Abstract
The topic of the special series of reviews in this issue will be nanobiology and nanomedicine, with a focus on the impact of nanotechnology on children and their health; hence, the title of this collection and this introduction, Nanopediatrics: Enabling Personalized Medicine for Children. We will address what is meant when we discuss these nanodisciplines and why we developed a NanoPediatrics Program at University of California, Los Angeles. We will consider the implications of these nanodisciplines for individuals and society. The nature of research, diagnosis, and screening in nanomedicine and nanopediatrics will be illustrated by selected projects in nanodiagnostics and nanotherapeutics by our group and our collaborators, and the combined use of diagnostics and therapeutics in a single nanodevice referred to as “theranostics.” We will conclude this introductory review with a summary of the reasons for developing the discipline of nanopediatrics.
Similar content being viewed by others
Main
Why Nanopediatrics?
In a talk on December 26, 1959, to the American Physical Society at Cal Tech that was subsequently published in 1961 as a chapter entitled “There's Plenty of Room at the Bottom,” Richard P. Feynman is credited with developing the concept of nanotechnology (1). Among the examples he cited was the use of DNA for information storage at the molecular level.
Nano, as in nanotechnology, nanobiology, nanomedicine, and nanopediatrics, refers to phenomena at the nanometer or molecular level. A nanometer (nm) is one-billionth of 1 m, which is an extremely small linear measurement. To put this in perspective, a typical human hair is ∼100,000 nm in diameter. An adenovirus is 90-nm wide.
Nanotechnology refers to the manipulation of materials at the atomic or molecular scale, typically 100–150 nm or less, with the National Institutes of Health recently codifying the nanoscale as ≤100 nm (http://www.asee.org/conferences/erc/presentations/upload/nanotechnology-at-the-national-institutes-of-health.pdf). Some have said nanotechnology will be similar to the “plastics revolution” of the 1950s and 1960s, in which plastics transformed the appearance and manufacturing of everything from household items and appliances to medical devices. There are numerous examples of products that currently use nanomaterials, including cosmetics, transparent sunscreens, and spill-proof fabrics.
Nanomedicine represents medicine at the nanotechnology scale with typical examples being nanoparticles and nanodevices. The economic force of nanomedicine is predicted to be substantial, with Kosterelos estimating the venture capital investment since 2000 at ∼$900M and noting a significant commitment by the National Institutes of Health (2). Nanomedicine sales were estimated to be $6.8B in 2004 with the market in nanotechnology enabled drug delivery estimated to reach $20.1B in 2011 (2).
Why would we want to develop a NanoPediatrics Program in the Mattel Children's Hospital, University of California, Los Angeles and collect a series of reviews on this topic for Pediatric Research? Why differentiate nanopediatrics from nanomedicine? One of the mantras of pediatrics is “Children are not small adults.” If nanotechnology and nanomedicine are anticipated to be the drivers for personalized medicine, then their promises and risks must be addressed in children. By developing this program, we believe that we are emphasizing the importance of the child in the applications of nanotechnology. In this way, we will improve children's health care by determining whether children have particular vulnerabilities to, for example, certain nanoparticles, thus benefiting them by protecting them from unanticipated and unintended consequences.
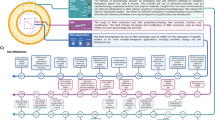
The Mattel Children's Hospital NanoPediatrics Program consists of a variety of clinical targets that are intended to meet the clinical needs of patients using distinct approaches and tools (Fig. 1). We will review a subset of these projects to introduce you to the types of applications representative of nanopediatrics.
Implications of Nanopediatrics for Individuals and Society
Genomics and the related disciplines, such as transcriptomics, proteomics, metabolomics, and other “-omics” referred to collectively as “xomics,” will enable personalized medicine. We have had success in monitoring and managing populations through the application of epidemiologic methods. Now, it is time to begin to tailor the treatment to the individual through a medical approach that will be predictive, preventive, and personalized (3). Personalized medicine will involve collection of tremendous volumes of xomic and other personal health information on each individual who engages in this medical paradigm, and many are concerned about the risks to the individual if this information is misused (4).
