ significantly between the two groups. Linear discriminant effect size analysis revealed significantly enriched species of Prevotella_uncultured, Streptococcus_anginosus, Prevotella_7_Prevotella_enoeca, and Mogibacterium_timidum.
Conclusion
EA/TEF patients frequently suffer from restrictive ventilation disorders and impaired cardiopulmonary function associated with minor alterations of the airway microbiome. Long-term examinations of EA/TEF patients seem to be necessary in order to detect impaired cardiopulmonary function.
Impact
-
The key messages of the present study are a significantly decreased VCmax and exercise performance, as well as airway microbiome differences in EA/TEF patients.
-
This study is the first to present parameters of lung function and exercise performance in combination with airway microbiome analysis with a mean follow-up of 24 years in EA/TEF patients.
-
Prospective, long-term studies are needed to unravel possible interactions between alterations of the airway microbiome and impaired pulmonary function in EA/TEF patients.
Similar content being viewed by others
Introduction
The importance of the respiratory long-term outcome of patients with EA/TEF (esophageal atresia/tracheoesophagel fistula) has been increasingly recognized in recent years.1,2,3,4,5 The determinants of reduced exercise capacity are still unclear, but comorbidities like tracheomalacia or ventilation disorders may represent underlying causes.2,3,4,6 Children following EA repair more frequently suffer from respiratory symptoms and impairment of cardiopulmonary function compared to the healthy population.1,2,3,5,6 While the negative impact of those health conditions on quality of life in adolescence and adulthood has been demonstrated in several studies, there is no exact idea about the relationship between early childhood disease progression and later pulmonary impairment.2,5,7 Recurrent respiratory tract infections may contribute to impaired pulmonary function.
The common opinion that lungs are sterile has been refuted in several studies identifying up to 2000 bacterial genomes per cm2 lung tissue.8,9,10 In detail, the airways get colonized soon after birth and the airway microbiome continues to evolve during growth.8,11,12 Comparisons of the airway microbiome between infants with chronic lung diseases, such as asthma, chronic obstructive pulmonary disease (COPD), bronchiectasis, or cystic fibrosis, and healthy controls have shown significant differences.12,13,14,15 The importance of the human microbiome has been increasingly recognized in recent years and the microbiome seems to play a substantial role in pulmonary health and immune response.10,13,15,16 The scientific work of airway microbiota composition, however, mainly focuses on chronic pulmonary diseases, and microbiome studies for EA/TEF patients are currently lacking. In addition, it remains unknown whether there is an association between cardiopulmonary performance capacity and alterations of the airway microbiome in EA/TEF patients.
Therefore, the aim of this prospective cohort study was to investigate the cardiopulmonary performance capacity and the airway microbiome of adolescent and adult patients after corrected EA and provide a comparison to a healthy age- and sex-matched control group.
Patients and methods
All consecutive children treated at the Department of Pediatric and Adolescent Surgery of the Medical University of Graz for EA/TEF between 1980 and 2010 were invited to participate in a prospective study consisting of spirometry, cardiopulmonary exercise performance testing, and analysis of the airway microbiome. Exclusion criteria were antibiotic and pre-/probiotic treatment within the past 2 weeks and respiratory or gastrointestinal infections with the past 2 weeks.
This study was performed according to the Declaration of Helsinki. Informed written consent was obtained from all patients and controls and/or their legal guardians. The study was approved by the institutional review board (EK 29-276 ex 16/17).
The medical records of the EA/TEF patients were reviewed retrospectively for associated congenital anomalies, postoperative complications, and consecutive interventions or disease-related comorbidities. Gastroesophageal reflux disease (GERD) was assessed by pHmetry or 24 h impedance pHmetry. All patients were classified according to Spitz risk groups.17 The results were compared to healthy age- and sex-matched control group recruited from the personal environment of the Department’s employees or patients.
All participants were asked to assess their quality of life regarding the presence of abdominal discomfort, swallowing difficulties, regurgitation, acid belching, recurrent cough allergies, or food intolerances.
