Abstract
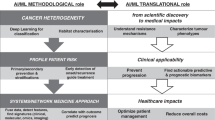
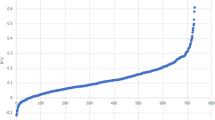
Advances in biotechnology and machine learning have created an enhanced environment for unearthing and exploiting previously unrecognized relationships between genomic and epigenetic data with potential therapeutic implications. We applied advanced algorithms to data from the Cancer Dependency Map to uncover increasingly complex relationships. Specifically, we investigate characteristics of tumor cell lines with varying levels of telomerase reverse transcriptase (TERT) expression in liver cancer. The findings indicate that the effect of CRISPR knockout of Histone Deacetylase 1 (HDAC1) and numerous individual respiratory complex I genes is strongly related to the level of TERT expression, with knockout being particularly efficacious at killing or inhibiting growth of tumor cells with low levels of TERT expression for HDAC1 and high levels for Complex I genes. These findings suggest key biomarkers for therapeutic efficacy and yield novel potential pathways for drug development and provide further proof of principle for the potential of artificial intelligence in oncology.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Bazaga A, Leggate D, Weisser H. Genome-wide investigation of gene-cancer associations for the prediction of novel therapeutic targets in oncology. Sci Rep. 2020;10:10787.
Dezső Z, Ceccarelli M. Machine learning prediction of oncology drug targets based on protein and network properties. BMC Bioinforma. 2020;21:104.
Madhukar NS, Khade PK, Huang L, Gayvert L, Galletti G, Stogniew M, et al. A Bayesian machine learning approach for drug target identification using diverse data types. Nat Commun. 2019;10:5221.
Sherman, J, Verstandig, G, Rowe, JW, Brumer Y. Application of machine learning to large in vitro databases to identify drug–cancer cell interactions: azithromycin and KLK6 mutation status. Oncogene. 2021. https://doi.org/10.1038/s41388-021-01807-4.
Tsherniak A, Vazquez F, Montgomery PG, Weir BA, Kryukov G, Cowley GS, et al. Defining a cancer dependency map. Cell. 2017;170:564.
Travis J. Making the cut. Science 2015;350:1456–7.
Jinek M, Chylinski K, Fonfara I, Hauer M, Doudna JA, Charpentier EA. Programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science, 2012;337:816–21.
Ghandi M, Huang FW, Jané-Valbuena J, Kryukov GV, Lo CC, McDonald R III, et al. Next-generation characterization of the Cancer Cell Line Encyclopedia. Nature. 2019;569:503.
DepMap, Broad (2020): DepMap 21Q1 Public. figshare. Dataset https://doi.org/10.6084/m9.figshare.13681534.v1.
Saretzki G. Extra-telomeric functions of human telomerase: cancer, mitochondria and oxidative stress. Curr Pharm Des. 2014;20:6386–403.
Koh CM, Khattar E, Leow SC, Liu CY, Muller J, Ang WX, et al. Telomerase regulates MYC-driven oncogenesis independent of its reverse transcriptase activity. J Clin Investig. 2015;125:2109–22.
Yuan X, Larsson C, Xu D. Mechanisms underlying the activation of TERT transcription and telomerase activity in human cancer: old actors and new players. Oncogene. 2019;38:6172–83.
Amisaki M, Tsuchiya H, Sakabe T, Fujiwara Y, Shiota G. Identification of genes involved in the regulation of TERT in hepatocellular carcinoma. Cancer Sci. 2019;110:550–60.
Liu T, Yuan X, Xu D. Cancer-specific telomerase reverse transcriptase (Tert) promoter mutations: biological and clinical implications. Genes. 2016;7:38.
Seynnaeve B, Lee S, Borah S, Park Y, Pappo A, Kirkwood JM, et al. Genetic and epigenetic alterations of TERT are associated with inferior outcome in adolescent and young adult patients with melanoma. Sci Rep. 2017;7:45704.
Yuan X, Xu D. Telomerase reverse transcriptase (TERT) iN Action: Cross-talking with Epigenetics. Int J Mol Sci. 2019;20:3338.
Shimada K, Muhlich JL, Mitchison TJ. A tool for browsing the Cancer Dependency Map reveals functional connections between genes and helps predict the efficacy and selectivity of candidate cancer drugs. bioRxiv. 2019. https://doi.org/10.1101/2019.12.13.874776.
Rosen J, Jakobs P, Ale-Agha N, Altschmied J, Haendeler J. Non-canonical functions of Telomerase Reverse Transcriptase—Impact on redox homeostasis. Redox Biol. 2020;34:101543.
Indran IR, Hande MP, Pervaiz S. Htert overexpression alleviates intracellular ros production, improves mitochondrial function, and inhibits ros-mediated apoptosis in cancer cells. Cancer Res. 2011;71:266–76.
Urra FA, Muñoz F, Lovy A, Cárdenas C. The mitochondrial complex(I)ty of cancer. Front Oncol. 2017;7:118.
Acknowledgements
The authors would like to thank Eoin McDonnell, Kris Wood, and Jack Rowe for relevant conversations.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors have roles at Red Cell Partners and Zephyr AI that involve the application of AI to cancer drug discovery. The authors report no known financial interest in HDAC1 or complex I inhibitors.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Sherman, J., Verstandig, G. & Brumer, Y. Application of machine learning to large in-vitro databases to identify cancer cell characteristics: telomerase reverse transcriptase (TERT) expression. Oncogene 40, 5038–5041 (2021). https://doi.org/10.1038/s41388-021-01894-3
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41388-021-01894-3