To prevent the misuse of this extraordinary complete body of information about the individuals, the US government has taken a variety of approaches to protect against genetic discrimination considered by many to be a new civil right (4). President Clinton signed an executive order on February 8, 2000, banning genetic discrimination in employment and health insurance for nearly all individuals in the federal workforce. President George W. Bush signed a law, the Genetic Information Nondiscrimination Act (GINA), on May 21, 2008. The Coalition for Genetic Fairness has referred to GINA as the initial legislative civil rights protection of the 21st century.
With the passage of GINA, it has been said that genetic discrimination is over and there is no need to pursue discussions on this topic (5). However, the Civil Rights Act of 1964 had broad-ranging implications, including outlawing racial segregation, extending the US Commission on Civil Rights, and creating the Equal Employment Opportunity Commission. Despite more than 25 years since passage of that landmark civil rights legislation, the US Commission on Civil Rights is still quite active, for example, announcing a public education campaign to end anti-Semitism on university campuses on April 10, 2007 (http://www.usccr.gov/campusanti-semitism.html).
Examples of Nanopediatrics Projects
Whole genome analyses.
Genomic microarrays are nanotechnology platforms. In at least one embodiment of this technology, the capture probe is a 20-bp oligomer (Fig. 2). The genome of the individual being interrogated is digested and labeled. When a sequence within the individual's DNA finds an identical match in a capture probe, the labeled genomic DNA is held onto the microarray and the label is quantified by using a very sensitive camera. This 20-bp long double helix is 6.8 nm in length and 2.0 nm in diameter.
The uses of genomic microarrays for copy number variations (CNVs) and genome-wide association studies (GWAS) represent, at this time, lower price proxys for whole genome sequencing. When the Human Genome Project was completed in 2003, 3 years were required to sequence the first genome at a cost of $2.7 billion (http://www.genome.gov/11006929). The National Human Genome Research Institute has funded a group of investigator-initiated grants that responded to the request for proposals, Revolutionary Sequencing Technologies—The $1000 Genome (http://grants.nih.gov/grants/guide/rfa-files/rfa-hg-09-011.html). It is anticipated that the goal of achieving a cost of ≤$1000 to sequence an individual's genome will be reached within the next decade. Genomic DNA mapping arrays are already successful in achieving costs less than this $1000 threshold in the research arena.
Genomic microarrays offer less expensive technological platforms compared with the current status of whole genome sequencing. These microarrays were originally designed for GWAS and therefore used single nucleotide polymorphisms (SNPs) to allow association of polymorphic markers with phenotypic traits. Any given SNP can be represented by one of three genotypes, AA, AB, or BB, where the A and B alleles are one of the two nucleotides, A, T, G, or C. Our group and others showed that the microarrays and the software used to analyze the data from them were adequate to determine whether one SNP allele was deleted, that is, to identify the presence of the A/null or B/null genotype (6–8). More recently, because these genomic microarrays have become more routinely used for assay of CNVs, array manufacturers have added to their chips copy number probes (CNPs), including probes targeting known CNV regions and additional evenly spaced CNPs, achieving median intermarker distances of <700 bp (9).
The use of genomic microarrays also represents a proxy for genomic sequencing in terms of the information and its use. For example, we are learning to live with uncertainty in terms of what represents an error versus a benign change, to improve our ability to develop efficacious correlations with clinical findings using large population studies, and to handle huge volumes of data with respect to processing and storage (3,4,10–12).
Research.
Xp21 contiguous gene deletion syndromes have been central to my research career (13), and therefore, we elected to use SNP microarrays to attempt to identify these deletions (7). We showed that “SNP chips” could be used for X chromosomal CNVs. The normalized X chromosomal copy numbers were ∼2.0 and ∼1.0 for normal women and men, respectively. With DNA from males with Xp21 deletions, the mean normalized copy number in the deleted regions was <0.4, and the reasons for detectable signal in the deleted region were unclear.
Identification of a copy number loss from a single X chromosome in an affected male would seem to be an easier discrimination than the loss of a single copy number from one of two autosomes. We chose the del22q11.2 DiGeorge syndrome to test this application and determined that representational oligonucleotide microarray analysis was effective using SNP chips (8). We showed that the choice of informatics algorithms was critical with circular binary segmentation performing better than other available algorithms. We speculated that genome-wide assessments might improve our understanding of phenotypic variability in disorders such as DiGeorge syndrome.
