Abstract
Microbial translocation contributes to persistent inflammation in both treated and untreated HIV infection. Although translocation is due in part to a disintegration of the intestinal epithelial barrier, there is a bias towards the translocation of Proteobacteria. We hypothesized that intestinal epithelial microvesicle cargo differs after HIV infection and contributes to biased translocation. We isolated gastrointestinal luminal microvesicles before and after progressive simian immunodeficiency virus (SIV) infection in rhesus macaques and measured miRNA and antimicrobial peptide content. We demonstrate that these microvesicles display decreased miR-28-5p, -484, -584-3p, and -584-5p, and let-7b-3p, as well as increased beta-defensin 1 after SIV infection. We further observed dose-dependent growth sensitivity of commensal Lactobacillus salivarius upon co-culture with isolated microvesicles. Infection-associated microvesicle differences were not mirrored in non-progressively SIV-infected sooty mangabeys. Our findings describe novel alterations of antimicrobial control after progressive SIV infection that influence the growth of translocating bacterial taxa. These studies may lead to the development of novel therapeutics for treating chronic HIV infection, microbial translocation, and inflammation.
Similar content being viewed by others
Introduction
Microbial translocation—the systemic dissemination of microbes and microbial products absent of sepsis—has emerged as a key contributor to disease progression in people living with HIV (PLWH). In addition to viral infection itself, microbial translocation profoundly stimulates the immune system, promoting a persistent regeneration of activated CD4 + T-cells and over time, exhaustion of the immune system1. Although the initiation of antiretroviral therapy lessens both microbial translocation and immune activation, both phenomena persist and are leading contributors to co-morbidities in treated PLWH1,2. As there is yet no available cure for HIV, a fundamental understanding of the biological processes that lead to microbial translocation is essential for the design of improved prophylaxes.
The human gastrointestinal tract harbors an estimated 1013 bacteria in addition to an incompletely quantified abundance of viruses, fungi, and other eukaryotes3. In healthy individuals, the intestinal epithelium co-exists as part of a symbiotic ecosystem with the commensal microbiome. Both commensals and host epithelia contribute to localized oxygen regulation, provide trans-beneficial nutrients, and produce antimicrobial factors that minimize pathobiont colonization4. In both PLWH and SIV-infected nonhuman primate models for HIV infection, lentiviral infection induces a profound physical and immunological dysfunction of the gastrointestinal tract barrier5,6. In part, this damage stems both from the exquisite sensitivity of epithelial cells to inflammation as well as the loss of IL-22-producing lymphocytes necessary for epithelial regeneration7,8,9,10. Importantly, as the commensal microbiome exists in synchrony with the epithelial barrier, damage to the epithelial barrier destabilizes the intestinal microbiome culminating in the invasion of pathobionts.
Surprisingly and despite overt physical damage to the epithelial barrier, microbial translocation in lentiviral infections is not stochastic. We and others have described a distinct bias in the translocation of bacteria in macaques favoring the dissemination of Proteobacteria and other disease-associated taxa across the breached epithelium and into distal anatomic sites11,12. While the causes for this imbalance remain unclear, lentiviral-associated changes in the intestinal ecosystem may promote metagenomic moreso than phylogenetic changes in the commensal microbiome13,14. Such changes may discriminately prime certain taxa to translocate or otherwise provide a competitive advantage in the lentivirus-remodeled intestinal ecosystem.
Although the capacity of epithelial cells to actively secrete antimicrobial peptides (AMPs) and mucus that regulate bacterial growth is well-characterized, the role of epithelial-derived microvesicles is only recently being appreciated. Microvesicles are small (0–1000 nm), cell-derived extracellular vesicles that serve as a means for the intercellular transport of proteins, nucleic acid, and lipids15. Microvesicles show superior stability under many biological conditions and are thus uniquely disposed to facilitate interkingdom communication within the intestinal lumen. Recently, several publications have begun to define a complex interkingdom correspondence system mediated by microvesicles. Luminal microvesicles contribute an essential and direct role in regulating the composition of the intestinal microbiome, in part through the transmission of miRNA16. Pro-coagulant microvesicles facilitate opsonization of Streptococcus pyogenes after infection, thereby limiting systemic bacterial dissemination17. Reciprocally, extracellular vesicles from some vaginal Lactobacilli inhibit HIV-1 attachment to cellular targets in situ18. Although there has been intense interest in characterizing the composition and contribution of microvesicles relative to HIV transmission and immune activation19,20,21,22, there has been no assessment to date of a potential role for host microvesicles in regulating microbial translocation.
Herein, we characterized intestinal microvesicle concentration, size, cargo, and activity against bacterial taxa in the SIV nonhuman primate HIV model. We demonstrate that macaque luminal microvesicles display decreased miR-28-5p, -484, -584-3p, and -584-5p, and let-7b-3p, as well as increased beta defensin 1 (DEFB1) after SIV infection. We further demonstrate that luminal microvesicles differentially alter the growth of bacteria known to translocate, in vitro. These data describe a novel mode of antimicrobial dysregulation in lentiviral infection and highlight the need to further investigate host:commensal:pathogen communication in lentiviral infection.
Results
SIV infection does not alter the physical phenotype of luminal microvesicles
To assess the potential effects of HIV infection on luminal microvesicles, we began by isolating fecal extracellular vesicles from 13 rhesus macaques —a nonhuman primate model of progressive HIV infection—prior to and during chronic SIV infection. Post-SIV samples were collected during chronic infection from 6 animals, or at AIDS progression from the remaining 7. We observed a wide distribution in microvesicle recovery (Fig. 1a), with no significant differences in either concentration (Fig. 1b) or median diameter (Fig. 1c) by SIV infection status.
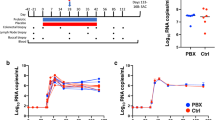
a Luminal microvesicle (MV) size distribution in paired uninfected (black) and SIV-infected (red) macaques. Light lines correspond to individual samples and heavy lines to mean sample distribution by infection status. Luminal microvesicle concentration (b), median diameter (c), CD63 expression (d), and 18 S:16 S transcript ratio (e) from paired uninfected and SIV-infected macaques, with chronic (closed squares) and AIDS (open squares) disease progression denoted. Microvesicle concentration, diameter, and distribution measured by Nanosight. CD63 measured by ELISA. 18S and 16S measured by quantitative reverse transcription (qRT)-PCR, with the ratio reflective of raw threshold cycle (Ct) values. ELISA and qRT-PCR values represent the mean of 3 technical replicates. Significance in b–e assessed by paired, two-way t-test.
Extracellular vesicle categorization remains incompletely defined, with some proposed categorizations relying on size and other categorizations employing surface molecule characterizations15. With a mean composite median diameter of 272.11 nm, many of the isolated vesicles from these macaques can broadly be classified as nonendosomal microvesicles (100–1000 nm), with no significant differences observed by infection status (Fig. 1c). As some of the isolated vesicles fell under 200 nm we considered that SIV infection might shift released microvesicles towards or away from endosomal origin—i.e., exosomes (0–150 nm). To address this, we measured for the presence of the putative exosomal marker CD63. CD63 expression varied among microvesicles irrespective of SIV infection status, with no significant differences observed (Fig. 1d).
Microvesicles are not unique to eukaryotes, with many prokaryotes and viruses releasing either membraned- or enveloped-nanovesicles23,24. As the intestinal lumen is home to upwards of 1013 bacteria3, we considered that significant shifts in bacterial membrane-vesicle shedding may confound our analyses. To assess this potential, we assessed the ratio of 18S (eukaryotic) to 16S (prokaryotic) rRNA in our microvesicle preparations but observed no significant differences by infection status (Fig. 1e). These results suggest that intestinal microvesicle generation remains broadly preserved following SIV infection.
