Abstract
The detection of mismatched base pairs in DNA plays a crucial role in the diagnosis of genetic-related diseases and conditions, especially for early stage treatment. Among the various biosensors that have been used for DNA detection, EC sensors show great promise because they are capable of precise DNA recognition and efficient signal transduction. Advancements in micro- and nanotechnologies, specifically fabrication techniques and new nanomaterials, have enabled for the development of highly sensitive, highly specific sensors making them attractive for the detection of small sequence variations. Furthermore, the integration of sensors with sample preparation and fluidic processes enables for rapid, multiplexed DNA detection essential for POC clinical diagnostics.
Similar content being viewed by others
Main
The recent discovery and sequencing of the human genome has provided valuable insight into understanding how genetic factors contribute to the development of disease. Specifically, the detection of DNA sequence variations plays an important role in the diagnosis of genetic-related diseases and conditions, especially for early stage treatment and monitoring. Among the different types of diseases caused by DNA alterations, sequence-specific mismatch has the most importance, yet is extremely difficult to detect (1), especially for single-nucleotide polymorphism (SNP). Furthermore, sequence-specific detection has great importance in various medical and scientific applications such as the diagnosis of inherited diseases and the study of pathogen response and bacterial/viral detection.
Because of the complex nature of DNA, the detection of single or small numbers of base mismatches requires high sensitivity and specificity (2–4). Current detection methods rely on sample amplification combined with meticulous experimental stringency control (5). For example, polymerase chain reaction (PCR) requires careful primer design and accurate temperature control to obtain sensitivities in the fM range with single-base mismatch specificity (1,3,4). Although these conventional technologies provide the golden standard for laboratory-based DNA diagnostics, they cannot meet the requirements of POC clinical diagnostics (4).
EC sensors, initially developed to detect biomolecules in a laboratory setting, have recently found extensive applications for on-site biosensing and detection (6,7), especially for medical and clinical diagnostics (8–12). While offering simplicity in operation and sample manipulation, the contemporary EC biosensor also provides highly sensitive and specific measurements for a broad spectrum of biomolecules (13–17). The sample size required for current EC sensors is small, ranging from several microliters to hundreds of nanoliters, which includes the sample pretreatment reagents. Additionally, the detection time is relatively fast, varying from a few minutes to tens of seconds. However, the most important feature of EC sensors is their potential to be easily transformed from a laboratory-based instrument to a commercializable POC device. Because of all these advantages, EC biosensing for DNA diagnostics is becoming a very promising area of research and development.
Recently, micro- and nanotechnologies have shown emerging potential in EC DNA diagnostics. EC sensors offer a perfect interface for incorporating these technologies, which includes a variety of new materials and fabrication processes. Nanomaterials can be used in various aspects of the detection system including capture probes, reporting molecules, electrode fabrication, and electrode coatings (18–25). These materials offer improved biocompatibility, additional binding sites and higher signal intensities (via enhanced electrical properties) compared with traditional materials in EC sensors (17,21,25–28). Nanofabrication allows for miniaturization of the sensor, which improves the sensitivity and reduces the sample and reagent volumes, making the detection process more efficient. Although nanomaterials and nanofabrication are described here as two separated categories, recent trends combine both of these elements in the design of new EC sensors for DNA diagnostics (Fig. 1). With contributions from microfluidics and MEMS technologies, EC sensors can be integrated onto portable platforms incorporating all the necessary preparation and fluidic processes (10,29,30), giving way to commercializable devices for clinical diagnostics (31,32). Ultimately, the end goal of EC sensor development is to construct a total analysis system for rapid DNA biosensing, which incorporates sample pretreatment, sample delivery, and detection.
Principles for EC DNA Sensors
EC detection.
The EC detection of biologic species is based on EC reactions that occur during biorecognition processes (33). These reactions can be exhibited as changes of EC properties (i.e. current/potential, redox kinetics, impedance, etc.) or changes of non-EC properties (i.e. conformation changes, mass transportation, van der Waals interactions, etc.), resulting in fluctuations of an EC signal. Such fluctuations, which usually contribute to high background noise, are not sequence-specific and need to be suppressed during the detection process. The resultant signal readouts can take the form as electrical currents, potentials, or impedances in steady state or changes in these parameters during the recognition process, which correspond to the kinetics of recognition (34). Currently, ex situ EC sensors, in which sample pretreatment and fluidic processing are performed “off-chip,” are most commonly used because they generate a better SNR, resulting from the detection of purified, concentrated biomolecules. However, these sensors have limited applications in POC diagnostics. Therefore, in situ EC sensors, which incorporate all the sample processing steps “on-chip,” are more desirable for clinical application; however, they require higher sensitivity and specificity for non-pretreated samples. Additionally, in situ EC sensors can monitor changes of EC properties, which is more desirable for studying biologic processes during DNA recognition (35).
EC DNA sensors.
A typical EC DNA sensor consists of an electrode, capture probe and reporter probe. A capture probe is an element used to recognize and bind to the target DNA and is usually immobilized onto a solid substrate, such as the electrode surface. However, they can also be immobilized on nanomaterials or other biomolecules. A reporter probe is a molecule that generates the EC signal in response to EC reactions. Both the capture probe and reporter probe are created with high specificity to the target DNA. Additional components, such as electrode coatings and intermediate molecular linkers, are also commonly integrated for improved sensor performance. Common molecules used as probes (capture and reporter) include single-stranded oligonucleotides, aptamers, peptides, and DNA-related proteins (14). In some sensors, the capture and reporter probes are combined together as a single unit for improved integration. Probe, target, and reporter molecules can all be modified or linked with properly integrated nanomaterials, as shown in Figure 2. Because of their high surface-to-volume ratios and biologic compatibilities, nanomaterials not only increase the signal intensity but also help to accumulate/separate specific DNA molecules during EC reactions, which greatly improves the SNR, especially for sequence-specific recognition (28). A wide variety of nanomaterials can be applied, where the most common include metal nanoparticles, cadmium sulfide nanoparticles, CNTs and SiNWs. An extensive and detailed review on the applications of nanomaterials for DNA biosensors can be found in literature (21,28,36).
Common nanomaterials used in EC biosensors for DNA/RNA diagnostics; (A) nanomaterials for electrode coatings, (B) nanomaterials for probe labeling, (C) nanomaterials for target labeling, and (D) nanomaterials for signal reporting. Reprinted from Xu K et al. 2009 Sensors 9:5534–5557. Copyright © 2009 by authors, with permission.
Current EC sensors for DNA diagnostics include two schemes for biologic recognition. The most common scheme, hybridization-based DNA detection, uses nucleotides as the probe and targets elements (Fig. 3A–D) (37). The performance of EC sensors based on this method is highly dependent on the affinity between the probe and target molecules, which can be tuned by the probe design, environmental conditions, and additional amplification processing. This detection system can either use a one probe versus one target scheme, a multiple probe versus single target scheme, or vice versa. For example, EC sensors based on a sandwich detection mode are composed of one capture probe and one reporter probe for each DNA target. Ultimately, no matter what method is used for detection, the output signal is caused by changes in EC properties or EC-related properties.
Schematic representation illustrating the principles for EC DNA sensors. A, Direct oxidation/reduction of nucleotide bases. B, Detection of intercalating complex for single/duplex stands. C, Detection of specific DNA with labeled reporting molecules. D, Detection of specific DNA with integrated capture probe and reporter probe. E, Direct detection after specific DNA enzymatic process. F, Detection of extra labeled reporter after specific DNA enzymatic process.
The second detection scheme, which has emerged in recent years, is enzymatic-based DNA detection (Fig. 3E and F) (16,38,39). In this scheme, DNA-related enzymes are introduced into the biorecognition system and changes in amount of these enzymes correlate to specific biologic process (i.e. deletion/fusion of the target DNA). For example, when an EC sensor experiences a specific process, the enzyme level either increases or decreases, resulting in amplification or reduction of the signal. The enzymatic process is highly specific to a DNA sequence, which makes it ideal for DNA mismatch detection.
Advantages of EC sensors for DNA diagnostics.
EC sensors offer several advantages over other detection methods, making them attractive for DNA biosensing. In addition to being highly specific and sensitive, EC sensors are extremely efficient, in terms of fast detection times and low power consumption. EC DNA sensors largely rely on nucleotide hybridization during the detection process, which involves specific electrostatic charge distributions and strong hydrogen bonding. Because the backbone of a nucleotide is composed of phosphoric acids and base units, the entire molecule is heavily charged with a negative potential. Therefore, hybridization between nucleotides needs to overcome the strong repulsion force between each other. In traditional DNA detection methods, temperature and chemicals are used to reduce the repulsion of these molecules; however, both of these modulations are not very effective. Based on the thermodynamics of DNA hybridization/denaturation, the Gibbs free energy for such processes is in the range of 1 to 10 kcal/mol. Therefore, very high temperatures or ion buffer concentrations are required to overcome this energy barrier and such conditions are likely to interfere with the bio-system. In contrast, EC sensors are capable of producing strong electrical fields, where only several hundred millivolts of potential can overcome the reaction barrier (10,40). Similar to the melting temperature (Tm) for traditional temperature control, the melting potential (Vm), which denotes the voltage at which 50% hybridization/denaturation occurs, is an important parameter useful for characterizing different DNA strands and provides an additional tool for controlling the specificity (41).
