Abstract
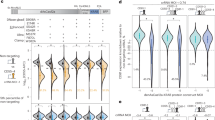
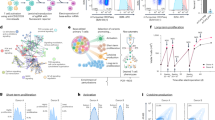
Mutations in U2AF1 are relatively common in myelodysplastic neoplasms (MDS) and are associated with an inferior prognosis, but the molecular mechanisms underlying this are not fully elucidated. Circular RNAs (circRNAs) have been implicated in cancer, but it is unknown how mutations in splicing factors may impact on circRNA biogenesis. Here, we used RNA-sequencing to investigate the effects of U2AF1 mutations on circRNA expression in K562 cells with a doxycycline-inducible U2AF1S34 mutation, in a mouse model with a doxycycline-inducible U2AF1S34 mutation, and in FACS-sorted CD34+ bone marrow cells from MDS patients with either U2AF1S34 or U2AF1Q157 mutations. In all contexts, we found an increase in global circRNA levels in the U2AF1-mutated setting, which was independent of expression changes in the cognate linear host genes. In patients, the U2AF1S34 and U2AF1Q157 mutations were both associated with an overall increased expression of circRNAs. circRNAs generated by a non-Alu-mediated mechanism generally showed the largest increase in expression levels. Several well-described cancer-associated circRNAs, including circZNF609 and circCSNK1G3, were upregulated in MDS patients with U2AF1 mutations compared to U2AF1-wildtype MDS controls. In conclusion, high circRNA expression is observed in association with U2AF1 mutations in three biological systems, presenting an interesting possibility for biomarker and therapeutic investigation.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
Raw RNAseq data from the K562 cell line with doxycycline-induced U2AF1S34F/WT is available at accession number GSE224576 on GEO datasets, whilst RNAseq data from the mouse model are available at accession number GSE89834. The raw RNA sequencing data from patients and healthy controls generated in this study are available under restricted access, since all genomic data are considered sensitive personal data according to Danish Law and the European Union General Data Protection Regulation (GDPR), and thus cannot be shared with third parties without prior approval. To access the RNA data, an application must be sent to Kirsten.Groenbaek@regionh.dk. Applications will be reviewed by the DCCC/PTH board and subsequently by the Danish Data Protection Agency (DDPA). Access can only be granted for research purposes, and only if a data processor or data transfer agreement can be made in accordance with Danish and European law at the given time.
Code availability
The novel code for polypyrimidine tract detection and evaluation has been deposited at https://github.com/omiics/PPT_finder10. All other code utilized previously published methods (see Bioinformatics above, and Supplementary Methods).
Change history
24 March 2023
Figure 1 has been corrected.
References
U2AF1 Mutation - My Cancer Genome. https://www.mycancergenome.org/content/alteration/u2af1-mutation/ (accessed 27 Jan 2021).
Thol F, Kade S, Schlarmann C, Löffeld P, Morgan M, Krauter J, et al. Frequency and prognostic impact of mutations in SRSF2, U2AF1, and ZRSR2 in patients with myelodysplastic syndromes. Blood. 2012;119:3578–84.
Papaemmanuil E, Gerstung M, Malcovati L, Tauro S, Gundem G, Van Loo P, et al. Clinical and biological implications of driver mutations in myelodysplastic syndromes. Blood. 2013;122:3616–27.
Yoshida K, Sanada M, Shiraishi Y, Nowak D, Nagata Y, Yamamoto R, et al. Frequent pathway mutations of splicing machinery in myelodysplasia. Nature. 2011;478:64–69.
Graubert TA, Shen D, Ding L, Okeyo-Owuor T, Lunn CL, Shao J, et al. Recurrent mutations in the U2AF1 splicing factor in myelodysplastic syndromes. Nat Genet. 2012;44:53–57.
Fabre MA, de Almeida JG, Fiorillo E, Mitchell E, Damaskou A, Rak J, et al. The longitudinal dynamics and natural history of clonal haematopoiesis. Nature. 2022;606:335–42.
Li B, Zou D, Yang S, Ouyang G, Mu Q. Prognostic significance of U2AF1 mutations in myelodysplastic syndromes: a meta-analysis. J Int Med Res. 2020;48:30006051989101.
Wang H, Zhang N, Wu X, Zheng X, Ling Y, Gong Y. Prognostic value of U2AF1 mutant in patients with de novo myelodysplastic syndromes: a meta-analysis. Ann Hematol. 2019;98:2629–39.
Ilagan JO, Ramakrishnan A, Hayes B, Murphy ME, Zebari AS, Bradley P, et al. U2AF1 mutations alter splice site recognition in hematological malignancies. Genome Res. 2015;25:14–26.
Przychodzen B, Jerez A, Guinta K, Sekeres MA, Padgett R, Maciejewski JP, et al. Patterns of missplicing due to somatic U2AF1 mutations in myeloid neoplasms. Blood. 2013;122:999–1006.