At the study visit, a 12-lead resting electrocardiography (ECG) was obtained to exclude cardiac arrhythmias and non-invasive blood pressure measurement at rest was performed. Height and body weight (BW) were assessed and body mass index (BMI) was calculated. The body fat in % was determined by the caliper method using a four-site skin fold procedure and estimated according to a standardized table.18,19
Appendicular muscle mass was measured by a multifrequency impedance spectroscopy (CombynTM ECG, Academic Technologies at the Institute of Cardiovascular Medicine GmbH, Graz, Austria) as previously described in the literature.20
The individual physical activity level (PAL) was assessed as previously published and expressed as PAL values between 1.1–1.2 and 2.0–2.5, according to inactive-sedentary, moderate, or vigorous active lifestyle.21
Spirometry
The lung function was measured by a small spirometry (Oxycon Pro® Carl Reiner GmbH, Vienna, Austria) at rest and after exercise with a determination of the maximum vital capacity (VCmax) and the forced expiratory volume in 1 s (FEV 1). The VCmax was expressed as observed over age- and gender-corrected expected maximum vital capacity. The Tiffeneau index was calculated as FEV 1/VCmax. An obstructive ventilation disorder was documented if the Tiffeneau index was <75%, a restrictive disorder if VCmax was <80%. The spirometry was repeated after the cardiopulmonary exercise performance testing to rule out an exercise-induced asthma by showing a decreased Tiffeneau index following intense physical activity.22
Cardiopulmonary exercise performance testing
The cardiopulmonary exercise performance was determined by cardiopulmonary exercise testing (CPET) with a bicycle ergometer (Excalibur Sport®, Lode B.V., Groningen, The Netherlands) and the spirometer in an upright position. Spiroergometry was performed with a gender- and age-adapted protocol using a stepwise load increase as previously published.19 The spiroergometry was carried out to subjective exhaustion or until the participants were unable to maintain the required pedaling speed (cadence) of >60 revolutions per minute (r.p.m.). The exercise phase was followed by a 3-min recovery period of slow pedaling (60 r.p.m.) with the same workload as at the beginning of the test.
The respiratory parameters determined during the bicycle spiroergometry test included the minute ventilation (VE), the oxygen uptake (VO2), the oxygen pulse (O2/heart rate (HR)), the respiratory equivalent for oxygen (EQO2), and the respiratory exchange ratio (RER).19 A RER > 1.10 was used as a criterion to determine that the peak VO2 reflects a peak physiological workload.23
Relative performance capacity was calculated from the achieved maximal wattage in relation to the age- and gender-specific standard values.24 The peak oxygen uptake (peak VO2) was defined as the average VO2 over the past 30 s prior to subjective exhaustion and was expressed in ml/kg/min.24 HR was measured by continuous 12-lead ECG (CombynTM ECG, Academic Technologies at the Institute of Cardiovascular Medicine GmbH Graz, Austria) and oxygen saturation was continuously monitored (finger pulse oximeter Habel Medizintechnik®, Vienna, Austria). Lactate levels were obtained by earlobe sampling (20 µl of blood per measurement were sampled to heparinized capillaries per test) before the test, at the end of each step and after the recovery phase (enzymatically amperometric measurement with a Biosen C_line®, EKF Diagnostics for life, Cardiff, UK).
Airway microbiome sampling, total DNA isolation, 16S amplicon-based library preparation and sequencing
Deep induced sputum samples of the patients and age- and sex-matched controls were harvested as previously described.25 Samples were stored at −80 °C until total DNA isolation. Before nucleic acid extraction, samples were treated with an equal volume of 1 M (100 µg/ml) dl-dithiothreitol solution in water and incubated at 37 °C for 20 min in a water bath. After centrifugation at 4000 × g and 30 min at room temperature, the supernatant was decanted and the cell pellet was resuspended in phosphate-buffered saline (Roth) up to a volume of 500 µl. Two hundred and fifty microliters of the suspension were used for total DNA extraction with the MagNA Pure LC DNA Isolation Kit III (Bacteria and Fungi) (Roche, Mannheim, Germany) according to the manufacturer’s instructions and as published in Klymiuk et al.26 The cell suspension of each sample was mixed with 250 µl bacterial lysis buffer and bead beaten in a MagNA Lyser tube in a MagNA Lyser instrument (Roche, Mannheim, Germany) at 6000 r.p.m. for 30 s twice.