A second advantage of EC sensors is their simplicity in manipulating molecules within the sample fluid, which is performed through electrical fields generated by the electrode. Thorough mixing and precise manipulation of molecules are crucial for achieving high hybridization/denaturation efficiency. Traditional temperature or chemical-based control schemes require additional mixing and separation procedures, which greatly hinders advancement toward a POC DNA diagnostic platform. Specifically, the speed of these procedures is limited by chemical reaction times and heat/mass transfer processes within the solution. In contrast, EC sensors can generate high electrical fields within a very short time (34). By using this scheme, molecules near the electrodes can be manipulated by applying different electrical profiles. Additionally, thin dielectric double layers generated in high intensity electric fields can be used to enhance mixing and sample manipulation. Such accurate control circumvents the need for additional components and greatly simplifies the detection process, making in situ DNA detection possible.
A third advantage of EC sensors is their ability to achieve precise DNA recognition due to localization effects. Traditional detection methods, based on temperature or chemical control schemes, lack precision because their effects are dispersed within the entire sample solution rather than localized near the DNA molecules. In addition, the strong hydrogen bonds within DNA makes precise control over hybridization/denaturation quite difficult. In contrast, EC sensors can generate well-defined, localized electrical fields within the electrode domains where DNA recognition occurs. Additionally, nano-sized electrodes can produce electric fields that are concentrated within a small region surrounding the electrode, which allows for even greater precision and localization. Nanoelectrodes also require much smaller electrical potentials and reduces the overall power consumption of the sensor.
Because of the advantages listed above, there are several key features that EC sensors offer for DNA diagnostics:
-
1
Detection speed: the detection of DNA involves several processes including DNA recognition, sample manipulation, and signal readout. By using EC sensors, DNA recognition can occur within minutes and in some cases, several seconds. The entire detection process, including sample preparation, sample delivery, and signal readout, can be completed in several minutes. Such rapid detection is ideal for POC clinical diagnostics.
-
2
Sensitivity: with appropriate electrical field profiles, EC DNA sensors can achieve sensitivities down to several fM and in some cases, aM for short oligonucleotides. Such sensitivities can be achieved without sample amplification (i.e. PCR), which is a major advantage for POC systems. Additionally, the total amount of sample required for detection is on the orders of several microliters.
-
3
Specificity: the stringency of EC-controlled DNA recognition enables for single-base mismatch specificity, even in clinical samples.
-
4
Convenience: with contributions from MEMS and nanotechnology, EC sensors can be integrated onto portable platforms, enabling for POC monitoring and on-site biosensing.
-
5
Multiplexing: nowadays, a single biomarker is not sufficient for high specificity detection in clinical samples. By combining several biomarkers, the detection accuracy can be greatly improved, making multiplex detection for DNA diagnostics very important. Because the electrical fields generated by electrodes are highly localized, they can be used for simultaneous detection within a miniaturized platform, where the detection condition for each DNA target can be optimized separately (32,42).
EC DNA Sensor Types
Traditional EC biosensors are based on detecting changes of EC properties, such as current/potential, redox kinetics, or electrical impedance (34). However, because most biorecognition is influenced by non-EC properties (mass transportation or conformational changes) in addition to EC reactions, EC sensors can also be based on controlling the biorecognition process. From these two aspects, we will distinguish EC DNA sensors in two categories: EC biosensors as transducers and EC biosensors as controllers. In the following section, each type of sensor will be discussed in further detail.
EC sensor as a transducer.
Most EC biosensors directly detect changes in EC signals, in which case, the sensor acts as a transducer. We separate EC sensors into two categories: labeled detection and label-free detection. Usually EC sensors use labeled reporters, which can result in high levels of background noise because of mass transportation processes or even conformational changes. Therefore, prelabeled reporters can help to increase the SNR. The second category, label-free detection, usually refers to detection without any sort of labeling. This scheme also includes detection which incorporates a labeled reporter in conjunction with nonlabeled targets because the final detection process is label-free.
Labeled detection.
Labeling molecules are electrochemically active in that they exhibit specific EC properties, which correlate to the status of DNA targets on the electrodes. Usually, the amount of the labeling molecules indicates the amount of DNA targets in the sample either in a direct detecting mode or a competitive detecting mode (Fig. 3C and F). Nanomaterials, because of their high surface-to-volume ratios, provide more binding sites for nucleic acids. Additionally, applying nanomaterials to electrode coatings and/or labeling tags greatly improves the signal intensity. Metal and magnetic nanoparticles are commonly used because they can be easily accumulated onto the sensor surface via electrical or magnetic fields. By incorporating Au nanoparticles to DNA probes, Ozsoz et al. (24) detected Leiden mutations with a sensitivity down to 0.78 fmol. Castaneda et al. (25) successfully detected the single-base mismatched BRCA1 breast cancer gene and a cystic fibrosis-related gene with a combination of sandwich reactions of Au nanoparticles and magnetic beads. Using magnetic nanoparticles as labels also improves the separation efficiency, where specific DNA/RNA targets can be accumulated while removing nonspecific molecules (25,43,44).
Similar to a direct detection mode, a competitive detection mode measures changes in the output signal, which indicates increased DNA target levels. Liao and Ho (23) recently detected enterohemorrhagic Escherichia coli O157, a verocytotoxin (VT1/2)-producing pathogen, using a competitive EC sensor. The electrodes were modified with Au nanoparticles and a self-assembled monolayer (SAM) of thiol-capped single-stranded DNA (capture probe) for the detection of the rfbE gene, which is specific to E. coli O157. This assay is based on a competition between the target gene and reporter DNA-tagged liposomes. The sensitivity of detection for the rfbE gene was 0.75 aM.
In addition to detection based on changes in the amount of DNA targets, the detection process can be based on conformation changes of the labeling molecules (Fig. 3D). During DNA recognition, conformational change of the DNA molecule alters the distance between the labeling molecules and the electrode, which affects the EC signal. Fan et al. (45) first reported an ultrasensitive EC sensor based on DNA folding during hybridization. The DNA probe is designed as a hairpin structure with a ferrocene-tag at one end, which remains in a closed configuration before hybridization. After hybridizing with a complementary strand, the hairpin probe opens up, enlarging the distance between the ferrocene and the electrode, which generates a significant EC current. The sensitivity of this sensor is approximately 10 pM, even in bodily fluids (45,46). By combining this folding process with an additional amplification process, Wei et al. (10,11,20) detected an oral cancer mRNA gene in whole saliva. In their approach, DNA dendrimers were applied to the surface probe for improved biocompatibility and sensitivities as low as 3.9 fM of mRNA could be obtained.
Label-free detection.
Label-based EC sensors require a prelabeling process, which greatly limits the variance of available reporters and the possibility for in situ detection. Recently, label-free EC sensors using nanomaterials have attracted much attention (47,48). Although eliminating the labeling process does not significantly improve the detection efficiency, it provides the only means to realize universal in situ and in vivo DNA detection. This approach eliminates the need for additional sample separation processes or reporting molecules and allows for DNA levels to be monitored real-time within biologic systems. Label-free detection can be performed without any sort of labeling (9,13,49) or can incorporate a labeled reporter in conjunction with nonlabeled targets (10,50).
One of the simplest ways in which label-free DNA sensors operate is in the direct detection of EC reactions of nucleotide bases (Fig. 3A and E) or intercalating molecules (Fig. 3B). The bases in DNA have specific EC redox properties, which can be used to indicate their quantity on an electrode. However, the redox of DNA bases can be affected by other factors including the environment, hybridization events, and buffer solutions, which results in higher noise. Single/duplex-specific intercalating complexes are another type of label-free EC sensor. These intercalating molecules can either bind to the groove of a DNA duplex or insert into the planes between stacking base pairs. After the dissociation of the duplex, these molecules are released and cause a change in the EC properties. Commonly used DNA-intercalating metallic complexes include Co(2, 2′-bipyridy1)33+, Co(1,10-phenanthroline)33+, methylene blue, daunomycin, and aromatic amines. These compounds have been used for several clinical-related diagnostic applications, such as the detection of cystic fibrosis (51), pathogenic E. coli bacteria (52), Mycobacterium tuberculosis (52), hepatitis B virus (53), the human immunodeficiency virus (HIV) (52), and familial dysbetalipoproteinemia (54). An extensive review of this topic can be found in literature (55).