Luminal intestinal microvesicles display altered miRNA content after SIV infection
Epithelial-derived microvesicles have been shown to convey miRNAs capable of regulating bacterial gene transcripts and affecting bacterial growth16. We sought to assess if, like intestinal mucosa itself25,26, SIV infection alters the miRNA profile of luminal intestinal microvesicles. To assess miRNA content, we longitudinally isolated miRNA from intestinal microvesicles isolated from 4 macaques before SIV infection, at acute day 35 post-infection (p.i.), and chronic day 168 p.i. and we subsequently quantified equimolar complementary DNA (cDNA) utilizing human miRNA Arrays. Of 768 assessed targets, 100 were expressed by more than two animals at more than one time point (Supplementary Table S1). By threshold cycle (Ct) value alone, we observed a global upregulation (lower ΔCt) in microvesicle miRNA content during chronic SIV infection (90/100 compared to uninfected; two-way Wilcoxon signed-rank test, p < 0.0001) with several upregulated targets (30/90, p = 0.0348) showing upregulation during acute infection (Fig. 2a). To determine if miRNA profiles distinguished samples by infection status, we further compared these profiles by principal component analysis (PCA). Chronic samples clearly segregated from baseline samples by PC1 (57.5% of sample variance captured, Fig. 2b).
a Heatmap displaying longitudinal host miRNA expression from luminal microvesicles isolated at baseline and days 35 and 168 post-infection (p.i.). Expression (Ct values) assessed by miRNA Array. Data ordered from highest to lowest Ct value at baseline. Represented Ct values are the average of non-replicated, raw Ct values from 4 animals per timepoint, with per-animal values below threshold normalized to 45. b Principal component analysis considering animal, day post-infection, and Ct values of miRNA considered in a. c Normalized expression (ΔCt) of 6 follow-up miRNA from paired uninfected (black) and SIV-infected (red) luminal microvesicle samples as measured by qRT-PCR, with chronic (closed squares) and AIDS (open squares) disease progression denoted. Values are normalized to spike-in control and represent the mean of 3 technical replicates. d miRNA network analysis as assessed by miRnet. Network highlights known genes (circles) targeted by individual miRNA (squares), with shared targets joined by spanning lines. Genes targeted by >2 miRNA (green) and cell-cycle genes (yellow) are uniquely denoted. Significance in c assessed by paired, one-way t-test.
We sought to verify these changes by assessing the top ten differentially regulated miRNAs by quantitative reverse transcription (qRT)-PCR. Here, we normalized miRNA expression to input microvesicle number utilizing a spike-in miRNA control and included an assessment of both sense and antisense transcripts where appropriate. A preliminary assessment of the resultant 15 targets demonstrated that 13/15 were decreased (higher ΔCt) among intestinal microvesicles after SIV infection relative to baseline, six of which were chosen for a more in-depth analysis (Supplementary Fig. 1 and Fig. 2c). Of these six miRNAs, five showed significant differences from baseline during chronic SIV infection—miR-28-5p (p = 0.0188, one-way paired t-test), miR-484 (p = 0.0292), miR-584-3p (p = 0.0484), miR-584-5p (p = 0.0230), and let-7b-3p (p = 0.0123). These differences were not apparent upon AIDS progression (Fig. 2c).
Having seen significant differences in intestinal microvesicle miRNA profile after SIV infection, we next sought to determine which genes these miRNA might target. We analyzed known miRNA interaction data utilizing miRNet and identified targeted networks by Kyoto encyclopedia of genes and genomes (KEGG) enrichment analysis27. Our analysis identified few shared genes between the five miRNAs (Fig. 2d). Only E3-ubiquitin ligase RING finger protein 11 (RNF11) showed shared targeting by >2 miRNA. KEGG enrichment identified several targeted networks, with a significant enrichment for cell-cycle regulating genes (Supplementary Table S2). The targeted cell-cycle genes were not necessarily shared among the five queried miRNAs but rather, were largely dispersed as targets of individual miRNA. These data suggest that rather than targeting a specific biological process per se, these packaged miRNAs broadly reflect components of cell-cycle dysregulation occurring at the intestinal epithelium during SIV infection25,26,28.
DEFB1 is increased among luminal intestinal microvesicle AMPs
In vitro, epithelial-derived microvesicles have been described to transmit AMPs which in turn reduce the viability of infectious microbes in vitro29. Intestinal epithelial cells express an abundance of AMPs—in particular, DEFB1 which is both directly bactericidal and capable of forming bacterial entrapment nets30,31. To characterize the effect of SIV infection on DEFB1 production, we first quantified DEFB1 by ELISA on whole microvesicle lysates (Fig. 3a). We observed a significant increase in DEFB1 concentration upon AIDS progression (p = 0.0013, two-way t-test) and a trend towards increased DEFB1 during chronic infection, with all but 1 animal showing increased expression (p = 0.1390). We additionally examined the expression of Alpha-defensin 1 (DEFA1), Lysozyme (LYZ), Phospholipase A2 group IIA (PLA2G2a), and regenerating islet-derived protein III-gamma (REG3γ) in unpaired, RM MVs. No significant differences were observed by infection status (Supplementary Figure 2).
a Beta-defensin 1 (DEFB1) load in luminal microvesicles as measured by ELISA from paired uninfected (black) and SIV-infected (red) macaques, with chronic (closed squares) and AIDS (open squares) disease progression denoted. b Normalized DEFB1 expression (ΔCt) from paired uninfected and chronic SIV-infected rectal biopsy (RB, left) and jejunal biopsy (Jej, right) macaque homogenates as measured by qRT-PCR. Rectal (c) and jejunal (d) homogenate hypoxia-inducible factor 1-alpha (HIF1A, left) and peroxisome proliferator-activated receptor-gamma (PPARG, right) expression from paired uninfected and chronic SIV-infected macaques as measured by qRT-PCR. e Association between jejunal homogenate DEFB1 and PPARG in chronic SIV-infected macaques. All values represent the mean of 3 technical replicates. qRT-PCR ΔCt values are normalized to housekeeping gene (monkey) beta-glucuronidase (mGusB). Significance in b–d assessed by paired, two-way t-test. Association in e assessed by Pearson correlation.
To determine if increased microvesicle expression of DEFB1 after SIV infection might result from altered expression by parental cells, we quantified DEFB1 mRNA in paired intestinal tissue homogenates, before and after chronic SIV infection (Fig. 3b). Consistent with previous reports, we noted higher expression (lower ΔCt) of DEFB1 within the large intestine as compared to the small intestine32,33. No differences were seen in DEFB1 expression in rectal homogenates by SIV infection status; however, a significant increase (lower ΔCt) was observed in jejunal homogenates (p = 0.0075). We next assessed whether increased DEFB1 transcription and microvesicle packaging might stem from alterations in the DEFB1 transcriptional network. The DEFB1 promoter has response elements for both hypoxia-inducible factor 1-alpha (HIF-1α) and peroxisome proliferator-activated receptor-gamma (PPARγ)34,35. Mirroring the absence of changes in rectal DEFB1 expression, no SIV-induced differences were observed for rectal homogenate HIF1A or PPARG (Fig. 3c). Jejunal homogenates, however, demonstrated a significant decrease (higher ΔCt) in PPARG expression alone after chronic SIV-infection (p = 0.0069; Fig. 3d). Within the small intestine, DEFB1 and PPARG expression trended towards an inverse correlation (Pearson r = −0.6720, p = 0.0679, Fig. 3e). Collectively these results demonstrate a perturbation in microvesicle DEFB1 and jejunal DEFB1 expression after SIV infection, coincident with a disruption in jejunal PPARG.