Okeyo-Owuor T, White BS, Chatrikhi R, Mohan DR, Kim S, Griffith M, et al. U2AF1 mutations alter sequence specificity of pre-mRNA binding and splicing. Leukemia. 2015;29:909–17.
Wadugu BA, Nonavinkere Srivatsan S, Heard A, Alberti MO, Ndonwi M, Liu J, et al. U2af1 is a haplo-essential gene required for hematopoietic cancer cell survival in mice. J Clin Invest. 2021;131:e141401.
Shirai CL, Ley JN, White BS, Kim S, Tibbitts J, Shao J, et al. Mutant U2AF1 Expression Alters Hematopoiesis and Pre-mRNA Splicing In Vivo. Cancer Cell. 2015;27:631–43.
Fei DL, Zhen T, Durham B, Ferrarone J, Zhang T, Garrett L, et al. Impaired hematopoiesis and leukemia development in mice with a conditional knock-in allele of a mutant splicing factor gene U2af1. Proc Natl Acad Sci. 2018;115:E10437–E10446.
Wu S, Romfo CM, Nilsen TW, Green MR. Functional recognition of the 3′ splice site AG by the splicing factor U2AF35. Nature. 1999;402:832–5.
Zamore PD, Patton JG, Green MR. Cloning and domain structure of the mammalian splicing factor U2AF. Nature. 1992;355:609–14.
Voith von Voithenberg L, Sánchez-Rico C, Kang H-S, Madl T, Zanier K, Barth A, et al. Recognition of the 3′ splice site RNA by the U2AF heterodimer involves a dynamic population shift. Proc Natl Acad Sci. 2016;113:E7169–E7175.
Taylor J, Lee SC. Mutations in spliceosome genes and therapeutic opportunities in myeloid malignancies. Genes, Chromosom Cancer. 2019;58:889–902.
Brooks AN, Choi PS, de Waal L, Sharifnia T, Imielinski M, Saksena G, et al. A Pan-Cancer Analysis of Transcriptome Changes Associated with Somatic Mutations in U2AF1 Reveals Commonly Altered Splicing Events. PLoS One. 2014;9:e87361.
Pellagatti A, Armstrong RN, Steeples V, Sharma E, Repapi E, Singh S, et al. Impact of spliceosome mutations on RNA splicing in myelodysplasia: dysregulated genes/pathways and clinical associations. Blood. 2018;132:1225–40.
Li W, Zhong C, Jiao J, Li P, Cui B, Ji C, et al. Characterization of hsa_circ_0004277 as a New Biomarker for Acute Myeloid Leukemia via Circular RNA Profile and Bioinformatics Analysis. Int J Mol Sci. 2017;18:597.
Shang J, Chen W-M, Liu S, Wang Z-H, Wei T-N, Chen Z-Z, et al. CircPAN3 contributes to drug resistance in acute myeloid leukemia through regulation of autophagy. Leuk Res. 2019;85:106198.
Sun Y-M, Wang W-T, Zeng Z-C, Chen T-Q, Han C, Pan Q, et al. circMYBL2, a circRNA from MYBL2, regulates FLT3 translation by recruiting PTBP1 to promote FLT3-ITD AML progression. Blood. 2019;134:1533–46.
Ping L, Jian-Jun C, Chu-Shu L, Guang-Hua L, Ming Z. Silencing of circ_0009910 inhibits acute myeloid leukemia cell growth through increasing miR-20a-5p. Blood Cells, Mol Dis. 2019;75:41–47.
Yi Y, Yi J, Zhu X, Zhang J, Zhou J, Tang X, et al. Circular RNA of vimentin expression as a valuable predictor for acute myeloid leukemia development and prognosis. J Cell Physiol. 2019;234:3711–9.
Papaioannou D, Volinia S, Nicolet D, Świerniak M, Petri A, Mrózek K, et al. Clinical and functional significance of circular RNAs in cytogenetically normal AML. Blood Adv. 2020;4:239–51.
Merkerova MD, Klema J, Kundrat D, Szikszai K, Krejcik Z, Hrustincova A, et al. Noncoding RNAs and Their Response Predictive Value in Azacitidine-treated Patients With Myelodysplastic Syndrome and Acute Myeloid Leukemia With Myelodysplasia-related Changes. Cancer Genomics Proteomics. 2022;19:205–28.
Lux S, Blätte TJ, Cocciardi S, Schwarz K, Döhner H, Döhner K, et al. Deregulated Expression of Circular RNAs in Acute Myeloid Leukemia. Blood. 2018;132:3894–3894.
Wu W, Li S, Zhao G, Li N, Wang X. Identification of circular RNAs as novel biomarkers and potentially functional competing endogenous RNA network for myelodysplastic syndrome patients. Cancer Sci. 2021;112:1888–98.