After incubation with lysozyme (25 µl, 100 mg/ml) at 37 °C for 30 min and proteinase K (43.4 µl, 20 mg/ml) digestion at 65 °C for 1 h, samples were heat inactivated at 95 °C for 10 min. After centrifugation for 5 min at 14,000 r.p.m., 200 µl of the sample was loaded to the MagNA Pure LC 2.0 instrument (Roche, Mannheim, Germany) and isolated according to manufacturer’s instructions with the MagNA Pure LC DNA Isolation Kit III (Bacteria, Fungi) (Roche, Mannheim, Germany). Total DNA was eluted in 100 µl elution buffer. For PCR reaction, 5 µl of total DNA was used in a 25 µl PCR reaction in triplicates using a Fast Start High Fidelity PCR System (Roche, Mannheim, Germany) according to Klymiuk et al.26 with the target-specific primers F27-AGAGTTTGATCCTGGCTCAG and R357-CTGCTGCCTYCCGTA.27 Finally, the triplicates of each sample were pooled and the PCR for indexing was performed as described in Klymiuk et al.26 with eight amplification cycles. Five microliters of the indexing PCR of each sample were pooled and the final library was purified by conventional gel electrophoresis and quantified using the QuantiFluor ONE dsDNA Dye (Promega, Hilden, Germany) on a Quantus™ Fluorometer (Promega, Hilden, Germany). Quality control was performed on a BioAnalyzer 2100 instrument (Agilent Technologies Inc. 2020, USA) using a DNA 7500 LabChip. For sequencing, the pool was diluted according to standard procedures and the 6 pM library was sequenced on an Illumina MiSeq desktop sequencer (Illumina, Eindhoven, Netherlands) with 20% PhiX control DNA (Illumina, Eindhoven, Netherlands) and v3 chemistry for 600 cycles in paired-end mode according to the manufacturer’s instructions and FastQ raw reads were used for data analysis. For data analysis, a total of 2,363,371 (per sample minimum 39,514, maximum 81,288, median 63,061) raw sequence reads were used in the Galaxy-based workflow (Medical University of Graz, funded by the Austrian Federal Ministry of Education, Science and Research, Hochschulraum-Strukturmittel 2016 grant as part of BioTechMed Graz). Briefly, raw reads were quality-filtered, de-noised, de-replicated, merged, and checked for chimeras using DADA2 pipeline with standard settings in QIIME2.0.28,29 For taxonomic assignment, SILVA rRNA database 1.3 with 97% identity was used. Low abundant taxa with <0.01% abundance across all samples were eliminated from the analysis.
Statistical analysis
All data were entered into an Excel 2019® (Microsoft Corporation, Microsoft Excel [Internet], 2018, USA) spreadsheet and SPSS Statistics 21© (IBM Corp., released 2012, IBM SPSS Statistics for Windows, Version 21.0, IBM Corp, Armonk, NY) was used for data analysis. A Kolmogorov–Smirnov test was used to assess normal distribution. In case of normal distribution, data are displayed as mean and standard deviation and statistical group comparison was performed using a two-sided, unpaired t test. In case of absent normal distribution, data are displayed as median and interquartile range and a Mann–Whitney U test was used for group comparison. The Fisher’s exact test or the χ2 test was used to compare the categorical data. Statistical significance was defined as p < 0.05.
To interpret and compare taxonomic information from the 16S rRNA datasets online software Calypso® (Version 8.84) was used.30 Species richness was calculated using the Chao1 estimator, the Shannon’s index, and inversed Simpson index. As a measure for beta-diversity redundancy analysis (RDA), color-coded principal component analysis (PCoA) plot and the Anosim (analysis of similarities) Bray–Curtis dissimilarity score were used. Differences in taxa abundance were calculated with Calypso® after log 10 transformation of data. P values were adjusted for multiple testing by false discovery rate. The top 100 most abundant taxa were included in the analysis. Linear discriminant effect size (LEfSe) analysis was performed to detect statistically relevant bacteria in different groups.
Results
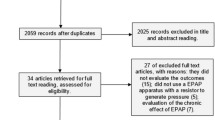
Out of 47 eligible EA/TEF patients treated in the regarded time period, 21 agreed to participate. Three patients refused to take part and 23 patients were unavailable. At the follow-up examinations, one patient had to be excluded because of mental impairment and another one was unable to donate a sufficient sputum sample, leaving 19 patients for further analysis.
EA/TEF patient group
The mean age of the EA/TEF group (10 males, 9 females) was 24.7 ± 7 years (range: 14–40 years). Eleven of them (57.9%) were born preterm. Sixteen patients (84.2%) had a TEF Gross type C, 1 a type B and 1 a type D. One patient had a pure EA (Gross type A). All patients underwent right thoracotomy for EA/TEF repair.