CNVs are present in normal individuals and even differ between monozygotic twins (14). These investigators estimated the frequency of posttwinning de novo CNVs to be as high as 5% per individual or 10% per twinning occurrence for monozygotic twins and CNVs in the ≥1 Mb range. The relationship between CNVs and disease pathogenesis remains uncertain, given their presence in phenotypically concordant and discordant monozygotic twins. Similarly, somatic tissues from the same individual may be mosaic for CNVs indicating that these events are not limited to the germline, and the CNVs involved known genes, therefore presumably affecting function (15). We know that CNVs involving the drug-metabolizing enzyme, CYP2D6, influence the metabolism of certain psychotropic drugs, with some who have duplicated/multiduplicated (up to 13 copies) CYP2D6 genes being ultrarapid metabolizers (16).
Microarrays are also valuable for their originally designated use, GWAS. SNP microarrays can complement CNVs to attempt to understand phenotypic variability, for example, as we have noted in patients with del22q11.2 (8,10), but such studies will require large groups of patients for these analyses (5,6,12–14). This approach can also be used to identify genomic regions associated with disease phenotypes. A recent example of this type of investigation involved patients with autosomal recessive (AR) hyperimmunoglobulin E recurrent infection syndrome (HEIS; OMIM 243700), a primary immune deficiency disorder characterized not only by hyper-IgE but also by eosinophilia and recurrent Staphylococcus aureus skin abscesses. Two recent reports have shown that AR HEIS is because of loss-of-function mutations in the gene encoding the protein, dedicator of cytokinesis 8 (DOCK8), which maps to the subtelomeric region of chromosome 9, 9p24 (17,18). Somewhat surprisingly, in a GWAS investigation, of the 15 affected individuals examined by microarray, 17 of 30 alleles were detected as deletions representing pathologic CNVs in >50% of alleles in these families with AR HEIS.
Diagnosis, screening, and management.
The use of microarrays as clinical nanodiagnostic tools is increasing. There are a number of academically based and corporate organizations providing these services. We believe that it is essential to link microarray testing with genetic counseling services, particularly given the uncertainty inherent in genome-wide evaluations.
The clinician is buffered from the microarray data by using algorithms that provide interpretation of the results. Inherent in these algorithms are assumptions that may profoundly influence these interpretations. For example, if identification of CNVs is a clinical target for a diagnostic microarray laboratory, then it is important that the clinician and informatician team have agreed on the threshold size above which the duplication or deletion will be called. Other assumptions embedded in the software algorithms may be more subtle, but the team must understand their implications and reach a consensus on how these assumptions will be resolved.
Other clinical applications of genomic microarrays are speculative but may well begin to be used in the very near future. One example that remains extremely controversial is the use of microarrays for identification of CNVs in the setting of newborn screening (NBS). NBS involves identification of relatively rare disorders, such as phenylketonuria (PKU; incidence 1/20–25,000 livebirths), with a cumulative incidence of ∼1.5/1000 births (In 2008 at California, 835 patients were identified from 575,645 births for incidence of 1.45055/1000 births; http://www2.uthscsa.edu/nnsis). Using standard karyotypic methods, the incidence of cytogenetic abnormalities is 1/160 or >4-fold the incidence of all currently tested NBS disorders at 6.25/1000 births (7). Although treatment for cytogenetic abnormalities may be less specific than for PKU and other traditional NBS disorders, the value of early intervention and infant stimulation for many of these patients has been clearly demonstrated. With additional high profile disorders such as autism spectrum disorders being recognized to be associated with CNVs (19–21), the clinical indications for microarray analyses are increasing. The evidence for the improved efficacy of early intervention among at least some young children and the need for better tools for earlier identification of individuals with autism (22–26) may lead to increased pressure for NBS using microarray nanotechnologies.