Another type of indirect, label-free EC sensor is based on a sandwich detection mode. This sensor does not require labeling of the target biomolecules; rather, only the reporting molecules are labeled (56–58). The biorecognition process is performed by monitoring changes in the amount or EC properties of the reporter molecules.
Traditional EC biosensors are based on the measurement of electrical current and potential (amperometry, voltammetry, and coulometry); however, dielectric properties can also be used for impedance-based detection (59–62). Although labeling of target molecules is not mandatory for impedance-based detection, in some cases, it can enhance the signal. Dielectric properties are highly sensitive to the conditions at the electrode interface and exhibit very unique features under certain electrical parameters such as current, potential, frequency, magnetic, and optical fields. Because EC impedance detection is a label-free method and capable of high sensitivity, it is very desirable for POC DNA sensors (49). Currently, impedance-based EC sensors can achieve sensitivities in the nM to pM range. However, obtaining such high sensitivities is still a challenging feat for impedance detection. Fundamentals of EIS and impedance-based detection can be found in literature (34,49).
Nanomaterials are important elements in impedance-based DNA sensors, especially those that poses semiconductor properties (38,59,63–68). Such materials enhance the sensor performance through increasing the electrode surface area, electrical conductivity and connectivity, chemical accessibility and electrocatalysis. The most widely used nanomaterials in impedance sensors are Au nanoparticles and CNTs, which amplify the impedance signals by forming nanoparticle-biomolecule conjugates in a solution phase. Feng et al. (64) reported an EIS sensor for sequence-specific DNA detection of the phosphinothricin acetyltransferase (PAT) gene. The dynamic detection range was from 1 pM to 1 μM with a detection limit of 0.3 pM.
Polymers, including chitosan and dendrimer, are another category of emerging material for impedance-based sensors (69–71). By combining CNT-Au composite nanoparticles with a polyaniline nanofiber (PANnano)-carbon paste electrode (CPE), Zhou et al. (59) detected the PAT gene using a label-free EIS detection scheme. The dynamic range of this sensor ranged from 1.0 × 10−12 mol/L to 1.0 × 10−6 mol/L with a detection limit of 5.6 × 10−13 mol/L. Liao and Cui (61) reported a reagentless impedance biosensor for the detection of neuro-inflammatory cytokine PDGF with a limit of detection around 40 nM. Tiwari and Gong (67) developed an EC biosensor for detecting the breast cancer susceptible gene BRCA1 based on a chitosan-co-polyaniline (CHIT-co-PANI) coating on top of indium–tin-oxide. This sensor had a sensitivity of 2.1 μA/fM with an impressive response time of 16 s.
EC sensor as a controller.
Traditionally, EC sensors do not interact with the biologic system during the detection process and are known as “passive mode” sensors. However, as discussed above, the detection process can be affected by the EC profile, taking into account factors such as electrolysis, electrode potential, and current. This type of sensor is known as an “active mode” sensor and acts as a controller as well as a transducer. In this scheme, the output signal is not restricted to an EC signal but can include other types of signals (i.e. optical, thermometric, piezoelectric, magnetic, micromechanical, and mass change), which greatly expands the application of EC sensors (72,73). Both DNA probe-target and DNA electrode interactions can be controlled by EC fields (Fig. 4). Additionally, the interactions between the electrode and DNA recognition elements can occur before, during, or after the recognition process.
Schematic illustration depicting the various electrical field effects during DNA recognition, including (A) orientation changes, (B) conformational changes, (C) separation from interferents, (D) accumulation to local domain, (E) hybridization with complementary sequence, and (F) denaturation of nonspecific sequence.
Before recognition.
Applications of EC fields before DNA recognition include sample accumulation (Fig. 4C) (74), sample separation (Fig. 4D) (75–78), and controlling probe surface densities (79). AC electrical fields can generate dielectrophoretic (DEP) forces for manipulating molecules within liquids, which can be applied to EC biosensing systems (80). Wang et al. (81) has reported an electrical focusing system for the detection of DNA/RNA at the single-molecule level by using specially-designed three-dimensional (3D) electrodes within a microfluidic reactor.
Understanding the conformation of DNA is crucial for detection in clinical samples because most model systems for EC detection use short oligonucleotides. In its natural state, DNA is in a coiled or quasi-hybridized state, taking on a complex, 3D structure. For improved hybridization efficiency, the complimentary inner/inter-structure is normally removed, so that the molecule is in a stretched conformation. In traditional DNA detection methods, high temperatures and detergent are applied to achieve this goal. By using EC sensors, this process can be accomplished by applying an electric potential, which is faster and more efficient. Within minutes, coiled DNA can be stretched out and ready for subsequent hybridization. For EC sensors with surface immobilized DNA probes, the prerecognition electrical field also arranges the probe in a more uniform angle to the surface (Fig. 4A and B) (82).
During recognition.
There are numerous examples of EC field-assisted hybridization in literature (10,11,20,40,41,83,84). As previously discussed, a positive potential improves the hybridization efficiency in a short time and with a localized pattern (Fig. 4E) (40). In addition, the distance between the reporter and the electrode surface can also be controlled by the EC field. Based on this property, the surface of a metallic electrode can act as a quencher for the output signal. For example, in a sandwich configuration with a hairpin probe, the reporter near the electrode cannot bind with the amplifier and prevents the generation of an output signal. Only when the reporter is far from the surface it can bind with the amplifier and generate output signals (11). Analogously, optical EC sensors are based on the quenching of fluorescent signals from optical labels by the surfaces of metallic electrodes. By adjusting the EC field, the optical signal can be easily modulated. In addition, the specificity of EC DNA sensors can be guaranteed by applying the exact electrical profile corresponding to Vm of specific DNA (i.e. only probes and targets with the same Vm can hybridize) (41,84). Based on this concept, mismatched sequences, which have lower Vm, are differentiated from the “normal” matched sequences.
Furthermore, multiplexed detection is advantageous for the analysis of DNA and DNA-containing biomolecules (10). In this scheme, an electrode array is used where each electrode contains a specific DNA probe/target that has a different Vm, which requires a different electrical profile. This method is more precise than a temperature-based control, where the heating domain is widespread, making clearly defined boundaries difficult to achieve.
After recognition.
EC fields applied after DNA recognition can transport nonspecific species away from the electrode, resulting in higher SNRs. When a positive potential is applied to accumulate DNA near the electrode, it also collects nonspecific molecules, which increases the background noise. By applying a specific negative EC field after DNA recognition, nonspecific adsorbed molecules can be removed from the electrode surface while allowing the target molecules to remain, thereby increasing the SNR (Fig. 4F).
Controlling the EC field during DNA recognition can be performed throughout the entire detection process and is not limited to certain stages or steps. The combinational effects of manipulating molecules and assisting DNA hybridization from EC fields results in fast and effective sensing. For example, a properly designed EC profile with turn-overs between positive and negative potentials enables for thorough mixing (11,20). The positive potential push negatively charged DNA toward the electrode whereas the negative potential cause DNA to repel. Thorough mixing before and during detection is important because it improves the reaction possibility between the probes and targets. For samples containing biomolecules other than DNA (i.e. peptides, proteins, etc.), the mixing process is mainly driven by ions inside the solution. Ultimately, appropriate EC field profiles allow for the entire recognition process to be completed within seconds with high SNRs (10,20).
Sensor Fabrication
Recent advancements in micro- and nanofabrication technologies have allowed for the development of EC DNA sensors which can precisely detect, convert, and amplify signals using various electrode configurations and nanomaterials. Adapted from the manufacturing of integrated circuits (IC) and semiconductors, surface micromachining and nanofabrication offer many advantages for DNA sensors including device miniaturization, high precision, and batch-fabrication capabilities. Additionally, microfluidics and MEMS technologies enable for sensors to be integrated into total analytical systems, allowing for rapid multiplexed detection on a portable platform for the eventual realization of POC clinical diagnostics. The following section will present an overview on technologies used in the fabrication of EC sensors, mainly for DNA biosensing applications. In-depth reviews on micro/nanofabrication technologies for EC sensors can be found in literature (85,86).
MEMS fabrication and micromachining.
The fabrication of EC sensors is largely influenced by semiconductor and MEMS manufacturing due to their abilities to create high quality, high precision structures and devices. These techniques heavily rely on surface micromachining, a top-down fabrication scheme, in which a bulk material is shaped through subsequent patterning, cutting, and etching using externally controlled machines and processes. Sensors constructed using this approach are traditionally fabricated on rigid substrates where silicon, silicon-on-insulator (SOI), and glass are the most common materials due to their compatibility with existing micromachining and nanofabrication processes. Recently, microfabricated devices are widely adapting plastics as the substrate material because they are more cost effective and suitable for batch-fabrication (87). However, plastics are not compatible with many MEMS and nanofabrication processes and therefore cannot achieve very high precision or nanoscale resolution.