Natural host luminal intestinal microvesicles are not perturbed as in progressive hosts
Natural hosts of SIV have co-evolved with SIV and do not progress to AIDS or manifest microbial translocation or immune activation36. We postulated that—reflecting the absence of AIDS progression—sooty mangabeys would not exhibit deviations in intestinal microvesicle phenotype or content after SIV infection. To assess this, we isolated fecal microvesicles from uninfected and chronically SIV-infected sooty mangabeys. As with our macaque cohort, mangabeys exhibited no significant differences in basic measures of microvesicle phenotype in response to SIV infection—neither distribution, concentration, diameter, CD63 expression, nor 18S:16S ratio (Fig. 4a-e) differed. Investigation into microvesicle levels of DEFB1 showed no significant differences by SIV status in mangabeys (Fig. 4f). Surprisingly, however, a cross-species comparison of SIV-uninfected mangabeys versus macaques demonstrated that mangabeys naturally express significantly higher levels of luminal microvesicle DEFB1 (Fig. 4g, unpaired two-way t-test, p = 0.0003). This difference was not skewed by the two highly expressing samples as their removal did not negate significance. miRNAs differentially regulated in macaques showed no significant differences relative to SIV infection status in mangabeys (Fig. 4h), with no significant cross-species differences noted in SIV-uninfected animals (direct comparisons not shown). These results confirm our hypothesis that SIV-infected natural hosts show a preservation of microvesicle cargo identified as differentially regulated during progressive infection, whilst naturally expressing higher levels of DEFB1.
a Luminal microvesicle size distribution in unpaired uninfected (black) and SIV-infected (green) mangabeys. Light lines correspond to individual samples and heavy lines to mean sample distribution by infection status. Luminal microvesicle concentration (b), median diameter (c), CD63 concentration (d), 18S:16S transcript ratio (e), and DEFB1 concentration (f) from unpaired uninfected and SIV-infected mangabeys. Microvesicle concentration, diameter, and distribution measured by Nanosight. CD63 and DEFB1 measured by ELISA. 18S and 16S measured by qRT-PCR, with the ratio reflective of raw Ct values. g Direct comparison of DEFB1 expression in uninfected rhesus macaques (RM, circles) and sooty mangabeys (SM, triangles) as assessed by ELISA. h Expression (ΔCt) of 6 miRNAs from unpaired uninfected and SIV-infected mangabey luminal microvesicle samples as measured by qRT-PCR with values normalized to spike-in control. ELISA and qRT-PCR values represent the mean of three technical replicates. Significance in b–h assessed by unpaired, two-way t-test.
Luminal intestinal microvesicles isolated after progressive SIV infection differentially regulate the growth of translocating bacterial taxa
We postulated that the differences in luminal intestinal microvesicle cargo observed after SIV infection might uniquely alter the growth of commensal microbes. To address this, we co-cultured paired luminal microvesicles with bacteria known to translocate in SIV-infected macaques and assessed bacterial growth in vitro. Specifically, we assessed the growth of the two commensal taxa we commonly observe to translocate—Lactobacillus salivarius and Escherichia coli—and two less common translocators within the same bacterial classes—Enterococcus gallinarum and Klebsiella pneumoniae, respectively.
Of our co-cultures, only L. salivarius showed altered growth in the presence of microvesicles co-cultured aerobically, with further evidence of differential growth attributable to SIV infection status (Fig. 5a). No differences were seen by SIV status when co-cultures were assessed anaerobically (Fig. 5a). To assess whether the apparent differential growth seen by infection status with aerobically grown L. salivarius was relevant, we calculated the generation-time (doubling-time) of bacteria in the presence of microvesicles isolated from SIV- versus SIV + animals (Fig. 5b). We observed a significant increase in generation-time by SIV-infection status (p = 0.0389), driven largely by delayed replication in the presence of microvesicles isolated after AIDS progression (p = 0.0397). The microvesicle effect on L. salivarius growth was dose-dependent, suggesting the action of a discrete, transmissible agent (Fig. 5c).
a Growth curves of translocating bacterial isolates in aerobic (top) or anaerobic (bottom) conditions upon co-culture with luminal microvesicles (1:2 dilution) isolated from paired uninfected and SIV-infected macaques. Growth of L. salivarius (n = 12 pairs), Escherichia coli (n = 8), Enterococcus gallinarum (n = 11), and Klebsiella pneumoniae (n = 8) measured by spectrophotometer at OD600. Values represent the mean of 8–12 paired biological and 3 technical replicates ±SEM. b Generation time of L. salivarius aerobically co-cultured in a, with chronic (closed squares) and AIDS (open squares) disease progression denoted. c Growth curves of L. salivarius aerobically co-cultured with serial dilutions of luminal microvesicles isolated from uninfected macaques. Values represent the mean of four paired biological replicates ±SEM. Significance in growth generation times in b, c assessed by paired, two-way t-test.
Luminal intestinal microvesicles do not influence colonic epithelial cell proliferation or apoptosis during progressive SIV infection
In addition to altering the translocation potential of intestinal microbes, luminal microvesicles—whether host or microbe derived—may contribute to physical alterations of the gastrointestinal epithelial barrier. To address this possibility, we co-cultured human colonic organoids with paired luminal microvesicles isolated before and after SIV infection and assessed cellular proliferation and epithelial apoptosis by immunofluorescence staining for Ki-67 and active caspase-3, respectively. Microvesicles from SIV-infected or uninfected animals did not influence expression of either Ki-67 (Fig. 6a, b) or active caspase-3 (Fig. 6a, c). Thus, our results do not indicate a contribution of luminal microvesicles in epithelial barrier dysfunction.
a Representative immunofluorescence images of colonic organoids cultured with or without microvesicles isolated from paired uninfected and SIV-infected macaques, stained for Ki-67, DAPI, active caspase-3, and EPCAM. b Percent Ki-67+ of DAPI + nuclei of colonic organoids stained as in a, with chronic (closed squares) and AIDS (open squares) disease progression denoted. c Intensity (sum/volume) of active caspase-3 of colonic organoids stained as in b. Values represent the mean of 5 technical replicates from four control or seven paired biological samples ±SEM.
Discussion
Microbial translocation has emerged as a key contributor to disease progression in both treated and untreated progressive lentiviral infections. Although the effects of microbial translocation on systemic inflammation have been extensively characterized, mechanisms contributing to translocation remain unclear. In particular, it is uncertain why there is a bias in translocation. Herein, we postulated that HIV infection may result in altered intestinal microvesicle cargo which may, in turn, alter their biological influence on commensal stasis. Utilizing the rhesus macaque SIV model, we demonstrate that luminal microvesicles have significantly altered miRNA and DEFB1 content following progressive SIV infection and further show that SIV infection differentially influences the ability of microvesicles to alter the growth of the commensal L. salivarius ex vivo.
Microvesicles have recently gained appreciation as an important method of intracellular communication. It is unclear whether microvesicles have evolved as an intentional method of communication or whether these effects are a byproduct of cellular disposal; regardless, microvesicle cargo plays an essential role in disease progression. In SIV and HIV infection respectively, plasma microvesicles activate resting CD4 + T-cells and induce interferon-stimulated gene responses in monocytes in vitro20,22. Importantly, microvesicle release is not limited to eukaryotes23,24. Although our methodology did not separate host from non-host microvesicles, the preserved physical characteristics of our assayed microvesicles after SIV-infection (Figs. 1 and 4) in addition to our direct assessment of primate targets suggest that our findings are specific to the nonhuman primate lentiviral response.