Management using microarrays is speculative at this time but may be used in the future for a disorder such as obesity. We know from the work of David Barker and others on the fetal origins of adult disease that the fetal epigenome is quite plastic (4). Extreme under-nutrition of the fetus in utero can lead to imprinting that increases the risk of obesity, hyperlipidemia, diabetes, and coronary artery disease, and these effects may persist throughout the lifespan of the individual. Perhaps this plasticity continues beyond birth and could be used to monitor the efficacy of surgical and medical interventions and patient compliance in obesity management protocols. SNP microarrays can be used to interrogate the epigenome, for example, by genome-wide surveys of methylation sensitivity (27). Integration of information across various xomic (where the ‘x' stands for any prefix, e.g. gen-, transcript-, etc.) parameters, such as SNP and CNV associations, mRNA expression, and epigenomic analyses, could be used to develop personalized management strategies for disorders such as obesity.
Nanoparticles
Nanoparticles are particles with diameters in the range of 100–150 nm or less. They can be used to enhance imaging, to improve therapeutic delivery, and to combine these properties in therapeutics and diagnostics referred to as “theranostics.” The following subsections will review the work of collaborators in each of these areas as potential pediatric examples of the use of nanoparticles.
Enhanced imaging.
Akhtari et al (28) generated magnetonanoparticles (MNPs) by coprecipitation of maghemite (γ-Fe2O3) and dextran that had average diameters of 10 to 20 nm with average maghemite diameters of 2 to 3 nm. They functionalized the MNPs by covalently conjugating α-methyltryptophan (AMT) to the dextran. The functionalized AMT-MNPs were shown to cross the blood-brain barrier in a rat epilepsy model and appeared to localize to epileptogenic areas, where the AMT-MNPs enhanced MRI of these foci in live animals. These investigators suggested that other bioactive ligands could be used for functional MRI to distinguish and localize abnormal brain areas. Additional molecules conjugated to these and other nanoparticles could be used to identify other pathogenic lesions, such as specific cancers.
Improved therapeutic delivery systems.
A collaboration between the Zink and Tamanoi groups is developing fluorescent mesoporous silica nanoparticles (FMSNs) as therapeutic delivery systems for use in cancer (29). They were able to demonstrate that FMSNs effectively delivered the extremely hydrophobic anticancer drug, camptothecin, to human cancer cells in culture (30,31). These FMSNs were ∼130 nm in diameter and contained 2-nm pores into which the camptothecin was loaded. The camptothecin delivered by FMSNs resulted in apoptosis of the cultured cells. Conjugation of specific targeting ligands on the surface of FMSNs can increase uptake of the particles in the cancer cells (32,33) and converting FMSNs to nanomachines, for example, by the addition of environmentally sensitive snap-tops and photocontrolled nanoimpellers, improve the regulation of drug delivery (32).
Vaults are highly conserved, naturally occurring ribonucleoprotein particles originally described by the Rome laboratory in 1986 (33). These self-assembling 13-MDa nanoparticles can be engineered to sequester specifically exogenous proteins, such as luciferase and green fluorescent protein within the vault cavity, suggesting that vaults may be able to serve as a nanocapsule that would be biocompatible (34). These 72.5 × 41 nm barrel-shaped structures can be modified by adding: cell-specific ligands to their external structure to generate recombinant delivery nanoparticles (35); a recombinant viral domain to their internal structure to facilitate delivery of a soluble ribotoxin and cDNA to cells (36); and a bacterial outer membrane protein to their internal structure to generate a “smart” vaccine delivery particle that enhanced bacterial eradication of the targeted organism in mice without the destructive inflammation associated with traditional adjuvants (37). These examples demonstrate promise for vaults as protein and cDNA delivery vehicles.
Theranostics.
Theranostics link therapeutics and diagnostics in a single agent (38). One example would be a nanoparticle that could enhance an MRI to indicate the targeting of a drug-containing nanoparticle to the desired specific cell-type or tissue, such as a malignant cancer. These would couple the diagnostic properties of functionalized MNPs (FMNPs) with the therapeutic delivery of FMSNs or vaults, and such coupling is possible. FMNPs can be modified to incorporate drugs, proteins, or DNA (Akhtari, UCLA, personal communication, 2008). FMSNs can be modified to incorporate targeting ligands for specificity in drug delivery and iron oxide nanocrystals to identify specific cells and tissues with enhanced MRI (33,34). Similarly, vaults that can serve as tissue- and cell-specific therapeutic nanoparticles can be modified to incorporate metal molecules internally (39), and these could include doped iron oxide to increase the MRI signal (40).