Photolithography.
Photolithography is one of the most fundamental processes in semiconductor and MEMS manufacturing, combining high precision patterning with batch-fabrication capabilities. Ultraviolet (UV) light passes through a mask onto a substrate, which is coated with a UV-sensitive photoresist. The light exposes the photoresist, which transfers the pattern from the mask to the substrate. The photoresist is developed in a chemical etchant, resulting in a polymer structure. This structure is commonly used as a mask for subsequent processing steps, such as metal deposition or etching, or can be used as a mold for fabricating channels, wells, and spacers for the entire sensor. Conventional MEMS-based photolithography is capable of producing features down to several microns, which is primarily dictated by the resolution of the mask and the wavelength of the UV light.
Alternative lithography.
Smaller feature can be patterned through alternative lithography techniques, enabling for the fabrication of nanoelectrodes. By miniaturizing electrodes to a similar size scale as the molecules of interest (i.e. DNA, RNA, proteins), higher sensitivity can be achieved compared with macro-sized electrodes. Nanoelectrodes provide smaller effective surface areas for concentrating probe and target molecules, which aids in reducing the background noise. Similar to photolithography, deep UV lithography and x-ray lithography use UV lasers and x-rays, respectively, to expose the resist, which are capable of nanometer resolution. Chua et al. (47) fabricated SiNW arrays by first patterning the electrode fingers using x-ray lithography and performing subsequent thermal oxidation and wet etching to further define their profile (Fig. 5B). EBL is a mask-less approach for patterning resist, which is capable of producing feature sizes down to tens of nanometers. Rather than using light, a beam of electrons scans across the resist, exposing those regions. EBL is commonly used to pattern SiNWs electrodes (88–90), which is usually followed by thermal oxidation and wet etching. SiNW electrodes fabricated on SOI substrates enable for simplified fabrication and improved integration with semiconductor-based signal processing and communication circuits. In an alternative sensor configuration, Lee et al. (91) fabricated oriented nanowell (ONW) arrays within an Au electrode, which was designed so that each well could only accommodate for one or a few biomolecules (Fig. 5C). The nanowells were precisely patterned using EBL whereas the remainder of the electrode surface was passivated with a layer of resist, preventing nonspecific binding and enhancing the signal sensitivity. Analogous to EBL, ion-beam lithography uses a focused beam of ions to expose the resist. Such maskless approaches are relatively slow and require additional photolithographic processing for patterning larger sensor elements, such as the contact pads and electrical leads.
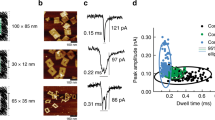
Images of nanomaterials and nanoelements used for EC sensors. A, Transmission electron microscopy (TEM) image of a CdTe-Au multi-segment nanowire. Reprinted from Wang and Ozkan, Nano Lett, 2008;8:398–404 Copyright © 2009 American Chemical Society, with permission. B, Scanning electron microscopy (SEM) image of patterned SiNWs, which are individually addressable by oxidepassivated metal contact lines. Reprinted from Chua et al., Anal Chem, 2009;81:6266–6271 Copyright © 2009 American Chemical Society, with permission. C, Atomic force microscopy (AFM) image of probe ssDNA immobilized inside an ONW array. Reprinted from Lee et al., Appl Phys Lett, 2006;89:113901 Copyright © 2006 American Institute of Physics, with permission. D and E, SEM images of multi-walled CNT (MWCNT) arrays patterned using UV lithography and e-beam lithography, respectively. Reprinted from Li et al., Nano Lett, 2003;3:597–602 Copyright © 2003 American Chemical Society, with permission. F, SEM image of a CNT-poly-l-lysine film on top of a CPE. Reproduced from Jiang et al., Electrochim Acta, 2008;53:2917–2924 Copyright © 2007 Elsevier Ltd., with permission.
Metal deposition.
Thin-film electrodes can be fabricated by depositing metals onto a patterned resist mask through evaporation or sputtering. Metal on top of the mask is removed by selectively etching the underlying resist whereas metal that is deposited directly onto the substrate remains. Most noble metals can be patterned using this approach, making it useful for fabricating a variety of electrodes. Au is commonly used for fabricating the sensing electrode because of its high electrical conductivity, which enables for enhanced sensitivity. Additionally, Au is extremely biocompatible and its surfaces can be easily modified using SAMs, allowing for the direct immobilization of thiolated-probes (92). Electrodes that are fabricated using this approach can have thicknesses ranging from tens of nanometers to several microns, which can be precisely controlled through the deposition process. Electrodeposition, an EC process widely used for industrial metal plating, is an alternative approach more commonly used for fabricating thick-film electrodes. An electrical current is applied to an electrolyte bath containing the substrate and source material. Cations from the source material are reduced and deposited onto a conductive seed layer, which is patterned using lithography to define the shape of the electrode. Alternatively, metals can be deposited on top of screen-printed carbon electrodes (23,93) or silver wool substrates (94). A wide variety of metals can be deposited using electrodeposition (Au, Ag, Ni, Ti, Pt, etc.) while having the capability of producing a wide range of electrode thickness.
Nanofabrication.
Electrodes fabricated using lithographic techniques are usually restricted to flat, two-dimensional (2D) structures. Free-standing electrodes offer larger working surface areas compared with planar electrodes, which enhances the diffusion of redox species/target molecules to the surface, enabling for more pronounced hybridization signals. 3D nanoelectrodes can be constructed using bottom-up approaches, in which nano-sized components and molecules are self-guided and assembled to form the final structure. This approach can generate electrodes with highly-ordered, defect-free atomic structures, enhancing their electrical properties for higher sensitivity measurements. Free-standing nanowires can be fabricated by “nanocasting,” where materials are electroplated or deposited within a mold containing nanopores. Selective etching is performed to remove the outer mold, thereby exposing the enclosed nanowires. Floating multi-segment nanowires consisting of CdTe-Au-CdTe segments were fabricated by sequential electrodeposition of metals within an alumina oxide template (Fig. 5A) (95). Alternatively, free-standing gold nanowires were fabricated through electroless deposition within a polycarbonate membrane followed by controlled plasma etching, allowing for the wires to remain securely embedded in the membrane (96). Nanocasting can also be used to fabricate nonmetallic nanotubes; Chang et al. (97) fabricated polyaniline (PANI) electrodes through polymerization within a thin nanoporous aluminum film. The alumina film was initially sputtered on top of a graphite electrode and subsequently etched in H2S04, resulting in vertically oriented PANI nanotubes arrays.
Recently, CNTs have shown great potential as an electrode material for EC DNA detection due to its superior mechanical and electrical properties (98,99). Specifically, CNTs demonstrate rapid electron transport, amplifying the detection signal and making them effective transducers. Additionally, the well-defined chemistry of CNTs allows for precise immobilization of probe molecules by adsorption or chemical grafting. CNTs can be grown using several processes, including arc discharge, laser ablation, and chemical vapor deposition (CVD). Of these, plasma-enhanced CVD is the most common method for fabricating CNT electrodes due to its ability to precisely pattern nanotubes with specific orientations (Fig. 5D and E). The growth of CNTs is initiated by metal nanostructure catalysts (Ni, Co, or Fe), which are typically patterned via lithography. This technique commonly results in a bundled, forest-like CNT configuration (Fig. 5D) (100,101), which increases the effective surface area; however, lacks the spatial resolution for single molecule detection (102). Improved sensitivity can be achieved by embedding CNT arrays within a SiO2 matrix, which enhances mechanical stability and electrical isolation of the electrodes (66,103,104).
Printing.
Screen printing is a thick-film patterning/deposition technique capable of large-scale sensor production. Screen printed electrodes (SPEs) have been used for various EC DNA sensors due to their straightforward fabrication, high uniformity, and material versatility (105,106). A paste or ink is spread over an emulsion or steel screen, containing the electrode pattern. The paste is usually a mixture of an organic binder, a solvent and the electrode material, which commonly are metallic (Au, Pt, Ag), ceramic (Al2O3, ZrO2), or carbon nanoparticles. The pattern is transferred onto the substrate by forcing the paste through the screen's openings. The paste is then set to dry, removing the solvents, followed by firing to burn off the organic binder. Alternatively, inks can be deposited using a printer for enhanced automation and precision (96,107,108). EC sensors commonly use CPEs composed of carbon nanomaterials (i.e. graphite, carbon fibers, CNTs), which does not require high-temperature processing, allowing them to be fabricated on plastic substrates. Screen printing can produce electrodes with thicknesses of several millimeters with minimum features of ∼100 μm, without the use of expensive equipment or a clean room facility.