Five patients (26.3%) suffered from at least one additional congenital anomaly: one patient was born with an imperforate anus and a unilateral renal agenesis. Four patients had a congenital cardiac anomaly; three of these patients underwent cardiac surgery: one patient with an atrial septal defect (ASD) type II, one patient with a muscular ventricular septal defect, and another patient with congenital aortic coarctation. The fourth patient with a congenital cardiac anomaly had a not hemodynamically relevant ASD type II. No patient had a birth weight below 1500 g. Only the three patients with hemodynamically relevant cardiac anomalies were therefore classified as risk group II, according to the Spitz risk classification.17
The following postoperative complications were recorded: one patient (5%) had an anastomotic insufficiency, and seven patients (37%) required dilatations because of esophageal strictures.
Clinically relevant tracheomalacia was diagnosed in six patients (32%). Fifteen patients (79%) had a documented GERD, and nine of them required a fundoplication in the further course.
At the study visit, nine EA/TEF patients (47%) complained about abdominal discomfort. Five patients (26%) reported swallowing difficulties. Regurgitation and acid belching were recorded in six patients (32%). The prevalence of recurrent cough was 21% (n = 4) and occurred only in the EA/TEF patient group (χ2 test, p = 0.034). Six patients (32%) mentioned allergies, another three (16%) reported food intolerances.
Control group
The mean age of the age- and sex-matched control group was 24.6 ± 8 years (range: 12–43 years) and consisted of ten male and nine female participants. Two of the control participants were born preterm. There was no statistically significant difference regarding patient age between EA/TEF and control groups (unpaired t test; p = 0.949).
Anthropometric data
There was no statistically significant difference in the amount of PAL and the height, weight, BMI, or body fat percentage between the study and the control group. However, EA/TEF patients showed a significantly lower muscle mass compared to the control group (unpaired t test; p < 0.001) (Table 1).
Spirometry
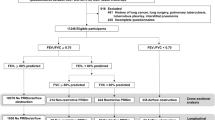
Spirometry revealed a significantly lower VCmax for EA/TEF patients compared to controls (p < 0.001; unpaired t test) (Table 1). Twelve patients showed a reduced VCmax; nine of these had a restrictive and three a combined ventilation disorder. Restrictive ventilation disorders occurred only in the EA/TEF patient group (EA/TEF n = 9 vs. controls n = 0; χ2 test, p = 0.001). Two EA/TEF patients were obstructive and one patient showed an exercise-induced asthma.
Cardiopulmonary exercise performance
All participants had a RER value >1.10. Spiroergometry revealed a significantly lower relative performance capacity (unpaired t test; p = 0.001) and a significantly lower peak VO2 (unpaired t test; p = 0.013) in the EA/TEF group compared to the control group (Table 1). The values for O2/HR and EQO2 were within the normal ranges in both groups. However, the O2/HR was significantly lower (Mann–Whitney U test; p = 0.012) and the EQO2 was significantly higher in the EA/TEF group (Mann–Whitney U test; p = 0.014) (Table 1).
Exclusion of the four EA/TEF patients with congenital cardiac anomalies and their respective controls still revealed a significantly decreased performance capacity (mean 116.7 ± 24.6 vs. 146.1 ± 31.5; unpaired t test, p = 0.008) and peak VO2 (mean 37.6 ± 8.8 vs. 44.7 ± 8.9, unpaired t test; p = 0.036) of EA/TEF patients in the remaining participants.
Patients with restrictive ventilation disorders showed a significantly lower relative performance capacity compared to participants without restrictive ventilation disorders (Mann–Whitney U test; p = 0.029) (Supplement 1). Moreover, neither tracheomalacia nor prematurity had a significant impact on the relative performance capacity or spirometry results in EA/TEF patients (Supplements 2 and 3).
16S rRNA-based airway microbiome
Alpha-diversity—as a measure of the diversity of microbes in a single sample—of the deep induced sputum samples as measured by Shannon index, inversed Simpson’s, and Chao1 did not differ significantly between EA/TEF patients and controls (Fig. 1). Likewise, beta-diversity as a measure of the variation of the species composition between two samples was not significantly different between the two groups in Anosim Bray–Curtis, PCoA, and RDA analysis (Fig. 1). Figure 2 depicts the relative abundances at the phylum and order levels comparing EA/TEF patients and controls.