SUMMARY
This review has introduced the motivation leading to the development of the discipline of nanopediatrics. Because children are not small adults and nanotechnology and nanomedicine are anticipated to be major drivers of personalized medicine, it is essential that we focus the power of these technologies to enable personalized medicine for children, although recognizing that there may be differential risks in children and these cannot be ignored.
Nanoanalytical platforms for whole genome analyses will improve research, diagnosis, screening, and management, nanoparticles will enhance imaging, therapeutic delivery, and theranostics.
Many of the reviews in this special issue will build on this introduction and will focus on applications in pediatrics. However, this is a new discipline, and it is impossible to round out this topic without inviting authors beyond the field of nanopediatrics. We would hope that, a decade from now, this field will have grown so as to fill a thick volume with accomplishments in the discipline of nanopediatrics.
This issue includes a number of articles that use nanotechnology in diagnostics (41,42), in food safety (43), in improved computer technology (44), in the study of disease pathogenesis (45), and in the development of new treatments (46–49).
Abbreviations
- AR:
-
autosomal recessive
- AMT:
-
α-methyltryptophan
- CNV:
-
copy number variation
- FMSN:
-
fluorescent mesoporous silica nanoparticles
- GINA:
-
Genetic Information Nondiscrimination Act
- GWAS:
-
genomewide association studies
- HEIS:
-
hyperimmunoglobulin E recurrent infection syndrome
- MNP:
-
magneto nanoparticle
- NBS:
-
newborn screening
- SNP:
-
single nucleotide polymorphism
References
Feynman RP 1961 There's plenty of room at the bottom. In: Gilbert HD (ed) Miniaturization. Reinhold Publishing, New York, NY, pp 282–296
Kostarelos K 2006 Establishing nanomedicine. Nanomed 1: 259–260
McCabe LL, McCabe ER 2008 Expanded newborn screening: implications for genomic medicine. Annu Rev Med 59: 163–175
McCabe LL, McCabe ER 2008 DNA: Promise and Peril. University of California Press, Berkeley, CA
Hudson K 2008 Testing times for genomics. Nature 454: 163–164
McGhee SA, McCabe ER 2006 Genome-wide testing: genomic medicine. Pediatr Res 60: 243–244
Stanczak CM, Chen Z, Nelson SF, Suchard M, McCabe ER, McGhee S 2007 Deletion mapping in Xp21 for patients with complex glycerol kinase deficiency using SNP mapping arrays. Hum Mutat 28: 235–242
Stanczak CM, Chen Z, Nelson SF, Suchard M, McCabe ER, McGhee S 2008 Representational oligonucleotide microarray analysis (ROMA) and comparison of binning and change-point methods of analysis: application to detection of del22q11.2 (DiGeorge) syndrome. Hum Mutat 29: 176–181
Li X, Quigg RJ, Zhou J, Gu W, Rao NP, Reed EF 2008 Clinical utility of microarrays: current status, existing challenges and future outlook. Curr Genomics 9: 466–474
McCabe ER 2002 Translational genomics in medical genetics. Genet Med 4: 468–471
Moore TB, McCabe ER 2006 National collaborative study groups: structure, benefits gained and potential for rare genetic diseases. Genet Med 8: 793–796
Watson MS, Epstein C, Howell RR, Jones MC, Korf BR, McCabe ER, Simpson JL 2008 Developing a national collaborative study system for rare genetic diseases. Genet Med 10: 325–329
McCabe ER 2001 Disorders of glycerol metabolism. In: Scriver CR, Beaudet AL, Sly WS, Valle D (eds) The Metabolic and Molecular Basis of Disease. McGraw Hill, New York, NY, pp 2217–2237