Surface chemical property modification.
Recent advancements in surface modification technologies have led to significant improvements in sensor performance. EC sensors commonly use multilayer electrodes, which consist of thin layers of polymers, nanoparticles, or nanoparticle-polymer composites stacked on top of the electrode. Such films can enhance the sensitivity and specificity of the sensor by acting as a 3D matrix for entrapping nucleotide probes and reducing the interference from nonspecific molecules, which can contribute to background noise. Additionally, conductive coatings can enhance the signal of redox species and minimize the loss of signal from the electrode to the electrical circuitry. Because of their high electrical conductivity and enlarged surface area, carbon and metallic nanoparticles are commonly used for particle-based coatings (59,64,109–111). In this approach, a solution containing nanoparticles is dispensed on the electrode surface and allowed to dry. As the solvent evaporates, the nanoparticles form SAMs, which are held together through intermolecular interactions (i.e. Van der Waals, electrostatic, etc.).
Alternatively, nanoparticles can be incorporated within polymer films, which allows for the particles to remain securely embedded within rigid matrix for enhanced robustness. Nanoparticles are commonly integrated into the polymer solution before polymerization, which can be initialized by heat, light, plasma, or electrical current. A CNT-poly-l-lysine film was fabricated by depositing a layer of CNTs on top of a CPE followed by application of poly-l-lysine solution and subsequent electro-polymerizing (Fig. 5F) (112). Additionally, conductive polymers [i.e. polypyrrol, polypyrrol propylic acid (PPA), PAN] can also be used as electrode coatings for signal enhancement (60,64,67,72,113). By applying pulsed electric fields, polymers can be locally electro-polymerized directly on the electrode surface, without the need for additional patterning procedures. Nonconductive polymer films, such as polyamidoamine dendrimers, can be deposited through various chemical bonding schemes (69,71).
Prospects
The future direction of EC DNA sensors is focused on the development of POC systems, which seek to integrate sample handling, fluidic processing, and detection on a portable platform. Although EC biosensors have been widely developed for laboratory-based detection within the past several years, there are very few successful POC devices for clinical diagnostics that are currently commercialized (i.e. glucometers). Clinical applications for EC DNA sensors are still far from reality due to several important issues. Although much work has been done to improve the performance of EC DNA sensors, the sensitivity/specificity is still a key issue. Specifically, the detection of clinical samples requires high sensitivity/specificity as well as high repeatability/reliability, which is still an unresolved problem. To address these issues, new nanomaterials with effective and stable performance are required along with higher stringency control during manufacturing. Additionally, the accuracy for clinical detection can be enhanced though bio-statistic support based on multiple DNA biomarkers. To improve the application of EC sensors for real clinical tests, a simple detection process is desired, which incorporates automatic sample processing or in situ detection. This can be achieved through using micro/nanotechnologies, which offers new materials and sensor fabrication processes. Furthermore, the safety of nanomaterials is becoming a significant issue, especially as applications for these materials become more widespread.
Sensitivity and specificity.
Current EC DNA sensors require labeling to achieve high sensitivity and specificity. Although label-free technology provides convenience for low cost and in situ detection, it suffers from high noise and false positives, which pose a serious problem for achieving high specificity. In most cases, signal amplification is necessary and in particular, specific-signal amplification is required for achieving a good SNR (10). Alternatively, new nanomaterials with DNA-specific binding properties can enhance signal amplification and improve overall sensor performance (114–116).
Repeatability and reliability.
The ability to produce repeatable and highly reliable measurements is one of the most important challenges facing EC DNA sensors, especially for clinical diagnostics and commercial usage. Clinical samples are prone to a high degree of variability, which results from physiologic and lifestyle differences between patients. Additionally, slight variations in the actual sensor (i.e. electrode geometry, uniformity of coatings and probes, etc.) can lead to inconsistencies in measurements, particularly for the detection of small sequence variations. In addition to improving the detection sensitivity and specificity through the use of nanomaterials and nanoelectrodes, optimized fabrication processes and higher stringency control during manufacturing can enhance the overall repeatability and reliability of the sensor.
Biostatistical support.
The performance of EC DNA sensors is largely dependent on the accuracy of the targeting biomarker(s). However, DNA biomarkers for clinical diagnostics still lack the accuracy needed for highly specific detection. Because of the complexity and nonlinearity of the human body, sequence mutations for single DNA/RNA do not necessarily correspond to one specific disease. Recently, a panel of multiple biomarkers resulted in improved accuracy for clinical diagnostics (10,117,118). Therefore, multiplexed detection will be very dominant in the future development of EC nanosensors for sequence-specific detection.
Sample pretreatment.
Clinical samples are complex mixtures, which contain a multitude of components and biologic species. Even in vitro detection of body fluids (i.e. blood, urine, and saliva) presents great challenges for simple detection systems. Usually, several pretreatment processes are required before detection, such as separation, purification, accumulation, and amplification. Application of nanomaterials can greatly simplify and improve the efficiency of such pretreatment processes. Additionally, MEMS and nanofabrication technologies enable for the construction of portable, automated devices with batch-fabrication capabilities. Current research is focused on total system integration where several promising devices have already been demonstrated (10,12,119,120).
Toxicity of nanomaterials.
Although EC DNA sensor performance is greatly enhanced by nanomaterials, their cross-linked structures and associated organic reagents can present serious toxicity problems in biomedical systems (121,122). Therefore, new nanomaterials with low toxicity are in high demand. Recently, new biocompatible nanomaterials have been developed (123–125), including ones which contain biologic backbones, which have attracted extensive attention (20,126,127).
CONCLUSIONS
EC sensors show great potential for DNA biosensing, offering high sensitivity and specificity essential for single-base mismatch detection. Advancements in micro/nanotechnologies, specifically fabrication techniques and new nanomaterials, are largely responsible for improvements in EC sensors. In particular, the detection sensitivity is enhanced through highly-specific molecular recognition (by appropriately-designed targets and probes), improved EC signal generation, transduction and amplification, and enhanced electrical conductivity for minimized background noise. Additionally, EC sensors are extremely efficient, in terms of fast detection times, low power consumption, and electrode multifunctionality (i.e. sample manipulation, polymer electro-polymerization, and DNA detection). Contributions from microfluidics and MEMS fabrication allow for EC sensors to be integrated with relevant sample handling and fluidic processes on a portable diagnostic platform, which enables for rapid, multiplexed, and high throughput analysis. With further development and integration of emerging technologies, EC DNA sensors will become more prominent clinical diagnostic tools for detecting a broad spectrum of genetic-related diseases and conditions.