LEfSe analysis at the species level is shown in Fig. 3. Prevotella uncultured (p = 0.0081), Streptococcus anginosus (p = 0.0078), Prevotella 7 prevotella enocea (p = 0.026), and Mogibacterium timidum (p = 0.047) were significantly enriched in the EA/TEF patient group. Species from Alloprevotella uncultured (p = 0.0079) and Campylobacter uncultured (p = 0.0072) were significantly higher in the control group than in the EA/TEF group (Fig. 3).
The top graph shows histograms of linear discriminant analysis (LDA) effect size (LEfSe) comparison of deep induced sputum samples at the species level comparing EA patients (n = 19) and controls (CTRL) (n = 19). Log-level changes in LDA score are displayed on the x-axis. The graphs below display charts of pairwise comparisons for selected bacteria; only results with p < 0.1 are shown as pairwise comparisons.
Analysis of the airway microbiome composition in EA/TEF patients with a reduced VCmax compared to their respective controls revealed no significant differences of alpha- and beta-diversity (Supplement 4). LEfSe analysis revealed significantly enriched Prevotella uncultured bacterium (p = 0.004) and S. anginosus (p = 0.021) and decreased Alloprevotella uncultured Prevotellacae (p = 0.035) of patients with a reduced VCmax (Supplement 4).
Comparison of the airway microbiome in formerly preterm EA/TEF patients (n = 11) to formerly full-term patients (n = 8) revealed no differences of alpha- and beta-diversity (Supplement 5). LEfSe analysis revealed significantly enriched Campylobacter uncultured bacterium (p = 0.0075), Bergeyella uncultured bacterium (p = 0.041) and Neisseria meningitides (p = 0.03) and significantly decreased Filifactor uncultured bacterium (p = 0.03) in formerly preterm patients.
Discussion
This study presents parameters of lung function and exercise performance in combination with airway microbiome analysis with a mean follow-up of 24 years in EA/TEF patients. The main findings of the present study were a significantly decreased VCmax and exercise performance, as well as minor airway microbiome differences in EA/TEF patients compared to healthy age- and sex-matched controls.
Anthropometric data
Our EA patients had a significantly lower muscle mass compared to the control group. This could be associated with a decreased physical fitness and impaired locomotor function as described in previous studies.6,7,31 Therefore, a routine locomotor function evaluation in school-aged patients and a referral to physiotherapy if required seems to be necessary in EA/TEF patients.
Spirometry
Spirometry revealed that the majority of EA/TEF patients in our study (78.9%) had some sort of ventilatory impairment confirming other reports of long-term pulmonary function in patients following EA/TEF repair.1,2,3,4,5,6,32,33 EA patients showed a significantly lower VCmax than their age- and sex-matched peers in our series. Reduced lung volumes following EA/TEF repair were found frequently in long-term pulmonary function tests.3,4
The proportion of restrictive pulmonary dysfunction in patients after esophageal atresia compared to healthy controls is strikingly high; it was documented in almost half of our EA/TEF patients. Nevertheless, it remains unclear whether an altered lung parenchyma or a restricted thoracic mobility as sequelae of the operation acts as the underlying cause. A comparison with patients who had thoracoscopic surgery would be very interesting in this regard. Furthermore, restrictive ventilatory defects were significantly correlated to the interpouch distance, the duration of postoperative ventilation, recurrent aspiration pneumonia during infancy, and GERD in recently published reports.2
Cardiopulmonary exercise performance
Patients following esophageal atresia repair showed a significantly lower relative performance capacity and peak VO2 compared to controls in bicycle spiroergometry. A significantly lower exercise duration and maximum exercise tolerance has also been found in previous studies.2,3,4,6,33,34 Only three groups have studied the maximum exercise capacity of EA/TEF patients using bicycle ergometry and presented divergent results; none of these studies, however, has prospectively compared the cardiopulmonary exercise results to a healthy age- and sex-matched control group.1,32,35
A strong correlation between ventilation disorders and exercise performance capacity has also been reported in previous studies.2,3,4,6 Furthermore, earlier exhaustion in CPET significantly correlated with the interpouch distance, duration of postoperative ventilation, GERD, and recurrent aspiration pneumonia during infancy.2
The largest number of CPET following EA/TEF repair was published by Toussaint-Duyster et al.6, showing a significantly lower mean endurance time in 55 eight-year-old EA/TEF patients. Seventeen of these patients had been previously tested at the age of 5 years and their exercise capacity was not significantly different, suggesting that a reduced exercise performance may persist over time.