Within the intestinal mucosa of SIV-infected macaques, a significant decrease in miRNA production has been noted, with a particular decrease in miRNA targeting epithelial homeostasis25,26. Herein, we demonstrate that luminal microvesicle miR-28-5p, -484, -584-3p, -584-5p, and let-7p-3p miRNA are significantly decreased after progressive SIV infection (Fig. 2). The predicted enriched targeting of cell-cycle genes among these significantly altered miRNA is complementary to the pattern described by Gaulk et. al25. and is consistent with numerous observations demonstrating intestinal epithelial dysregulation6,26,28. This is further corroborated by shared targeting of RNF11, which negatively regulates TNF- and LPS-mediated NF-κB signaling37. Collectively, the results of our luminal microvesicle miRNA profiling reflect a landscape of intestinal epithelial dysfunction.
Constitutively expressed by colonic epithelial cells, DEFB1 has been shown to restrict microbial growth in a redox-dependent manner30,31,32. Whereas in the hypoxic intestinal epithelium reduced DEFB1 forms entrapment nets which restrict the movement and growth of both gram-negative and -positive bacteria, oxidized DEFB1 shows bactericidal activity only against gram-negative taxa. Although no prior link to microbial translocation has been described in HIV-1 infection, the early type I interferon response has been shown to upregulate monocyte DEFB138 and polymorphisms in DEFB1 correlate with altered risk of HIV infection and mother-to-fetus transmission39,40. We observed that after progressive SIV infection, luminal microvesicles displayed an increase in DEFB1, which was paired with an increase in jejunal but not rectal DEFB1 transcripts (Fig. 3). Although the intestinal tract forms a continuous tract, regional variations in microbe colonization are apparent, with the small intestine being relatively enriched for Proteobacteria41. A jejunal-specific, post-infection change in DEFB1 production may indicate that the profile and magnitude of microbial translocation is disparate across the intestinal tract.
Importantly, when we explored potential mechanisms of DEFB1 transcriptional control, we observed a specific decline in jejunal PPARG coincident with a rise in jejunal DEFB1. In addition to regulating DEFB1 transcription, PPARγ drives the energy metabolism of epithelial cells toward β-oxidation and hypoxia in a commensal- and butyrate-dependent manner42,43. Although there has been no direct assessment of oxygenation within the lentiviral-infected intestinal tract, there is evidence to suggest that dysregulation has been overlooked. The transcriptome of colonic epithelial cells after SIV infection reveals a strong disruption in genes associated with oxidative injury and stress—including PPARG26—and intestinal fatty acid metabolism is highly modulated in SIV infection, reflecting a defect in mitochondrial fatty acid β-oxidation44. A rigorous assessment of intestinal hypoxia in the context of lentiviral infections will be essential to fully inform the extent of intestinal dysfunction present in HIV infection.
We observed that the miRNA and DEFB1 microvesicle alterations that distinguish uninfected from SIV-infected macaques are preserved in mangabeys post-SIV infection (Fig. 4). However, irrespective of SIV infection, mangabeys express levels of microvesicle DEFB1 comparable to SIV-infected macaques. As natural hosts have evolved to live with SIV, constitutively high levels of DEFB1 may limit pathobiont colonization, thereby restraining translocation and intestinal inflammation6. This is in keeping with findings by Bochart et. al. describing lower levels of fecal pathobionts in mangabeys as compared to macaques, irrespective of host infection status (Manuscript submitted—see Bochart bioRxiv 780825). Studies comparing captive to semi-captive nonhuman primate species demonstrate strikingly convergent intestinal microbiomes45 (see Bochart bioRxiv 780825). While we cannot completely discount a contribution of diet and housing towards the differences noted in DEFB1—particularly with regards to the maintenance of keystone species46,47—all animals were fed the same diet throughout the study. Taken together, we believe diet contributions to the differences we observe to be minor.
Although the biological function of microvesicles has canonically been studied within the context of a particular host (i.e., host–donor to host–target), microvesicles are capable of mediating inter-kingdom communication. We specifically investigated the role of microvesicles in modulating the growth of four species we have found to translocate in SIV-infected macaques. Of the four species investigated, only commensal L. salivarius demonstrated differential aerobic growth from controls, with SIV-infection stunting the growth advantage conferred by microvesicles isolated from SIV-uninfected macaques (Fig. 5). That microvesicles enhance the growth of the commensal L. salivarius may suggest that there is a symbiotic advantage to supporting the luminal growth of this particular commensal. Although widely touted as a probiotic, little is known about the specific role of L. salivarius in primate health. In vitro, L. salivarius exhibits specific antagonism towards pathogen Campylobacter jejuni48 and Bactofencin A produced by some L salivarius isolates displays bactericidal activity against S. aureus and Listeria spp49. As we did not see SIV-associated differential growth in anaerobic conditions, these results would be relevant upon the loss of physiologic hypoxia. A wider assessment of the contribution of microvesicle cargo to bacterial growth and function is necessary to fully characterize mechanisms of microbiome regulation in primate health and disease.
Microvesicles carry an array of cargo capable of influencing bacterial growth and function, including DNA, RNA (mRNA, miRNA, lncRNA, cirRNA), enzymes, metabolites, and lipids. Our study examined the direct influence of NHP microvesicles on bacterial growth with a particular focus on the potential roles of AMPs and miRNA. AMPs are, by-and-large, directly bactericidal, integrating into bacterial walls or membranes and causing their destabilization50. In contrast, miRNAs can either inhibit or promote growth at the transcriptional level. For example, co-culture of Fusobacterium nucleatum with miR-515-5p and E. coli with miR-1226-5p increase their growth in vitro by specifically increasing 16S:23S and yegH transcripts, respectively16. There are additional indirect mechanisms that might contribute to growth and colonization restriction in vivo. Neutrophil microvesicles facilitate formation of extracellular nets51 to impede the growth of Staphylococcus aureus. Intracellular and surface-bound enzymes are capable of degrading extracellular matrix, thereby altering mucosal density and biofilm integrity52,53. Furthermore, with their ability to transfer RNAs, microvesicles can modify host-interfacing bacterial functions, having secondary effects on bacterial growth or colonization54. We did not observe significant differences by SIV infection status on epithelial proliferation or apoptosis (Fig. 6); however, epithelial immunological function extends beyond the maintenance of a physical barrier4,55. Further investigation into the roles of intestinal microvesicles in regulating bacterial colonization and invasion at the epithelial barrier will undoubtedly lead to the identification of therapeutic targets for stemming microbial translocation.
Herein, we demonstrate that luminal microvesicles have significantly altered miRNA and DEFB1 content following progressive SIV infection and further show that progressive infection differentially influences the ability of luminal microvesicles to regulate the growth of the commensal L. salivarius ex vivo. These results reveal a novel mechanism by which primate hosts may regulate microbial translocation in HIV and SIV infection. Studies such as ours illuminate naturally occurring mechanisms of host: commensal regulation and significantly contribute to the ongoing research and development of synthetic microvesicles and nanovesicles in disease management56. We believe that a wider assessment of the contribution of microvesicle cargo to microbial regulation is warranted and essential for fully elucidating the mechanisms which contribute to pathogenesis in HIV infection.