Nicolet BP, Engels S, Aglialoro F, van den Akker E, von Lindern M, Wolkers MC. Circular RNA expression in human hematopoietic cells is widespread and cell-type specific. Nuc Acids Res. 2018;46:8168–80.
Kristensen LS, Jakobsen T, Hager H, Kjems J. The emerging roles of circRNAs in cancer and oncology. Nat Rev Clin Oncol. 2022;19:188–206.
Ivanov A, Memczak S, Wyler E, Torti F, Porath HT, Orejuela MR, et al. Analysis of Intron Sequences Reveals Hallmarks of Circular RNA Biogenesis in Animals. Cell Rep. 2015;10:170–7.
Zhang X-O, Wang H-B, Zhang Y, Lu X, Chen L-L, Yang L. Complementary Sequence-Mediated Exon Circularization. Cell. 2014;159:134–47.
Conn SJ, Pillman KA, Toubia J, Conn VM, Salmanidis M, Phillips CA, et al. The RNA Binding Protein Quaking Regulates Formation of circRNAs. Cell. 2015;160:1125–34.
Errichelli L, Dini Modigliani S, Laneve P, Colantoni A, Legnini I, Capauto D, et al. FUS affects circular RNA expression in murine embryonic stem cell-derived motor neurons. Nat Commun. 2017;8:14741.
Barrett SP, Wang PL, Salzman J. Circular RNA biogenesis can proceed through an exon-containing lariat precursor. Elife. 2015;4:1–18.
Starke S, Jost I, Rossbach O, Schneider T, Schreiner S, Hung L-H, et al. Exon Circularization Requires Canonical Splice Signals. Cell Rep. 2015;10:103–11.
Liang D, Tatomer DC, Luo Z, Wu H, Yang L, Chen L-L, et al. The Output of Protein-Coding Genes Shifts to Circular RNAs When the Pre-mRNA Processing Machinery Is Limiting. Mol Cell. 2017;68:940–954.e3.
Kelly S, Greenman C, Cook PR, Papantonis A. Exon Skipping Is Correlated with Exon Circularization. J Mol Biol. 2015;427:2414–7.
Smith MA, Choudhary GS, Pellagatti A, Choi K, Bolanos LC, Bhagat TD, et al. U2AF1 mutations induce oncogenic IRAK4 isoforms and activate innate immune pathways in myeloid malignancies. Nat Cell Biol. 2019;21:640–50.
Cheruiyot A, Li S, Nonavinkere Srivatsan S, Ahmed T, Chen Y, Lemacon DS, et al. Nonsense-Mediated RNA Decay Is a Unique Vulnerability of Cancer Cells Harboring SF3B1 or U2AF1 Mutations. Cancer Res. 2021;81:4499–513.
Shirai CL, White BS, Tripathi M, Tapia R, Ley JN, Ndonwi M, et al. Mutant U2AF1-expressing cells are sensitive to pharmacological modulation of the spliceosome. Nat Commun. 2017;8:14060.
Gao Y, Zhang J, Zhao F. Circular RNA identification based on multiple seed matching. Brief Bioinform. 2018;19:803–10.
Memczak S, Jens M, Elefsinioti A, Torti F, Krueger J, Rybak A, et al. Circular RNAs are a large class of animal RNAs with regulatory potency. Nature. 2013;495:333–8.
Stagsted LVW, O’Leary ET, Ebbesen KK, Hansen TB. The RNA-binding protein SFPQ preserves long-intron splicing and regulates circRNA biogenesis in mammals. Elife. 2021;10:1–26.
Stagsted LVW, Nielsen KM, Daugaard I, Hansen TB, Noncoding AUG. circRNAs constitute an abundant and conserved subclass of circles. Life Sci Alliance. 2019;2:e201900398.
Shen S, Park JW, Lu Z, Lin L, Henry MD, Wu YN, et al. rMATS: Robust and flexible detection of differential alternative splicing from replicate RNA-Seq data. Proc Natl Acad Sci. 2014;111:E5593–E5601.
Wang H, Guo Y, Dong Z, Li T, Xie X, Wan D, et al. Differential U2AF1 mutation sites, burden and co-mutation genes can predict prognosis in patients with myelodysplastic syndrome. Sci Rep. 2020;10:18622.
Cao D. Reverse complementary matches simultaneously promote both back-splicing and exon-skipping. BMC Genomics. 2021;22:586.
Koh W, Gonzalez V, Natarajan S, Carter R, Brown PO, Gawad C. Dynamic ASXL1 Exon Skipping and Alternative Circular Splicing in Single Human Cells. PLoS One. 2016;11:e0164085.
Zhang Y, Xue W, Li X, Zhang J, Chen S, Zhang J-L, et al. The Biogenesis of Nascent Circular RNAs. Cell Rep. 2016;15:611–24.