Airway microbiome
In our study, we have examined the airway microbiome following esophageal atresia repair and compared the results to an age- and sex-matched healthy control group for the first time. Alpha- and beta-diversity of the deep induced sputum samples did not differ significantly between EA/TEF patients and controls.
LEfSe analysis between the two groups showed marker bacteria such as Prevotella, S. anginosus, or M. timidum on the species level. These bacteria are considered normal bacterial inhabitants of the oral cavity.36 However, Prevotella and S. anginosus were also found among the respiratory microbiota of patients with chronic lung diseases.37,38 Furthermore, higher relative abundances of Prevotella in airway microbiota have been shown to be associated with an increased host inflammatory response, protracted bacterial bronchitis, asthma development, and a higher risk of progressing COPD.38,39,40 However, obstructive airway disease was uncommon in our study cohort, but recurrent cough occurred in 21% of our patients and may be related to prolonged respiratory tract infection. Recurrent respiratory tract infections are common in infants with EA/TEF, but are described to become less frequent with increasing age.41,42 Investigations of the aerodigestive microbial composition in children with chronic cough revealed that the lower airway microbiota was enriched with Prevotella in the bacterial bronchitis group.43
Airway microbiome studies have also found a high relative abundance of oral microbes, suggesting that recurrent microaspirations might influence the lower airway microbiota.36,43 Especially in EA/TEF patients with a high number of possible aspirations associated with esophageal dysmotility and gastroesophageal reflux, the oral microbiota could have a major impact on the composition of the airway microbiome. For instance, Veillonella, another oral commensal, was found significantly enriched in the lower airway microbiome of orally fed, neurologically impaired children who frequently suffer from aspiration of food, oral secretions, and common reflux of gastrointestinal contents into the airways.43 However, only little is known about the microbial functionality and the kinetics of microbiome changes.44
The microbiota composition can be influenced by exogenous forces, including diets, environmental biodiversity, infection, and antibiotics.45 For instance, antibiotic-induced changes in the microbiome composition may play an important role in the formation of allergies, autoimmunity, and infectious diseases.45 The exclusion criteria of our study were designed to avoid as many of these confounding factors as possible. However, the possibility to positively shape the airway microbiome in EA/TEF patients with for instance pro- or prebiotics is subject for further examinations.
Limitations
Possible limitations of this study include that deep induced sputum samples were used to examine the airway microbiome. There are concerns that these samples may be contaminated by the oropharyngeal microorganisms; however, deep induced sputum appears to be the only method that is ethically justifiable, since the assessment of the pulmonary microbiome is only possible with bronchoalveolar lavage or bronchoscopy, which can only be obtained under anesthesia. Thus, deep induced sputum is frequently used to describe the airway microbiome.10,44
Other possible limitations are the relatively small sample size and the mode of testing by bicycle spiroergometry instead of treadmill testing. Treadmill testing was preferred over bicycle testing in previous EA/TEF studies, because underdeveloped knee extensors could be a limiting factor in children.6 However, in our long-term follow-up, the youngest patient was already 14 years old and the mean age of the patients was 24.7 years. Large sample sizes in orphan pediatric diseases are difficult to achieve and other follow-up studies of EA/TEF patients have included between 8 and 63 patients.1,2,3,4,6,32,33,35
The strengths of our study are the long-term follow-up of 24.7 years and the age- and sex-matched control group.
Conclusion
EA/TEF patients frequently showed restrictive ventilation disorders and impaired cardiopulmonary function and discreet changes of the airway microbiome. Long-term examinations of patients with congenital esophageal atresia consisting of routine locomotor function evaluation, spirometry, and spiroergometry are necessary in order to detect impaired cardiopulmonary function and to prevent the progression of associated complications. Prospective, long-term studies are needed to unravel possible interactions between alterations of the airway microbiome and impaired pulmonary function.
References
Beucher, J. et al. Long-term evaluation of respiratory status after esophageal atresia repair. Pediatr. Pulmonol. 48, 188–194 (2013).
Dittrich, R., Stock, P., Rothe, K. & Degenhardt, P. Pulmonary outcome of esophageal atresia patients and its potential causes in early childhood. J. Pediatr. Surg. 52, 1255–1259 (2017).
Gischler, S. J. et al. A prospective comparative evaluation of persistent respiratory morbidity in esophageal atresia and congenital diaphragmatic hernia survivors. J. Pediatr. Surg. 44, 1683–1690 (2009).