Abbreviations
- CNT:
-
carbon nanotube
- EBL:
-
electron-beam lithography
- EC:
-
electrochemical
- EIS:
-
electrochemical impedance spectroscopy
- MEMS:
-
micro-electro-mechanical systems
- POC:
-
point-of-care
- SiNW:
-
silicon nanowire
- SNR:
-
signal-to-noise ratio
- Vm :
-
melting potential
References
Yang S, Rothman RE 2004 PCR-based diagnostics for infectious diseases: uses, limitations, and future applications in acute-care settings. Lancet Infect Dis 4: 337–348
Beaudet AL, Belmont JW 2008 Array-based DNA diagnostics: let the revolution begin. Annu Rev Med 59: 113–129
Roberts DG, Morrison TB, Liu-Cordero SN, Cho J, Garcia J, Kanigan TS, Munnelly K, Brenan CJ 2009 A nanoliter fluidic platform for large-scale single nucleotide polymorphism genotyping. Biotechniques 46: IX–XIII
Tost J, Gut IG 2005 Genotyping single nucleotide polymorphisms by MALDI mass spectrometry in clinical applications. Clin Biochem 38: 335–350
Murphy KM, Berg KD 2003 Mutation and single nucleotide polymorphism detection using temperature gradient capillary electrophoresis. Expert Rev Mol Diagn 3: 811–818
Zhang J, Wan Y, Wang LH, Song SP, Fan CH 2007 The electrochemical DNA biosensor. Progr Chem 19: 1576–1584
Cagnin S, Caraballo M, Guiducci C, Martini P, Ross M, SantaAna M, Danley D, West T, Lanfranchi G 2009 Overview of electrochemical DNA biosensors: new approaches to detect the expression of life. Sensors 9: 3122–3148
Ahmed MU, Hossain MM, Tamiya E 2008 Electrochemical biosensors for medical and food applications. Electroanalysis 20: 616–626
Ricci F, Bonham AJ, Mason AC, Reich NO, Plaxco KW 2009 Reagentless, electrochemical approach for the specific detection of double- and single-stranded DNA binding proteins. Anal Chem 81: 1608–1614
Wei F, Patel P, Liao W, Chaudhry K, Zhang L, Arellano-Garcia M, Hu S, Elashoff D, Zhou H, Shukla S, Shah F, Ho CM, Wong DT 2009 Electrochemical sensor for multiplex biomarkers detection. Clin Cancer Res 15: 4446–4452
Wei F, Wang JH, Liao W, Zimmermann BG, Wong DT, Ho CM 2008 Electrochemical detection of low-copy number salivary RNA based on specific signal amplification with a hairpin probe. Nucleic Acids Res 36: e65
Gau V, Wong D 2007 Oral fluid nanosensor test (OFNASET) with advanced electrochemical-based molecular analysis platform. Ann NY Acad Sci 1098: 401–410
Liepold P, Wieder H, Hillebrandt H, Friebel A, Hartwich G 2005 DNA-arrays with electrical detection: a label-free low cost technology for routine use in life sciences and diagnostics. Bioelectrochemistry 67: 143–150
Lucarelli F, Capponcelli S, Marrazza G, Sangiorgi L, Mascini M 2009 Split hybridisation probes for electrochemical typing of single-nucleotide polymorphisms. Analyst 134: 52–59
Pohlmann C, Wang YR, Humenik M, Heidenreich B, Gareis M, Sprinzl M 2009 Rapid, specific and sensitive electrochemical detection of foodborne bacteria. Biosens Bioelectron 24: 2766–2771
Wakai J, Takagi A, Nakayama M, Miya T, Miyahara T, Iwanaga T, Takenaka S, Ikeda Y, Amano M 2004 A novel method of identifying genetic mutations using an electrochemical DNA array. Nucleic Acids Res 32: e141
Zhang J, Song SP, Zhang LY, Wang LH, Wu HP, Pan D, Fan C 2006 Sequence-specific detection of femtomolar DNA via a chronocoulometric DNA sensor (CDS): effects of nanoparticle-mediated amplification and nanoscale control of DNA assembly at electrodes. J Am Chem Soc 128: 8575–8580
Lord H, Kelley SO 2009 Nanomaterials for ultrasensitive electrochemical nucleic acids biosensing. J Mater Chem 19: 3127–3134
Radwan SH, Azzazy HM 2009 Gold nanoparticles for molecular diagnostics. Expert Rev Mol Diagn 9: 511–524
Wei F, Liao W, Xu Z, Yang Y, Wong DT, Ho CM 2009 Bio/abiotic interface constructed from nanoscale DNA dendrimer and conducting polymer for ultrasensitive biomolecular diagnosis. Small 5: 1784–1790
Pandey P, Datta M, Malhotra BD 2008 Prospects of nanomaterials in biosensors. Anal Lett 41: 159–209
Kerman K, Morita Y, Takamura Y, Ozsoz M, Tamiya E 2004 Modification of Escherichia coli single-stranded DNA binding protein with gold nanoparticles for electrochemical detection of DNA hybridization. Anal Chim Acta 510: 169–174
Liao WC, Ho JA 2009 Attomole DNA electrochemical sensor for the detection of Escherichia coli O157. Anal Chem 81: 2470–2476
Ozsoz M, Erdem A, Kerman K, Ozkan D, Tugrul B, Topcuoglu N, Ekren H, Taylan M 2003 Electrochemical genosensor based on colloidal gold nanoparticles for the detection of Factor V Leiden mutation using disposable pencil graphite electrodes. Anal Chem 75: 2181–2187
Castaneda MT, Merkoci A, Pumera M, Alegret S 2007 Electrochemical genosensors for biomedical applications based on gold nanoparticles. Biosens Bioelectron 22: 1961–1967
Mao X, Liu GD 2008 Nanomaterial based electrochemical DNA biosensors and bioassays. J Biomed Nanotechnol 4: 419–431
Wang F, Hu SS 2009 Electrochemical sensors based on metal and semiconductor nanoparticles. Mikrochim Acta 165: 1–22
Xu K, Huang JR, Ye ZZ, Ying YB, Li YB 2009 Recent development of nano-materials used in DNA biosensors. Sensors 9: 5534–5557
Ferguson BS, Buchsbaum SF, Swensen JS, Hsieh K, Lou XH, Soh HT 2009 Integrated microfluidic electrochemical DNA sensor. Anal Chem 81: 6503–6508
Pedrero M, Campuzano S, Pingarron JM 2009 Electroanalytical sensors and devices for multiplexed detection of foodborne pathogen microorganisms. Sensors 9: 5503–5520
Kallioniemi OP 2001 Biochip technologies in cancer research. Ann Med 33: 142–147
Pavlovic E, Lai RY, Wu TT, Ferguson BS, Sun R, Plaxco KW, Soh HT 2008 Microfluidic device architecture for electrochemical patterning and detection of multiple DNA sequences. Langmuir 24: 1102–1107
Pohanka M, Skladai P 2008 Electrochemical biosensors—principles and applications. J Appl Biomed 6: 57–64
Bard AJ, Faulkner LR 1980 Electrochemical Methods: Fundamentals and Applications. John Wiley & Sons, New York, 1–44, 137-156
Wei F, Sun B, Guo Y, Zhao XS 2003 Monitoring DNA hybridization on alkyl modified silicon surface through capacitance measurement. Biosens Bioelectron 18: 1157–1163
Sadik OA, Aluoch AO, Zhou AL 2009 Status of biomolecular recognition using electrochemical techniques. Biosens Bioelectron 24: 2749–2765
Wang J 1999 Towards genoelectronics: electrochemical biosensing of DNA hybridization. Chem Eur J 5: 1681–1685
Jin Y 2009 Label-free monitoring of site-specific DNA cleavage by EcoRI endonuclease using cyclic voltammetry and electrochemical impedance. Anal Chim Acta 634: 44–48
Tang H, Yang X, Wang K, Tan W, Li H, He L, Liu B 2008 RNA-templated single-base mutation detection based on T4 DNA ligase and reverse molecular beacon. Talanta 75: 1388–1393
Heaton RJ, Peterson AW, Georgiadis RM 2001 Electrostatic surface plasmon resonance: direct electric field-induced hybridization and denaturation in monolayer nucleic acid films and label-free discrimination of base mismatches. Proc Natl Acad Sci USA 98: 3701–3704
Wei F, Chen CL, Zhai L, Zhang N, Zhao XS 2005 Recognition of single nucleotide polymorphisms using scanning potential hairpin denaturation. J Am Chem Soc 127: 5306–5307
Neugebauer S, Zimdars A, Liepold P, Gebala M, Schuhmann W, Hartwich G 2009 Optimization of an electrochemical DNA assay by using a 48-electrode array and redox amplification studies by means of scanning electrochemical microscopy. Chembiochem 10: 1193–1199
Kuramitz H 2009 Magnetic microbead-based electrochemical immunoassays. Anal Bioanal Chem 394: 61–69
Loaiza OA, Campuzano S, Pedrero M, Pividori MI, Garcia P, Pingarron JM 2008 Disposable magnetic DNA sensors for the determination at the attomolar level of a specific enterobacteriaceae family gene. Anal Chem 80: 8239–8245
Fan CH, Plaxco KW, Heeger AJ 2003 Electrochemical interrogation of conformational changes as a reagentless method for the sequence-specific detection of DNA. Proc Natl Acad Sci USA 100: 9134–9137