Methods
Animal selection and care
Animals selected for inclusion in this study were chosen from among cryogenically banked fecal and tissue samples. SIV-infected rhesus macaques (Macaca mulatta) were infected with SIVmac239, SIVmac239X57, or SIVmac251 whilst SIV-infected sooty mangabeys (Cercocebus atys) were experimentally infected with SIVsmm prior to 2007 (Supplementary Table S3). Selected animals had not received antibiotics within 6 months and were greater than 2.5 years from experimental antiretroviral therapy. The National Institute of Allergy and Infectious Diseases (NIAID) Division of Intramural Research Animal Care and Use Program, as part of the National Institute of Health (NIH) Intramural Research Program, approved all experimental procedures pertaining to the macaques (protocol LVD 26). The Yerkes National Primate Research Center (YNPRC) institutional animal care and use committee (IACUC), as part of Emory University, approved all experimental procedures pertaining to the mangabeys (protocol PROTO201700393).
The macaques in this study were housed and cared for at the NIH Animal Center, under the supervision of the Association for the Assessment and Accreditation of Laboratory Animal Care (AAALAC)-accredited Division of Veterinary Resources and as recommended by the Office of Animal Care and Use Nonhuman Primate Management Plan. The mangabeys in this study were housed and cared for at the Yerkes Main Station, under the supervision of the AAALAC-accredited Division of Animal Resources. Care at both facilities met the standards set forth by the Animal Welfare Act, Animal Welfare Regulations, United States Fish and Wildlife Services Regulations, as well as The Guide for the Care and Use of Laboratory Animals (8th Edition).
The physical conditions of the animals were monitored daily. Animals in this study were exempt from contact social housing due to scientific justification, per respective IACUC protocols, and were housed in noncontact, social housing where primary enclosures consisted of stainless-steel primate caging. The animals were provided continuous access to water and offered commercial monkey biscuits twice daily as well as fresh produce, eggs, and bread products and a foraging mix consisting of raisins, nuts, and rice. Enrichment to stimulate foraging and play activity was provided in the form of food puzzles, toys, cage furniture, and mirrors.
Sample collection
Frozen feces and intestinal tissue homogenates were accessed for this study. Approximately 1 mL of macaque stool was originally collected fresh from each animal by inserting a sterile swab 2 cm into the rectum and spinning to collect available sample. Approximately 1 g of mangabey feces was collected opportunistically from all animals within one hour of defecation on grossly uncontaminated surfaces (areas with no other preexisting urine, water, food, natural substrates, or feces). Collected feces were snap-frozen and stored at −80 °C until accession. To obtain intestinal biopsies, animals were sedated with 3–4 mg/kg telazol administered intramuscular and isoflurane gas by intubation, to effect. Successful anesthetization was monitored by response to stimuli. For rectal biopsies, fecal material was removed from the rectum and biopsies obtained with biopsy forceps. Jejunal biopsies were obtained by video-guided endoscopy. 10 pinch biopsies were obtained per animal. Biopsies were maintained in RPMI-1640 medium for transport, rinsed twice with PBS, and transferred to soil-grinding Precellys tubes (Bertin Technologies, France). Samples were homogenized in 1 mL TRIzol (ThermoFisher Scientific, USA) at room temperature on a Precellys 24 homogenizer at 5000 revolutions per minute (RPM) in four successive 20 s intervals and immediately transferred to −80 °C for storage.
Microvesicle isolation from stool
Frozen feces were allowed to thaw to semi-solid at room temperature and vortexed into slurries with 3 mL cold PBS per gram stool (minimum 10 mL per isolation). Fecal slurries were clarified by sequential centrifugation at room temperature at 300 × g for 10 min and at 2000 × g for 15 min. Clarified supernatants were filtered at 70 μm and microvesicles isolated using the exoEasy Maxi kit (Qiagen, Germany), with a final elution at 300 μL Buffer XE/g of starting feces. Microvesicle concentration, diameter, and distribution were obtained by NanoSight (Malvern Panalytical, Netherlands). Microvesicles were standardized to 3 × 1010 microvesicles/mL in Buffer XE (Qiagen) and frozen in individual aliquots for subsequent analyses.
RNA extraction and quantification
miRNA analyzed by Array were isolated from microvesicles using the miRCURY RNA isolation kit (Exiqon, Denmark). cDNA was generated from 150 ng of purified RNA using the TaqMan MicroRNA Reverse Transcription Kit (ThermoFisher Scientific) with Megaplex RT Primers, Human Pool Set 3.0. cDNA pools were quantified using the TaqMan Array Human MicroRNA A + B Cards Set 3.0 (ThermoFisher Scientific) with TaqMan Universal PCR Master Mix II, no UNG.
For individual miRNA quantification, one 500 μL thawed aliquot of microvesicles (1.5 × 109 microvesicles) was spiked with 10 μL of 10μM Caenorhabditis elegans cel-miR-39-3p mimic (478293_mir; ThermoFisher Scientific). miRNA was isolated from the entirety of the microvesicle preparation using either the miRCURY RNA isolation kit or Qiagen miRNAeasy kit. miRNA was converted to cDNA using the TaqMan Advanced miRNA cDNA synthesis kit (ThermoFisher Scientific), with miR-Amp pre-amplification. miRNAs of interest were quantified using transcript specific TaqMan Advanced miRNA Assays (ThermoFisher Scientific) with TaqMan Fast Advanced Master Mix. Resultant Ct values were normalized to cel-miR-39-3p.
Microvesicle mRNA was quantified from the generated miRNA preparations (above). For intestinal mRNA, TRIzol-preserved samples were thawed and treated with 200 μL chloroform to separate nucleic acid into an aqueous phase. Following separation, Total RNA was isolated from the aqueous phase using the MagMAX-96 total RNA isolation kit (ThermoFisher Scientific). For both microvesicle and intestinal transcript quantification, cDNA was generated using the SuperScript III reverse transcriptase kit (ThermoFisher Scientific). qRT-PCR for microvesicle 16S and 18S transcripts were preformed using SYBR green and reported as a 18S:16S ratio. qRT-PCR for intestinal DEFB1, HIF1A, and PPARG were preformed using transcript-specific Gene Expression Assays (ThermoFisher Scientific) with TaqMan Fast Advanced Master Mix and were normalized to housekeeping gene (monkey) beta-glucuronidase (mGusB) utilizing a custom-designed mGusB TaqMan Gene Expression Assay (ThermoFisher Scientific). Noncommercial/custom primers and probes were as follows: 16S forward primer 5′-GGACTACGGATTAGATACCCTGGTAGTCC-3′, 16S reverse primer 5′-CTTGTGCGGGTCCCCGTCAAT-3′, 18S forward primer 5′-TAGAGGGACAAGTGGCGTTC-3′, 18S reverse primer 5′-CGCTGAGCCAGTCAGTGT-3′, mGusB forward primer 5′-CTCATTTGGAATTTTGCCGATT-3′, mGusB reverse primer 5′-CCGAGTGAAGATCCCCTTTTTA-3′, and mGusB probe 5′-TGAACAGTCACCGACGAGAGTGCTGG-3′.
All miRNA and total RNA were isolated per the specific manufacturers’ protocols, with optional on-column recombinant DNase treatment as appropriate. miRNA and RNA concentration and purity (A260/280 ≥1.8) were assessed by spectrophotometer prior to cDNA generation and normalized to 50–100 ng/μL in PCR-grade H2O. miRNA Array quantifications were carried out on the ViiATM 7 System with the Applied Biosystems QuantStudio Real-Time PCR Software (ThermoFisher Scientific), using the recommended TaqMan MicroRNA Array cycling parameters. Individual miRNA/mRNA assessments were assessed in triplicate on the Applied Biosystems StepOne Plus Real-Time PCR System with StepOne Software (ThermoFisher Scientific), using the recommended TaqMan Fast Advanced or SYBR Green cycling parameters, as appropriate.