Peetsold, M. G., Heij, H. A., Nagelkerke, A. F., Deurloo, J. A. & Gemke, R. J. Pulmonary function impairment after trachea-esophageal fistula: a minor role for gastro-esophageal reflux disease. Pediatr. Pulmonol. 46, 348–355 (2011).
Sistonen, S. et al. Repaired oesophageal atresia: respiratory morbidity and pulmonary function in adults. Eur. Respir. J. 36, 1106–1112 (2010).
Toussaint-Duyster, L. C. C. et al. Determinants of exercise capacity in school-aged esophageal atresia patients. Pediatr. Pulmonol. 52, 1198–1205 (2017).
Konig, T. T. & Muensterer, O. J. Physical fitness and locomotor skills in children with esophageal atresia—a case control pilot study. Front. Pediatr. 6, 337 (2018).
Dickson, R. P., Erb-Downward, J. R., Martinez, F. J. & Huffnagle, G. B. The microbiome and the respiratory tract. Annu. Rev. Physiol. 78, 481–504 (2016).
Hasegawa, K. & Camargo, C. A. Jr. Airway microbiota and acute respiratory infection in children. Expert Rev. Clin. Immunol. 11, 789–792 (2015).
Mitchell, A. B. & Glanville, A. R. The human respiratory microbiome: implications and impact. Semin. Respir. Crit. Care Med. 39, 199–212 (2018).
Gallacher, D. J. & Kotecha, S. Respiratory microbiome of new-born infants. Front. Pediatr. 4, 10 (2016).
Tracy, M., Cogen, J. & Hoffman, L. R. The pediatric microbiome and the lung. Curr. Opin. Pediatr. 27, 348–355 (2015).
Hahn, A., Warnken, S., Perez-Losada, M., Freishtat, R. J. & Crandall, K. A. Microbial diversity within the airway microbiome in chronic pediatric lung diseases. Infect. Genet. Evol. 63, 316–325 (2018).
Lal, C. V. et al. The airway microbiome at birth. Sci. Rep. 6, 31023 (2016).
Lynch, S. V. The lung microbiome and airway disease. Ann. Am. Thorac. Soc. 13(Suppl. 2), S462–S465 (2016).
Cui, L. et al. The microbiome and the lung. Ann. Am. Thorac. Soc. 11(Suppl. 4), S227–S232 (2014).
Spitz, L., Kiely, E. M., Morecroft, J. A. & Drake, D. P. Oesophageal atresia: at-risk groups for the 1990s. J. Pediatr. Surg. 29, 723–725 (1994).
Durnin, J. V. & Womersley, J. Body fat assessed from total body density and its estimation from skinfold thickness: measurements on 481 men and women aged from 16 to 72 years. Br. J. Nutr. 32, 77–97 (1974).
Windhaber, J. et al. Do anthropometric and aerobic parameters predict a professional career for adolescent skiers? Int. J. Sports Med. 40, 409–415 (2019).
Skrabal, F. et al. The Combyn ECG: adding haemodynamic and fluid leads for the ECG. Part II: prediction of total body water (TBW), extracellular fluid (ECF), ECF overload, fat mass (FM) and “dry” appendicular muscle mass (AppMM). Med. Eng. Phys. 44, 44–52 (2017).
Westerterp, K. R. Physical activity and physical activity induced energy expenditure in humans: measurement, determinants, and effects. Front. Physiol. 4, 90 (2013).
Ali, Z. How to diagnose exercise induced asthma? Asian J. Sports Med. 2, 63–67 (2011).
Mezzani, A., Corra, U., Bosimini, E., Giordano, A. & Giannuzzi, P. Contribution of peak respiratory exchange ratio to peak VO2 prognostic reliability in patients with chronic heart failure and severely reduced exercise capacity. Am. Heart J. 145, 1102–1107 (2003).
Wonisch, M. et al. Spiroergometrie in der Kardiologie—Grundlagen der Physiologie und Terminologie. Austrian J. Cardiol. 10, 383–390 (2003).
Pettigrew, M. M. et al. Association of sputum microbiota profiles with severity of community-acquired pneumonia in children. BMC Infect. Dis. 16, 317 (2016).
Klymiuk, I. et al. The human gastric microbiome is predicated upon infection with Helicobacter pylori. Front Microbiol. 8, 2508 (2017).
McKenna, P. et al. The macaque gut microbiome in health, lentiviral infection, and chronic enterocolitis. PLoS Pathog. 4, e20 (2008).