Lai RY, Plaxco KW, Heeger AJ 2007 Aptamer-based electrochemical detection of picomolar platelet-derived growth factor directly in blood serum. Anal Chem 79: 229–233
Chua JH, Chee RE, Agarwal A, Wong SM, Zhang GJ 2009 Label-free electrical detection of cardiac biomarker with complementary metal-oxide semiconductor-compatible silicon nanowire sensor arrays. Anal Chem 81: 6266–6271
Maehashi K, Matsumoto K 2009 Label-free electrical detection using carbon nanotube-based biosensors. Sensors 9: 5368–5378
Daniels JS, Pourmand N 2007 Label-free impedance biosensors: opportunities and challenges. Electroanalysis 19: 1239–1257
Pohlmann C, Humenik M, Sprinzl M 2009 Detection of bacterial 16S rRNA using multivalent dendrimer-reporter enzyme conjugates. Biosens Bioelectron 24: 3383–3386
Millan KM, Mikkelsen SR 1993 Sequence-selective biosensor for DNA-based on electroactive hybridization indicators. Anal Chem 65: 2317–2323
Wang J, Rivas G, Cai X, Palecek E, Nielsen P, Shiraishi H, Dontha N, Luo D, Parrado C, Chicharro M, Farias PA, Valera FS, Grant DH, Ozsoz M, Flair MN 1997 DNA electrochemical biosensors for environmental monitoring. A review. Anal Chim Acta 347: 1–8
Erdem A, Kerman K, Meric B, Akarca US, Ozsoz M 1999 DNA electrochemical biosensor for the detection of short DNA sequences related to the hepatitis B virus. Electroanalysis 11: 586–588
Mascini M, Palchetti I, Marrazza G 2001 DNA electrochemical biosensors. Fresenius J Anal Chem 369: 15–22
Belluzo MS, Ribone ME, Lagier CM 2008 Assembling amperometric biosensors for clinical diagnostics. Sensors 8: 1366–1399
Hajdukiewicz J, Boland S, Kavanagh P, Nowicka A, Stojek Z, Leech D 2009 Enzyme-amplified amperometric detection of DNA using redox mediating films on gold microelectrodes. Electroanalysis 21: 342–350
Wang K, Chen JH, Chen J, Liu AL, Li GW, Luo HB, Lin XH, Chen YZ 2009 A sandwich-type electrochemical biosensor for detection of BCR/ABL fusion gene using locked nucleic acids on gold electrode. Electroanalysis 21: 1159–1166
Zuo X, Xiao Y, Plaxco K 2009 High specificity, electrochemical sandwich assays based on single aptamer sequences and suitable for the direct detection of small-molecule targets in blood and other complex matrices. J Am Chem Soc 131: 6944–6945
Zhou N, Yang T, Jiang C, Du M, Jiao K 2009 Highly sensitive electrochemical impedance spectroscopic detection of DNA hybridization based on Au-nano-CNT/PAN(nano) films. Talanta 77: 1021–1026
Liao W, Randall BA, Alba NA, Cui XT 2008 Conducting polymer-based impedimetric aptamer biosensor for in situ detection. Anal Bioanal Chem 392: 861–864
Liao W, Cui XT 2007 Reagentless aptamer based impedance biosensor for monitoring a neuro-inflammatory cytokine PDGF. Biosens Bioelectron 23: 218–224
Vamvakaki V, Chaniotakis NA 2008 DNA stabilization and hybridization detection on porous silicon surface by EIS and total reflection FT-IR spectroscopy. Electroanalysis 20: 1845–1850
Suni II 2008 Impedance methods for electrochemical sensors using nanomaterials. Trends Anal Chem 27: 604–611
Feng Y, Yang T, Zhang W, Jiang C, Jiao K 2008 Enhanced sensitivity for deoxyribonucleic acid electrochemical impedance sensor: gold nanoparticle/polyaniline nanotube membranes. Anal Chim Acta 616: 144–151
K'Owino IO, Sadik OA 2005 Impedance spectroscopy: a powerful tool for rapid biomolecular screening and cell culture monitoring. Electroanalysis 17: 2101–2113
Li J, Koehne JE, Cassell AM, Chen H, Ng HT, Ye Q, Fan W, Han J, Meyyappan M 2005 Inlaid multi-walled carbon nanotube nanoelectrode arrays for electroanalysis. Electroanalysis 17: 15–27
Tiwari A, Gong SQ 2009 Electrochemical detection of a breast cancer susceptible gene using cDNA immobilized chitosan-co-polyaniline electrode. Talanta 77: 1217–1222
Deng CY, Chen JH, Nie Z, Wang MD, Chu XC, Chen XL, Xiao XL, Lei CY, Yao SZ 2009 Impedimetric aptasensor with femtomolar sensitivity based on the enlargement of surface-charged gold nanoparticles. Anal Chem 81: 739–745
Li G, Li X, Wan J, Zhang S 2009 Dendrimers-based DNA biosensors for highly sensitive electrochemical detection of DNA hybridization using reporter probe DNA modified with Au nanoparticles. Biosens Bioelectron 24: 3281–3287
Pan C, Guo M, Nie Z, Xiao X, Yao S 2009 Aptamer-based electrochemical sensor for label-free recognition and detection of cancer cells. Electroanalysis 21: 1321–1326
Zhang Z, Yang W, Wang J, Yang C, Yang F, Yang X 2009 A sensitive impedimetric thrombin aptasensor based on polyamidoamine dendrimer. Talanta 78: 1240–1245
Dong H, Cao XD, Li CM, Hu WH 2008 An in situ electrochemical surface plasmon resonance immunosensor with polypyrrole propylic acid film: comparison between SPR and electrochemical responses from polymer formation to protein immunosensing. Biosens Bioelectron 23: 1055–1062
Lucarelli F, Tombelli S, Minunni M, Marrazza G, Mascini M 2008 Electrochemical and piezoelectric DNA biosensors for hybridisation detection. Anal Chim Acta 609: 139–159
Stelzle M, Durr M, Cieplik M, Nisch W 2001 On-chip electrophoretic accumulation of DNA oligomers and streptavidin. Fresenius J Anal Chem 371: 112–119
Hestekin CN, Jakupciak JP, Chiesl TN, Kan CW, O'Connell CD, Barron AE 2006 An optimized microchip electrophoresis system for mutation detection by tandem SSCP and heteroduplex analysis for p53 gene exons 5–9. Electrophoresis 27: 3823–3835
Griess GA, Hardies SC, Serwer P 2005 Matrix conditioning for lengthened capillary DNA sequencing. Electrophoresis 26: 102–111
Nagarajan R, Liu W, Kumar J, Tripathy SK, Bruno FF, Samuelson LA 2001 Manipulating DNA conformation using intertwined conducting polymer chains. Macromolecules 34: 3921–3927
Han FT, Huynh BH, Ma YF, Lin BC 1999 High efficiency DNA separation by capillary electrophoresis in a polymer solution with ultralow viscosity. Anal Chem 71: 2385–2389
Lubin AA, Hunt BV, White RJ, Plaxco KW 2009 Effects of probe length, probe geometry, and redox-tag placement on the performance of the electrochemical E-DNA sensor. Anal Chem 81: 2150–2158
Liu C, De Palma R, Reekmans G, Laureyn W, Stakenborg T, Lagae L 2009 Discrimination of specific and non-specific bindings by dielectrophoretic repulsion in on-chip magnetic bio-assays. Biosens Bioelectron 24: 2294–2297
Wang TH, Peng YH, Zhang CY, Wong PK, Ho CM 2005 Single-molecule tracing on a fluidic microchip for quantitative detection of low-abundance nucleic acids. J Am Chem Soc 127: 5354–5359
Kelley SO, Barton JK, Jackson NM, McPherson LD, Potter AB, Spain EM, Allen MJ, Hill MG 1998 Orienting DNA helices on gold using applied electric fields. Langmuir 14: 6781–6784
Sin ML, Gau V, Liao JC, Haake DA, Wong PK 2009 Active manipulation of quantum dots using AC electrokinetics. J Phys Chem C 113: 6561–6565
Wei F, Qu P, Zhai L, Chen CL, Wang HF, Zhao XS 2006 Electric potential induced dissociation of hybridized DNA with hairpin motif immobilized on silicon surface. Langmuir 22: 6280–6285
Suzuki H 2000 Advances in the microfabrication of electrochemical sensors and systems. Electroanalysis 12: 703–715
Zhang S, Wright G, Yang Y 2000 Materials and techniques for electrochemical biosensor design and construction. Biosens Bioelectron 15: 273–282
Becker H, Gartner C 2008 Polymer microfabrication technologies for microfluidic systems. Anal Bioanal Chem 390: 89–111
Gao Z, Agarwal A, Trigg AD, Singh N, Fang C, Tung CH, Fan Y, Buddharaju KD, Kong J 2007 Silicon nanowire arrays for label-free detection of DNA. Anal Chem 79: 3291–3297
Hahm J, Lieber CM 2004 Direct ultrasensitive electrical detection of DNA and DNA sequence variations using nanowire nanosensors. Nano Lett 4: 51–54
Li Z, Chen Y, Li X, Kamins TI, Nauka K, Williams RS 2004 Sequence-specific label-free DNA sensors based on silicon nanowires. Nano Lett 4: 245–247