ELISAs
ELISAs were custom designed using cross-reactive, anti-human antibodies and recombinant human or macaque proteins or peptides (Supplementary Table 4). Samples were measured in triplicate on 7.5 × 10^7 microvesicles/replicate. Samples and standards were lysed on ice for 5 min in RIPA buffer with 1% v/v Calbiochem Protease and Phosphatase Inhibitor Cocktails (Millipore, USA), with further dilutions carried out in PBS. All washes were performed twice with 0.05% PBS-T (Tween-20) at room temperature on an orbital shaker.
CD63 and DEFB1 were measured as sandwich ELISAs. EIA/RIA plates (Corning, USA) were coated with capture antibody diluted in PBS or BupH™ (ThermoFisher Scientific) and incubated overnight with gentle rocking at 4 °C. Capture-coated plates were washed and blocked for 1 h at 37 °C with Blocking Buffer (0.1% PBS-T with 5% bovine serum albumin). Blocked plates were washed and incubated with freshly lysed samples or standards for 1 h at 37 °C. Antigen-loaded plates were washed and incubated with detection antibody for 1 h at 37 °C. Detection-loaded plates were washed and incubated with horseradish peroxidase-conjugated secondary antibodies for 1 h at 37 °C. Secondary-loaded plates were washed and treated with room temperature TMB (3,3′,5,5′- tetramethylbenzidine) substrate and Stop Solution (Biolegend, USA). Plates were immediately read by spectrophotometer (Molecular Devices SpectraMax i3 with SoftmaxPro v6.5.1, USA) at 450 nm for analyte detection, with background subtraction measurements taken at 570 nm.
DEFA1, LYZ, PLA2G2a, and REG3γ were measured as direct ELISAs. Freshly lysed samples or standards were added to EIA/RIA plates and incubated overnight with gentle rocking at 4 °C. Antigen-coated plates were blocked by the direct addition of AbGuard Plate Stabilizer (BioRad, USA) for 10 min at room temperature with gentle rocking. Plate contents were dumped. Plates were gently patted dry and incubated for 2 h at 37 °C in a dry chamber. Blocked plates were directly incubated with detection antibody for 1 h at 37 °C. All subsequent steps were performed as for the sandwich ELISAs, with the exception of the use of TMB at 4 °C.
miRNA target predictions
To identify genes targeted by specific miRNA (Fig. 2d), miRNA were queried against human target genes using the miRNA-centric network visual analytics platform, miRNet (v2.0, https://www.mirnet.ca/miRNet/home.xhtml)27. From all identified targets, enriched gene sets were identified by hypergeometric test using the KEGG database option in miRnet. miRnet was utilized with default parameters and no identified targets were excluded from publication.
Bacterial co-cultures
Bacterial isolates were chosen from among species isolated and amplified from the tissues of post-mortem, SIV-infected macaques. Isolates were identified by 16S rDNA sequencing as in Ortiz et al58. and stored as pure cultures in glycerol at −80 °C until use. Bacterial isolates are available upon request. For aerobic growth assessment, frozen glycerol isolates were chipped/scraped into 5 mL Brucella broth (Anaerobe Systems, USA) and incubated overnight at 37 °C with shaking at 225 RPM. Overnight bacterial cultures were diluted 1:10 in fresh Brucella broth and plated in triplicate in clear non-binding F-bottom 96-well plates (Greiner Bio-One, Austria) with 20 μL microvesicle dilutions or PBS (200 μL total/well). Microvesicle dilutions began at 1:2, representing 3.75 × 108 microvesicles/well. Plates were sealed with optical adhesive and incubated at 37 °C with 200 RPM shaking until growth curves plateaued (5–36 h). OD600 measurements were taken by spectrophotometer (Synergy HTX Multi-Mode Reader; BioTek, USA) every 5 min during incubation. For cultures assessed over more than one run, read times were initialized (normalized baseline) upon crossing OD600 0.12. For anaerobic growth assessment, frozen isolates were chipped/scraped into 5 mL Brucella broth, transferred into an anaerobic chamber (85% N2, 10% H2, 5% CO2), and incubated overnight at 37 °C. 3.75 × 108 microvesicles (10 μL) or Buffer XE were added to a clear non-binding F-bottom 96-well plate under aerobic conditions and transferred into the anaerobic chamber before addition of bacteria. Overnight bacterial cultures were diluted 1:100 in fresh Brucella broth and added to each plate (200 μL total/well), in duplicate replicates. Plates were sealed and incubated at 37 °C anaerobically until optical density was approximately 0.5, 0.8, and 1.0, as estimated by visual inspection and prior growth curve calibration. Plates were removed from the anaerobic chamber, unsealed, and OD600 measurements taken by spectrophotometer. Generation time (tgen) was measured as follows: tgen = (t In2)/(In Xt − In X0) where t is the elapsed time during which growth is measured and X0 and Xt are the number of bacteria at times zero and t, respectively, corresponding to readings taken at entry to and exit from log phase, respectively.
Expansion of human colon organoids
3dGRO human iPSC-derived colon organoids (MilliporeSigma) were thawed and washed with 9 mL of colon organoid growth media59 containing Advanced DMEM F12, 1X B27, 2 mM Glutamax, 100 U/mL Penicillin, and 100 μg/mL Streptomycin (ThermoFisher Scientific), 3 μM CHIR99021 and 300 nM LDN193189 (MilliporeSigma), 100 ng/mL human EGF (R&D, USA) and 10 μM ROCKi (StemCell Technologies, USA). The organoids were centrifuged at 1100 rpm for 5 min at 4 °C, the supernatant aspirated, and the pellet was resuspended in cold Matrigel Growth Factor Reduced Basement Membrane Matrix (Corning). Next, 25 μL of the organoid + Matrigel suspension was added to the center of a pre-warmed 48-well tissue culture plate (Corning) and the plate incubated in a 37 °C, 5% CO2 humidified incubator for 15 min to allow the Matrigel to harden. Then, 300 μL of colon organoid growth media supplemented with 100 U/mL Penicillin, 100 μg/mL Streptomycin, and 10 μM ROCKi was added to each well. The plate was then incubated overnight at 37 °C, 5% CO2 in a humidified incubator. On days 1 and 2 of culture, 300 μL of colon organoid growth media supplemented with 100 U/mL Penicillin, 100 μg/mL Streptomycin, and 10 μM ROCKi was replaced daily. On day 3 of culture, the media was replaced with 300 μL of colon organoid growth media supplemented with only 100 U/mL Penicillin and 100 μg/mL Streptomycin. Colon organoid growth media was then replaced every other day. Organoids were passaged on day 12. Colon organoids were passaged at least twice before use in experiments.
Stimulation of human colonic organoids with microvesicles
Human colon organoids were co-cultured with or without 4.5 × 108 luminal microvesicles from paired uninfected or SIV-infected macaques for 2 days. Microvesicles were diluted in colon organoid growth media. Control wells received colon organoid growth media with equivalent volume of Buffer XE only.