Jeck WR, Sorrentino JA, Wang K, Slevin MK, Burd CE, Liu J, et al. Circular RNAs are abundant, conserved, and associated with ALU repeats. RNA. 2013;19:141–57.
Bachmayr-Heyda A, Reiner AT, Auer K, Sukhbaatar N, Aust S, Bachleitner-Hofmann T, et al. Correlation of circular RNA abundance with proliferation – exemplified with colorectal and ovarian cancer, idiopathic lung fibrosis and normal human tissues. Sci Rep. 2015;5:8057.
Dahl M, Husby S, Eskelund CW, Besenbacher S, Fjelstrup S, Côme C, et al. Expression patterns and prognostic potential of circular RNAs in mantle cell lymphoma: a study of younger patients from the MCL2 and MCL3 clinical trials. Leukemia. 2022;36:177–88.
Moldovan L-I, Hansen TB, Venø MT, Okholm TLH, Andersen TL, Hager H, et al. High-throughput RNA sequencing from paired lesional- and non-lesional skin reveals major alterations in the psoriasis circRNAome. BMC Med Genomics. 2019;12:174.
Zhu Y, Song D, Guo J, Jin J, Tao Y, Zhang Z, et al. U2AF1 mutation promotes tumorigenicity through facilitating autophagy flux mediated by FOXO3a activation in myelodysplastic syndromes. Cell Death Dis. 2021;12:655.
Moldenhauer A, Futschik M, Lu H, Helmig M, Götze P, Bal G, et al. Interleukin 32 promotes hematopoietic progenitor expansion and attenuates bone marrow cytotoxicity. Eur J Immunol. 2011;41:1774–86.
Marcondes AM, Mhyre AJ, Stirewalt DL, Kim S-H, Dinarello CA, Deeg HJ. Dysregulation of IL-32 in myelodysplastic syndrome and chronic myelomonocytic leukemia modulates apoptosis and impairs NK function. Proc Natl Acad Sci. 2008;105:2865–70.
Liu C-X, Li X, Nan F, Jiang S, Gao X, Guo S-K, et al. Structure and Degradation of Circular RNAs Regulate PKR Activation in Innate Immunity. Cell. 2019;177:865-880.
Banerjee S, Gusho E, Gaughan C, Dong B, Gu X, Holvey-Bates E, et al. OAS-RNase L innate immune pathway mediates the cytotoxicity of a DNA-demethylating drug. Proc Natl Acad Sci. 2019;116:5071–6.
Acknowledgements
The study was supported by a grant to LSK from the Lundbeck Foundation (R307-2018-3433), a center grant from the Novo Nordisk Foundation (Novo Nordisk Foundation Center for Stem Cell Biology, DanStem; grant NNF17CC0027852) and the Greater Copenhagen Health Science Partners (Clinical Academic Group in Translational Hematology). The project is part of the Danish Research Center for Precision Medicine in Blood Cancers funded by Danish Cancer Society grant R223-A13071. EW received additional funding from Rigshospitalets Forskningsfond and the University of Copenhagen. M.J. Walter received the Siteman Investment Program (5124) from Washington University, a Developmental Research Program (DRP-1901) of the SPORE in Leukemia (NIH/NCI, P50CA171963), Edward P. Evans Foundation, and Leukemia and Lymphoma Society (7024-21). Bioinformatics were carried out with the assistance of Omiics (Aarhus, Denmark).
Author information
Authors and Affiliations
Contributions
EW, KG, and LSK conceived of the idea for the project. EW carried out laboratory work with patient samples, along with statistical analysis of all data sets and manuscript writing. UA carried out knockdown experiments, performed rMATS analyses, assisted with manuscript writing and editing, and figure production. TBH carried out bioinformatics and contributed to data interpretation. ZG and ROB carried out CRISPR-Cas9 laboratory experiments. MT contributed to data interpretation and analysis strategy, in addition to variant filtering. CC carried out FACS sorting of patient samples. SNS and MJW were involved in data interpretation as well as contributing the K562 cell line data. TA carried out the K562 cell line experiments. JSJ and BCS carried out NGS panel sequencing and variant filtering. CS and KRJ were involved in patient inclusion and characterization. NØ and JK contributed to project development and data interpretation. KG and LSK were involved in project planning, data interpretation and manuscript development. All authors contributed intellectual input and revised and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Wedge, E., Ahmadov, U., Hansen, T.B. et al. Impact of U2AF1 mutations on circular RNA expression in myelodysplastic neoplasms. Leukemia 37, 1113–1125 (2023). https://doi.org/10.1038/s41375-023-01866-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41375-023-01866-4
This article is cited by
-
CircZBTB46 predicts poor prognosis and promotes disease progression of myelodysplastic syndromes and acute myeloid leukemia
Clinical and Experimental Medicine (2023)