Bruder CE, Piotrowski A, Gijbers AA, Andersson R, Erickson S, Diaz de Stahl T, Menzel U, Sandgren J, von Tell D, Poplawski A, Crowley M, Crasto C, Partridge EC, Tiwari H, Allison DB, Komorowski J, van Ommen G-J, Bosmsma DI, Pederson NL, den Dunnen JT, Wirdefeldt K, Dumanski JP 2008 Phenotypically concordant and discordant monozygotic twins display different DNA copy-number-variation profiles. Am J Hum Genet 82: 763–771
Piotrowski A, Bruder CE, Andersson R, Erickson S, Diaz de Stahl T, Menzel U, Sandgren J, von Tell D, Poplawski A, Crasto C, Bogdan A, Bartoszewski R, Partridge EC, Komorowski J, Bebok Z, Krzyzanowsi M, Janokowski N, Dumanski JP 2008 Somatic mosaicism for copy number variation in differentiated human tissues. Hum Mutat 29: 1118–1124
Bertilsson L, Dahl M-L, Dalen P, Al-Shurbaji A 2002 Molecular genetics of CYP2D6: clinical relevance with focus on psychotropic drugs. Br J Clin Pharmacol 53: 111–122
Zhang Q, Davis JC, Lamborn IT, Freeman AF, Jing H, Favreau AJ, Matthews HF, Davis J, Turner ML, Uzel G, Holland SM, Su HC 2009 Combined immunodeficiency associated with DOCK8 mutations. N Engl J Med 361: 2046–2055
Engelhardt KR, McGhee S, Winkler S, Sassi A, Woellner C, Lopez-Herrera G, Chen A, Ehl S, Thiel J, Wellinghausen N, Niehues T, Weinspach S, Reisli I, Genel F, Kutukculer N, Camcioglu Y, Karakoc-Aydiner E, Keles S, Barlan I, Gennery A, Yeganeh M, Klein C, Puck JM, Holland SM, McCabe ER, Grimbacher B, Chatila T 2009 Large deletions and point mutations involving the dedicator of cytokinase 8(DOCK8) in the autosomal-recessive form of hyper-IgE syndrome. J Allergy Clin Immunol 124: 1289–1302
Gu W, Lupski JR 2008 CNV and nervous system diseases—what's new?. Cytogenet Genome Res 123: 54–64
Guilmatre A, Dubourg C, Mosca AL, Legallic S, Goldenberg A, Drouin-Garraud V, Layet V, Rosier A, Briault S, Bonnet-Brilhault F, Laumonnier F, Odent S, Le Vacon G, Joly-Helas G, David V, Bendavid C, Pinoit JM, Henry C, Impallomeni C, Germano E, Tortorella G, Di Rosa G, Barthelemy C, Andres C, Faivre L, Frebourg T, Verber PS, Campion D 2009 Recurrent rearrangements in synaptic and neurodevelopmental genes and shared biologic pathways in schizophrenia, autism, and mental redardation. Arch Gen Psychiatry 66: 947–956
Merikangas AK, Corvin AP, Gallagher L 2009 Copy-number variants in neurodevelopmental disorders: promises and challenges. Trends Genet 25: 536–544
Press LJ 2008 InfantSEE® as a portal to early intervention for autism spectrum disorders. Optometry 79: 627–630
Howlin P, Magiati I, Charman T 2009 Systematic review of early intensive behavioral interventions for children with autism. Am J Intellect Dev Disabil 114: 23–41
Altemeier WA, Altemeier LE 2009 How can early, intensive training help a genetic disorder?. Pediatr Ann 38: 167–170
Barbaro J, Dissanayake C 2009 Autism spectrum disorders in infancy and toddlerhood: a review of the evidence on early signs, early identification tools, and early diagnosis. J Dev Behav Pediatr 30: 447–459
Reichow B, Wolery M 2009 Comprehensive synthesis of early intensive behavioral interventions for children with autism based on the UCLA young autism project model. J Autism Dev Disord 39: 23–41
Kerkel K, Spandola A, Yuan E, Kosek J, Jiang L, Hod E, Li K, Murty VV, Schupf N, Vilain E, Morris M, Haghighi F, Tycko B 2008 Genomic surveys of methylation-sensitive SNP analysis identify sequence-dependent allele-specific DNA methylation. Nat Genet 40: 904–908
Akhtari M, Bragin A, Cohen M, Moats R, Brenker F, Lynch MD, Vinters HV, Engel J 2008 Functionalized magnetonanoparticles for MRI diagnosis and localization in epilepsy. Epilepsia 49: 1419–1430