Immunofluorescence staining of human colonic organoids
After two days of stimulation co-culture, media was discarded from the organoid wells. Each well received 1 mL of cold Gentle Cell Dissociation Reagent (StemCell Technologies). The Matrigel dome was broken up by pipetting and the suspension was transferred to a 1.5 mL Eppendorf tube that was pre-coated with fetal bovine serum. Organoids were incubated in Gentle Cell Dissociation Buffer for 30 min at 4 °C with rocking and then were pelleted by gravity. The supernatant was removed and the organoids were fixed in 1 mL of 4% paraformaldehyde (ThermoFisher Scientific) overnight at 4 °C with rocking. The next day, the organoids were pelleted by gravity. The supernatant was aspirated and the organoids were permeabilized in 1 mL of permeabilization buffer (1X PBS + 0.5% Triton X-100 (MilliporeSigma) + 2% BSA) for 30 min at 4 °C with rocking. The organoids were pelleted by gravity. The supernatant was aspirated and the organoids were blocked in 1 mL of blocking buffer (1X PBS + 0.1% Tween 20 (MilliporeSigma) + 2% BSA) for 1 h at room temperature with rocking. The organoids were then pelleted by gravity and the supernatant aspirated. The organoids were stained overnight at 4 °C with rocking in an antibody cocktail containing anti-human EPCAM-Alexa Fluor 488 (Biolegend; Clone: CO17-1A; 1:100), Ki-67-Alexa Fluor 700 (BD Pharmingen; Clone: B56; 1:100), and active caspase-3-Alexa Fluor 647 (BD Pharmingen; Clone: C92-605; 1:100) diluted in blocking buffer. The next day, the organoids were pelleted by gravity, washed once with 1X PBS, and stained with DAPI diluted in 1X PBS (ThermoFisher Scientific; 1 μg/mL) for 10 min at room temperature with rocking. The organoids were then washed twice with 1X PBS, pelleted by gravity and the supernatant was discarded. The organoids were then resuspended in 10 μL of Fluoromount-G (Southern Biotech, USA), and mounted on a microscope slide with a coverslip.
Human colon organoid imaging and quantification
Mosaic images of human colon organoids were acquired using a Leica SP8 confocal microscope (Leica, Germany) equipped with a ×40/1.32NA objective, HyD detectors, and appropriate excitation lasers. Quantification of Ki-67 positive cells and intensity of active caspase-3 was done using Bitplane (Imaris, Switzerland). In brief, nuclei, stained with DAPI, were quantified using the “spots” feature. The subset of nuclei positive for Ki-67-Alexa Fluor 700 was identified by further filtering for Alexa Fluor 700 mean intensity. Active caspase-3-Alexa Fluor 647 was quantified using the “surface” feature and obtaining the intensity sum of all Alexa Fluor 647 signal. This was normalized to the volume of the organoid, which was obtained by creating a surface on the EPCAM-Alexa Fluor 488 channel, which stains all cells in the organoid.
Statistics
Macaque microvesicle concentration and diameter, CD63 and DEFB1 concentrations, mRNA Ct values, L. salivarius generation time, and immunofluorescence staining were all compared by paired, two-way t-test in Prism (GraphPad, v8.3.1). miRNA chosen for follow-up by individual qRT-PCR were compared by paired, one-way t-test in macaques. All mangabey results were compared utilizing unpaired, two-way t-tests. miRNA Array variability was assessed by two-way Wilcoxon signed-rank test and visualized by PCA using the ggbiplot package (v0.55) in RStudio (v1.1.463). From all miRnet identified targets, enriched gene sets were identified by hypergeometric test using the KEGG database option, with inclusion of p value adjustments for multiple-comparisons. The association between DEFB1 and PPARG transcripts in jejunal homogenates was assessed by simple linear regression and Pearson correlation (Prism).
References
Marchetti, G., Tincati, C. & Silvestri, G. Microbial translocation in the pathogenesis of HIV infection and AIDS. Clin. Microbiol. Rev. 26, 2–18 (2013).
Hunt, P. W. HIV and inflammation: mechanisms and consequences. Curr. HIV/AIDS Rep. 9, 139–147 (2012).
Sender, R., Fuchs, S. & Milo, R. Revised estimates for the number of human and bacteria cells in the body. PLoS Biol. 14, e1002533 (2016).
Allaire, J. M. et al. The intestinal epithelium: central coordinator of mucosal immunity. Trends Immunol. 39, 677–696 (2018).
Brenchley, J. M. Mucosal immunity in human and simian immunodeficiency lentivirus infections. Mucosal Immunol. 6, 657–665 (2013).
Estes, J. D. et al. Damaged intestinal epithelial integrity linked to microbial translocation in pathogenic simian immunodeficiency virus infections. PLoS Pathog. 6, e1001052 (2010).
Delgado, M. E., Grabinger, T. & Brunner, T. Cell death at the intestinal epithelial front line. FEBS J. 283, 2701–2719 (2016).
Klatt, N. R. et al. Loss of mucosal CD103+ DCs and IL-17+ and IL-22+ lymphocytes is associated with mucosal damage in SIV infection. Mucosal Immunol. 5, 646–657 (2012).
Mudd, J. C. et al. Hallmarks of primate lentiviral immunodeficiency infection recapitulate loss of innate lymphoid cells. Nat. Commun. 9, 3967 (2018).
Hanash, A. M. et al. Interleukin-22 protects intestinal stem cells from immune-mediated tissue damage and regulates sensitivity to graft versus host disease. Immunity 37, 339–350 (2012).
Klase, Z. et al. Dysbiotic bacteria translocate in progressive SIV infection. Mucosal Immunol. 8, 1009–1020 (2015).
Ericsen, A. J. et al. Microbial translocation and inflammation occur in hyperacute immunodeficiency virus infection and compromise host control of virus replication. PLoS Pathog. 12, e1006048 (2016).
Guillen, Y. et al. Low nadir CD4+ T-cell counts predict gut dysbiosis in HIV-1 infection. Mucosal Immunol. 12, 232–246 (2019).
Vazquez-Castellanos, J. F. et al. Altered metabolism of gut microbiota contributes to chronic immune activation in HIV-infected individuals. Mucosal Immunol. 8, 760–772 (2015).
Samanta, S. et al. Exosomes: new molecular targets of diseases. Acta Pharm. Sin. 39, 501–513 (2018).
Liu, S. et al. The host shapes the gut microbiota via fecal MicroRNA. Cell Host Microbe 19, 32–43 (2016).
Oehmcke, S. et al. A novel role for pro-coagulant microvesicles in the early host defense against streptococcus pyogenes. PLoS Pathog. 9, e1003529 (2013).
Nahui Palomino, R. A. et al. Extracellular vesicles from symbiotic vaginal lactobacilli inhibit HIV-1 infection of human tissues. Nat. Commun. 10, 5656 (2019).
DeMarino, C. et al. Antiretroviral drugs alter the content of extracellular vesicles from HIV-1-infected cells. Sci. Rep. 8, 7653 (2018).
Hong, X., Schouest, B. & Xu, H. Effects of exosome on the activation of CD4+ T cells in rhesus macaques: a potential application for HIV latency reactivation. Sci. Rep. 7, 15611 (2017).
Arakelyan, A., Fitzgerald, W., Zicari, S., Vanpouille, C. & Margolis, L. Extracellular vesicles carry HIV Env and facilitate HIV infection of human lymphoid tissue. Sci. Rep. 7, 1695 (2017).
Chettimada, S. et al. Exosome markers associated with immune activation and oxidative stress in HIV patients on antiretroviral therapy. Sci. Rep. 8, 7227 (2018).
Lai, F. W., Lichty, B. D. & Bowdish, D. M. Microvesicles: ubiquitous contributors to infection and immunity. J. Leukoc. Biol. 97, 237–245 (2015).
Joffe L. S., Nimrichter L., Rodrigues M. L., Del Poeta M. Potential roles of fungal extracellular vesicles during infection. mSphere 1, e00099–16 (2016).
Gaulke, C. A. et al. Intestinal epithelial barrier disruption through altered mucosal microRNA expression in human immunodeficiency virus and simian immunodeficiency virus infections. J. Virol. 88, 6268–6280 (2014).
Kumar, V. et al. miR-130a and miR-212 disrupt the intestinal epithelial barrier through modulation of PPARgamma and occludin expression in chronic simian immunodeficiency virus-infected rhesus macaques. J. Immunol. 200, 2677–2689 (2018).