Lee H, Park J, Kim J, Jung H, Kawai T 2006 Well-oriented nanowell array metrics for integrated digital nanobiosensors. Appl Phys Lett 89: 113901
Yang M, Yau H, Chan H 1998 Adsorption kinetics and ligand-binding properties of thiol-modified double-stranded DNA on a gold surface. Langmuir 14: 6121–6129
Yang W, Gerasimov JY, Lai RY 2009 Folding-based electrochemical DNA sensor fabricated on a gold-plated screen-printed carbon electrode. Chem Commun (Camb) 2902–2904
Wu JT, Yin H, Zhou JZ, Jin L, Lin ZH 1997 Electrochemical behaviors of DNA at mercury film electrode. Bioelectrochem Bioenerg 44: 151–154
Wang X, Ozkan CS 2008 Multisegment nanowire sensors for the detection of DNA molecules. Nano Lett 8: 398–404
Lapierre-Devlin MA, Asher CL, Taft BJ, Gasparac R, Roberts MA, Kelley SO 2005 Amplified electrocatalysis at DNA-modified nanowires. Nano Lett 5: 1051–1055
Chang H, Yuan Y, Shi N, Guan Y 2007 Electrochemical DNA biosensor based on conducting polyaniline nanotube array. Anal Chem 79: 5111–5115
Baughman RH, Zakhidov AA, de Heer WA 2002 Carbon nanotubes—the route toward applications. Science 297: 787–792
Iijima S, Ichihashi T 1993 Single-shell carbon nanotubes of 1-nm diameter. Nature 363: 603–605
He P, Dai L 2004 Aligned carbon nanotube-DNA electrochemical sensors. Chem Commun (Camb) 348–349
Wang SG, Wang RL, Sellin PJ, Zhang Q 2004 DNA biosensors based on self-assembled carbon nanotubes. Biochem Biophys Res Commun 325: 1433–1437
Heller I, Kong J, Heering HA, Williams KA, Lemay SG, Dekker C 2005 Individual single-walled carbon nanotubes as nanoelectrodes for electrochemistry. Nano Lett 5: 137–142
Koehne J, Chen H, Li J, Cassell AM, Ye Q, Ng HT, Han J, Meyyappan M 2003 Ultrasensitive label-free DNA analysis using an electronic chip based on carbon nanotube nanoelectrode arrays. Nanotechnology 14: 1239–1245
Li J, Ng HT, Cassell A, Fan W, Chen H, Ye Q, Koehne J, Han J, Meyyappan M 2003 Carbon nanotube nanoelectrode array for ultrasensitive DNA detection. Nano Lett 3: 597–602
Azek F, Grossiord C, Joannes M, Limoges B, Brossier P 2000 Hybridization assay at a disposable electrochemical biosensor for the attomole detection of amplified human cytomegalovirus DNA. Anal Biochem 284: 107–113
Wang J, Rivas G, Ozsos M, Grant DH, Cai XH, Parrado C 1997 Microfabricated electrochemical sensor for the detection of radiation-induced DNA damage. Anal Chem 69: 1457–1460
Wang J, Cai XH, Rivas G, Shiraishi H, Farias PA, Dontha N 1996 DNA electrochemical biosensor for the detection of short DNA sequences related to the human immunodeficiency virus. Anal Chem 68: 2629–2634
Ye Y, Ju H 2005 Rapid detection of ssDNA and RNA using multi-walled carbon nanotubes modified screen-printed carbon electrode. Biosens Bioelectron 21: 735–741
Cai H, Cao XN, Jiang Y, He PG, Fang YZ 2003 Carbon nanotube-enhanced electrochemical DNA biosensor for DNA hybridization detection. Anal Bioanal Chem 375: 287–293
Yang Y, Wang Z, Yang M, Li J, Zheng F, Shen G, Yu R 2007 Electrical detection of deoxyribonucleic acid hybridization based on carbon-nanotubes/nano zirconium dioxide/chitosan-modified electrodes. Anal Chim Acta 584: 268–274
Zhu NN, Chang Z, He PG, Fang YZ 2005 Electrochemical DNA biosensors based on platinum nanoparticles combined carbon nanotubes. Anal Chim Acta 545: 21–26
Jiang C, Yang T, Jiao K, Gao H 2008 A DNA electrochemical sensor with poly-L-lysine/single-walled carbon nanotubes films and its application for the highly sensitive EIS detection of PAT gene fragment and PCR amplification of NOS gene. Electrochim Acta 53: 2917–2924
Ramanavicius A, Ramanaviciene A, Malinauskas A 2006 Electrochemical sensors based on conducting polymer—polypyrrole. Electrochim Acta 51: 6025–6037
Shen Q, Wang X, Fu D 2008 The amplification effect of functionalized gold nanoparticles on the binding of anticancer drug dacarbazine to DNA and DNA bases. Appl Surf Sci 255: 577–580
Pinijsuwan S, Rijiravanich P, Somasundrum M, Surareungchai W 2008 Sub-femtomolar electrochemical detection of DNA hybridization based on latex/gold nanoparticle-assisted signal amplification. Anal Chem 80: 6779–6784
Rijiravanich P, Somasundrum M, Surareungchai W 2008 Femtomolar electrochemical detection of DNA hybridization using hollow polyelectrolyte shells bearing silver nanoparticles. Anal Chem 80: 3904–3909
Malinowski DP 2007 Multiple biomarkers in molecular oncology. II. Molecular diagnostics applications in breast cancer management. Expert Rev Mol Diagn 7: 269–280
Malinowski DP 2007 Multiple biomarkers in molecular oncology. I. Molecular diagnostics applications in cervical cancer detection. Expert Rev Mol Diagn 7: 117–131
Liao JC, Mastali M, Li Y, Gau V, Suchard MA, Babbitt J, Gornbein J, Landaw EM, McCabe ER, Churchill BM, Haake DA 2007 Development of an advanced electrochemical DNA biosensor for bacterial pathogen detection. J Mol Diagn 9: 158–168
Soper SA, Brown K, Ellington A, Frazier B, Garcia-Manero G, Gau V, Gutman SI, Hayes DF, Korte B, Landers JL, Larson D, Ligler F, Majumdar A, Mascini M, Nolte D, Rosenzweig Z, Wang J, Wilson D 2006 Point-of-care biosensor systems for cancer diagnostics/prognostics. Biosens Bioelectron 21: 1932–1942
Nel A, Xia T, Madler L, Li N 2006 Toxic potential of materials at the nanolevel. Science 311: 622–627
Singh N, Manshian B, Jenkins GJ, Griffiths SM, Williams PM, Maffeis TG, Wright CJ, Doak SH 2009 NanoGenotoxicology: the DNA damaging potential of engineered nanomaterials. Biomaterials 30: 3891–3914
Variola F, Vetrone F, Richert L, Jedrzejowski P, Yi JH, Zalzal S, Clair S, Sarkissian A, Perepichka DF, Wuest JD, Rosei F, Nanci A 2009 Improving biocompatibitity of implantable metals by nanoscale modification of surfaces: an overview of strategies, fabrication methods, and challenges. Small 5: 996–1006
Zhou Y 2008 Lipid nanotubes: formation, templating nanostructures and drug nanocarriers. Crit Rev Solid State Mater Sci 33: 183–196
Bianco A, Kostarelos K, Prato M 2008 Opportunities and challenges of carbon-based nanomaterials for cancer therapy. Expert Opin Drug Deliv 5: 331–342
Rodriguez-Cabello JC, Prieto S, Reguera J, Arias FJ, Ribeiro A 2007 Biofunctional design of elastin-like polymers for advanced applications in nanobiotechnology. J Biomater Sci Polym Ed 18: 269–286
Banta S, Megeed Z, Casali M, Rege K, Yarmush ML 2007 Engineering protein and peptide building blocks for nanotechnology. J Nanosci Nanotechnol 7: 387–401
Acknowledgements
We thank Dr. T.S. Wong for his useful comments in reviewing the manuscript.
Author information
Authors and Affiliations
Corresponding author
Additional information
Supported by NASA/National Space Biomedical Research Institute (NSBRI) (TD01301), NIH/National Institute of Dental and Craniofacial Research (NIDCR) (U01 DE017790), NIH/NIDCR (DE007296), and NIH/Nanomedicine Development Center (NDC) (5PN2ey018228:03).
Rights and permissions
About this article
Cite this article
Wei, F., Lillehoj, P. & Ho, CM. DNA Diagnostics: Nanotechnology-Enhanced Electrochemical Detection of Nucleic Acids. Pediatr Res 67, 458–468 (2010). https://doi.org/10.1203/PDR.0b013e3181d361c3
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1203/PDR.0b013e3181d361c3
This article is cited by
-
Neuro-nanotechnology: diagnostic and therapeutic nano-based strategies in applied neuroscience
BioMedical Engineering OnLine (2023)
-
Aptamer-based rapid diagnosis for point-of-care application
Microfluidics and Nanofluidics (2023)
-
Zinc selenide based dual-channel SPR optical biosensor for HIV genome DNA hybridization detection
Optical and Quantum Electronics (2023)
-
DNA hybridization detection using graphene-MoSe2–Ag heterostructure-based surface plasmon resonance biosensor
Applied Physics A (2021)
-
Electrochemical DNA detection of hepatitis E virus genotype 3 using PbS quantum dot labelling
Analytical and Bioanalytical Chemistry (2021)