Bolyen, E. et al. Reproducible, interactive, scalable and extensible microbiome data science using QIIME 2. Nat. Biotechnol. 37, 852–857 (2019).
Callahan, B. J. et al. DADA2: high-resolution sample inference from Illumina amplicon data. Nat. Methods 13, 581–583 (2016).
Zakrzewski, M. et al. Calypso: a user-friendly web-server for mining and visualizing microbiome-environment interactions. Bioinformatics 33, 782–783 (2017).
Harmsen, W. J. et al. Developmental problems in patients with oesophageal atresia: a longitudinal follow-up study. Arch. Dis. Child Fetal Neonatal Ed. 102, F214–F219 (2017).
Olbers, J. et al. Physiological studies at 7 years of age in children born with esophageal atresia. Eur. J. Pediatr. Surg. 25, 397–404 (2015).
Zaccara, A. et al. Physical fitness testing in children operated on for tracheoesophageal fistula. J. Pediatr. Surg. 30, 1334–1337 (1995).
van der Cammen-van Zijp, M. H. et al. Motor-function and exercise capacity in children with major anatomical congenital anomalies: an evaluation at 5 years of age. Early Hum. Dev. 86, 523–528 (2010).
Montgomery, M., Frenckner, B., Freyschuss, U. & Mortensson, W. Esophageal atresia: long-term-follow-up of respiratory function, maximal working capacity, and esophageal function. Pediatr. Surg. Int. 10, 519–522 (1995).
Huffnagle, G. B., Dickson, R. P. & Lukacs, N. W. The respiratory tract microbiome and lung inflammation: a two-way street. Mucosal Immunol. 10, 299–306 (2017).
Asam, D. & Spellerberg, B. Molecular pathogenicity of Streptococcus anginosus. Mol. Oral. Microbiol. 29, 145–155 (2014).
Marsh, R. L. et al. Multiple respiratory microbiota profiles are associated with lower airway inflammation in children with protracted bacterial bronchitis. Chest 155, 778–786 (2019).
Caverly, L. J., Huang, Y. J. & Sze, M. A. Past, present, and future research on the lung microbiome in inflammatory airway disease. Chest 156, 376–382 (2019).
Thorsen, J. et al. Infant airway microbiota and topical immune perturbations in the origins of childhood asthma. Nat. Commun. 10, 5001 (2019).
Kovesi, T. & Rubin, S. Long-term complications of congenital esophageal atresia and/or tracheoesophageal fistula. Chest 126, 915–925 (2004).
Lacher, M., Froehlich, S., von Schweinitz, D. & Dietz, H. G. Early and long term outcome in children with esophageal atresia treated over the last 22 years. Klin. Padiatr. 222, 296–301 (2010).
Kazachkov, M. et al. Aerodigestive dysbiosis in children with chronic cough. Pediatr. Pulmonol. 53, 1288–1298 (2018).
Faner, R. et al. The microbiome in respiratory medicine: current challenges and future perspectives. Eur. Respir. J. 49, 1–12 (2017).
Wypych, T. P., Wickramasinghe, L. C. & Marsland, B. J. The influence of the microbiome on respiratory health. Nat. Immunol. 20, 1279–1290 (2019).
Acknowledgements
Parts of the project were funded by the Doctoral School “Bone, Muscle and Joint” of the Medical University of Graz.
Author information
Authors and Affiliations
Contributions
Each author has met the Pediatric Research authorship requirements.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Consent statement
Informed written consent was obtained from all patients and controls and/or their legal guardians.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, and provide a link to the Creative Commons license. You do not have permission under this license to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Arneitz, C., Windhaber, J., Castellani, C. et al. Cardiorespiratory performance capacity and airway microbiome in patients following primary repair of esophageal atresia. Pediatr Res 90, 66–73 (2021). https://doi.org/10.1038/s41390-020-01222-7
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41390-020-01222-7
This article is cited by
-
Long-term motor activity, cardiopulmonary performance and quality of life in abdominal wall defect patients
Pediatric Research (2023)
-
Habitual physical activity in patients born with oesophageal atresia: a multicenter cross-sectional study and comparison to a healthy reference cohort matched for gender and age
European Journal of Pediatrics (2023)
-
Systematic review of long term follow-up and transitional care in adolescents and adults with esophageal atresia - why is transitional care mandatory?
European Journal of Pediatrics (2023)