Fan, Y. et al. miRNet - dissecting miRNA-target interactions and functional associations through network-based visual analysis. Nucleic Acids Res 44, W135–W141 (2016).
George, M. D., Sankaran, S., Reay, E., Gelli, A. C. & Dandekar, S. High-throughput gene expression profiling indicates dysregulation of intestinal cell cycle mediators and growth factors during primary simian immunodeficiency virus infection. Virology 312, 84–94 (2003).
Hu, G. et al. Release of luminal exosomes contributes to TLR4-mediated epithelial antimicrobial defense. PLoS Pathog. 9, e1003261 (2013).
Raschig, J. et al. Ubiquitously expressed Human Beta Defensin 1 (hBD1) forms bacteria-entrapping nets in a redox dependent mode of action. PLoS Pathog. 13, e1006261 (2017).
Wendler, J. et al. Bacterial periplasmic oxidoreductases control the activity of oxidized human antimicrobial beta-Defensin 1. Infect. Immun. 86, e00875–17 (2018).
O’Neil, D. A. et al. Expression and regulation of the human beta-defensins hBD-1 and hBD-2 in intestinal epithelium. J. Immunol. 163, 6718–6724 (1999).
Malik, A. N. & Al-Kafaji, G. Glucose regulation of beta-defensin-1 mRNA in human renal cells. Biochem. Biophys. Res. Commun. 353, 318–323 (2007).
Kelly, C. J. et al. Fundamental role for HIF-1alpha in constitutive expression of human beta defensin-1. Mucosal Immunol. 6, 1110–1118 (2013).
Peyrin-Biroulet, L. et al. Peroxisome proliferator-activated receptor gamma activation is required for maintenance of innate antimicrobial immunity in the colon. Proc. Natl Acad. Sci. USA 107, 8772–8777 (2010).
Brenchley, J. M., Silvestri, G. & Douek, D. C. Nonprogressive and progressive primate immunodeficiency lentivirus infections. Immunity 32, 737–742 (2010).
Shembade, N., Parvatiyar, K., Harhaj, N. S. & Harhaj, E. W. The ubiquitin-editing enzyme A20 requires RNF11 to downregulate NF-kappaB signalling. EMBO J. 28, 513–522 (2009).
Corleis, B. et al. Early type I Interferon response induces upregulation of human beta-defensin 1 during acute HIV-1 infection. PLoS One 12, e0173161 (2017).
Braida, L. et al. A single-nucleotide polymorphism in the human beta-defensin 1 gene is associated with HIV-1 infection in Italian children. AIDS 18, 1598–1600 (2004).
Zapata, W. et al. Increased levels of human beta-defensins mRNA in sexually HIV-1 exposed but uninfected individuals. Curr. HIV Res. 6, 531–538 (2008).
Yasuda, K. et al. Biogeography of the intestinal mucosal and lumenal microbiome in the rhesus macaque. Cell Host Microbe 17, 385–391 (2015).
Nepelska, M. et al. Commensal gut bacteria modulate phosphorylation-dependent PPARgamma transcriptional activity in human intestinal epithelial cells. Sci. Rep. 7, 43199 (2017).
Byndloss, M. X. et al. Microbiota-activated PPAR-gamma signaling inhibits dysbiotic Enterobacteriaceae expansion. Science 357, 570–575 (2017).
Crakes, K. R. et al. PPARalpha-targeted mitochondrial bioenergetics mediate repair of intestinal barriers at the host-microbe intersection during SIV infection. Proc. Natl Acad. Sci. USA 116, 24819–24829 (2019).
Clayton, J. B. et al. Captivity humanizes the primate microbiome. Proc. Natl Acad. Sci. USA. 113, 10376–10381 (2016).
Tipton, L. et al. Fungi stabilize connectivity in the lung and skin microbial ecosystems. Microbiome 6, 12 (2018).
Velazquez, E. M. et al. Endogenous Enterobacteriaceae underlie variation in susceptibility to Salmonella infection. Nat. Microbiol. 4, 1057–1064 (2019).
Taha-Abdelaziz, K. et al. In vitro assessment of immunomodulatory and anti-Campylobacter activities of probiotic lactobacilli. Sci. Rep. 9, 17903 (2019).
O’Shea, E. F. et al. a new type of cationic bacteriocin with unusual immunity. mBio 4, e00498–00413 (2013).
Zhang, L. J. & Gallo, R. L. Antimicrobial peptides. Curr. Biol. 26, R14–R19 (2016).
Kraemer, B. F. et al. Novel anti-bacterial activities of beta-defensin 1 in human platelets: suppression of pathogen growth and signaling of neutrophil extracellular trap formation. PLoS Pathog. 7, e1002355 (2011).
Shimoda, M. & Khokha, R. Metalloproteinases in extracellular vesicles. Biochim. Biophys. Acta Mol. Cell Res. 1864, 1989–2000 (2017).
Kumar, L., Cox, C. R. & Sarkar, S. K. Matrix metalloprotease-1 inhibits and disrupts Enterococcus faecalis biofilms. PLoS One 14, e0210218 (2019).
Teng, Y. et al. Plant-derived exosomal MicroRNAs shape the gut microbiota. Cell Host Microbe 24, 637–652 e638 (2018).
Koyama, M. et al. MHC class II antigen presentation by the intestinal epithelium initiates graft-versus-host disease and is influenced by the microbiota. Immunity 51, 885–898 e887 (2019).
Agrahari, V., Agrahari, V., Burnouf, P. A., Chew, C. H. & Burnouf, T. Extracellular microvesicles as new industrial therapeutic frontiers. Trends Biotechnol. 37, 707–729 (2019).
Del Prete, G. Q. et al. Molecularly tagged simian immunodeficiency virus SIVmac239 synthetic swarm for tracking independent infection events. J. Virol. 88, 8077–8090 (2014).
Ortiz A. M., et al. Antiretroviral therapy administration in healthy rhesus macaques is associated with transient shifts in intestinal bacterial diversity and modest immunological perturbations. J. Virol. 93, e00472–19 (2019).
Crespo, M. et al. Colonic organoids derived from human induced pluripotent stem cells for modeling colorectal cancer and drug testing. Nat. Med. 23, 878–884 (2017).
Acknowledgements
We would like to acknowledge Heather Kendall, JoAnne Swerczek, Richard Herbert, Tracy Meeker, Amanda Cerqueda, Madison Prestipino and all the veterinary staff at the NIH and Yerkes animal centers for their excellent veterinary care. We would like to acknowledge Elise O’Connell (NIH) for valued technical assistance and discussion. Funding for this study was provided in part by the Division of Intramural Research/NIAID (NIH) and by ORIP/OD P51OD011132 (Yerkes). The content of this publication does not necessarily reflect the views or policies of DHHS, nor does the mention of trade names, commercial products, or organizations imply endorsement by the U.S. Government.
Author information
Authors and Affiliations
Contributions
J.K.F., C.A.L., E.P.K., P.J.B., L.P., E.G.G. and A.M.O. performed experiments. J.M.B. and A.M.O. conceived and designed the study and analyzed data. R.M.B. and S.E.B. provided valuable samples. M.S., M.G.S., T.B.N., N.A. and B.L.K. provided valued technical assistance and discussions. All authors contributed to writing the paper and approved the final version of this manuscript to be published and agree to be accountable for all aspects of the work.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Flynn, J.K., Langner, C.A., Karmele, E.P. et al. Luminal microvesicles uniquely influence translocating bacteria after SIV infection. Mucosal Immunol 14, 937–948 (2021). https://doi.org/10.1038/s41385-021-00393-8
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41385-021-00393-8