Lu J, Liong M, Zink JI, Tamanoi F 2007 Mesoporous silica nanoparticles as a delivery system for hydrophobic anticancer drugs. Small 3: 1341–1346
Liong M, Lu J, Kovochich M, Xia T, Ruehm SG, Nel AE, Tamanoi F, Zink JI 2008 Multifunctional inorganic nanoparticles for imaging, targeting, and drug delivery. ACS Nano 2: 889–896
Klichko Y, Liong M, Choi E, Angelos S, Nel AE, Stoddart F, Tamanoi F, Zink JI 2009 Mesostructured silica for optical functionality, nanomachines, and drug delivery. J Am Ceram Soc 92: s2–s10
Lu J, Choi E, Tamanoi F, Zink JI 2008 Light-activated nanoimpeller-controlled drug release in cancer cells. Small 4: 421–426
Kedersha NL, Rome LH 1986 Isolation and characterization of a novel ribonucleoprotein particle: large structures contain a single species of small RNA. J Cell Biol 103: 699–709
Kickhoefer VA, Garcia Y, Mikyas Y, Johansson E, Zhou JC, Raval-Fernandes S, Minoofar P, Zink JI, Dunn B, Stewart PL, Rome LH 2005 Engineering of vault nanocapsules with enzymatic and fluorescent properties. Proc Natl Acad Sci U S A 102: 4348–4352
Kickhoefer VA, Han M, Raval-Fernandes S, Poderycki MJ, Moniz RJ, Vaccari D, Silvestry M, Stewart PL, Kelly KA, Rome LH 2009 Targeting vault nanoparticles to specific cell surface receptors. ACS Nano 3: 27–36
Lai CY, Wiethoff CM, Kickhoefer VA, Rome LH, Nemerow GR 2009 Vault nanoparticles containing an adenovirus-derived membrane lytic protein facilitate toxin and gene transfer. ACS Nano 3: 691–699
Champion CI, Kickhoefer VA, Liu G, Moniz RJ, Freed AS, Bergmann LL, Vaccari D, Raval-Fernandes S, Chan AM, Rome LH, Kelly KA 2009 A vault nanoparticle vaccine induces protective mucosal immunity. PLoS One 4: e5409: 1–12
Bhaskar S, Pollack KM, Yoshida M, Lahann 2010 Towards designer microparticles: simultaneous control of anisotropy, shape, and size. Small 6: 404–411
Goldsmith LE, Pupols M, Kickhoefer VA, Rome LH, Monbouquette HG 2009 Utilization of a protein “shuttle” to load vault nanocapsules with gold probes and proteins. ACS Nano 3: 3175–3183
Jang JT, Nah H, Lee JH, Moon SH, Kim MG, Cheon J 2009 Critical enhancements of MRI contrast and hyperthermic effects by dopant-controlled magnetic nanoparticles. Angew Chem Int Ed Engl 48: 1234–1238
Wei F, Lillehoj PB, Ho C-M 2010 DNA diagnostics: nanotechnology-enhanced electrochemical detection of nucleic acids. Pediatr Res 67: 458–468
McCabe KM, Hernandez M 2010 Molecular thermometry. Pediatr Res 67: 469–475
Davis F, Higson SP 2010 Label-free immunochemistry approach to detect and identify antibiotics in milk. Pediatr Res 67: 476–480
Maojo V, Martin-Sanchez F, Kwlikowski C, Paton AR 2010 Nanoinformatics and DNA-based computing: catalyzing nanomedicine. Pediatr Res 67: 481–489
Ciftcioglu N, McKay DS 2010 Pathological calcification and replicating calcifying-nanoparticles (CNP): general approach and correlation. Pediatr Res 67: 490–499
Machadao M, Cheng D, Tarquinio K, Webster TJ 2010 Nanotechnology: pediatric applications. Pediatr Res 67: 500–504
Sakamoto KM 2010 Protacs for treatment of cancer. Pediatr Res 67: 505–508
Denny C 2010 Targeted liposomes for treatment of cancer. Pediatr Res 67: 514–519
Roth C 2010 Urologic tissue engineering in pediatrics: from nanostructures to bladders. Pediatr Res 67: 509–513
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
McCabe, E. Nanopediatrics: Enabling Personalized Medicine for Children. Pediatr Res 67, 453–457 (2010). https://doi.org/10.1203/PDR.0b013e3181d61ed2
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1203/PDR.0b013e3181d61ed2
This article is cited by
-
Health implications of engineered nanoparticles in infants and children
World Journal of Pediatrics (2015)