Abstract
Momentum around the era of genomic medicine is building, and with it, anticipation of the benefits that whole genome analysis (personalized or individualized genomics) will bring for the provision of health care. These technologies have the potential to revolutionize genetic diagnosis; however, the expansive data generated can lead to complex or unexpected findings, sometimes complicating clinical utility and patient benefit. Here, we use our experience with whole genome scanning microarrays, an early instance of whole genome analysis already in clinical use, to highlight fundamental challenges raised by these technologies and to discuss their medical, ethical, legal and social implications. We discuss issues that physicians and healthcare professionals will face, in particular, as the resolution of testing further increases toward full genome sequence determination. We emphasize that addressing these issues now, and starting to evolve our healthcare systems in response, will be pivotal in avoiding harms and realizing the promise of these new technologies.
Similar content being viewed by others
Main
We are experiencing a gathering storm in genomics. The last few years have seen the launch of whole genome research initiatives that are revealing the nature of individual human genomic variation and its corollaries to health and disease more fully than ever before (1–4). The cost of analysis is falling rapidly, with numerous individual genomes now sequenced to different levels of completion and many more in progress (5). Meanwhile, commercial ventures are offering whole genome microarray testing directly to consumers (DTC) (6). Publicity around these projects, as well as the U.S. President Barack Obama's underscoring of genomics and personalized medicine (7), has contributed to building anticipation of the benefits that genome-wide analysis will have for the provision of health care. Nevertheless, at the same time that genomic science is producing tremendous advances, medical, ethical, legal, and social questions are stemming from the information these new technologies unveil (8–11).
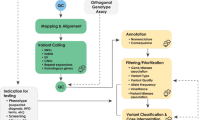
The concerns raised are perhaps the most critical in the clinical setting. In particular, the development of genome-scanning microarray technology (e.g. chromosome microarray analysis) (12,13) marks an early step in the move toward routine clinical genomics applications. These microarray-based diagnostics can readily scan the DNA for quantitative losses or gains of chromosomal segments or copy number variations (CNVs) identifying DNA anomalies that fall below the resolution of microscope-based cytogenetic analyses (14). Additionally, they have the capacity to survey variation at thousands to millions of loci at once, thus providing an assay of genomic scope combined with the sensitivity of targeted molecular testing. Importantly, the price of microarray screening is already in the $500 range nearing the cost of lower resolution methods, such as karyotyping. The unique ability to rapidly assay the entire genome in one test (at an affordable price) promises to revolutionize pre- and postnatal genetic diagnosis (15–17). However, at the same time, the massive amount of data generated will lead to unexpected findings and may complicate clinical utility and patient benefit. New paradigms to interpret and report this data and counsel individuals on its significance are urgently required.
WHOLE GENOME SCANNING DIAGNOSTICS
Microarrays can be of two types: either targeted to regions of known clinical significance or as is the focus of this piece, designed to screen the entire genome (whole genome scanning, WGS) for chromosomal abnormalities. Additionally, the sensitivity and resolution of arrays can be varied depending on the number, type, and density of their probe features. Within just a few years of being introduced into the clinical arena, there has been a steady progression toward higher resolution microarrays. Microarrays of various designs are now being used to personalize cancer treatment, screen for cancer susceptibility (18), identify etiology for constitutional disorders (19,20), conduct prenatal genetic diagnosis (21) and are increasingly being ordered to evaluate chromosomal abnormalities in specialities outside of genetics and oncology (22). Notably, DTC purveyors of personalized genomics are using the same microarray technologies; however, in this piece, we focus on issues arising from their use in the clinical realm.
Although microarray-based detection of CNVs has opened exciting potential for diagnosis and prognosis, it exemplifies a central challenge in the use of high-resolution whole genome methods. Our current understanding of detectable variation is still at an early stage (3,14,17,23,24). New CNVs continue to be discovered, and the realization that they represent a predominant form of human genomic variation, with a spectrum of phenotypic influences ranging from benign to pathogenic was only relatively recent (25–28). The presence of multiple CNVs in all human genomes has made distinguishing disease-associated variants from those without clinical significance a central challenge in medical genetics (23,29). Elsewhere, we discussed new and unforeseen complexities that the discovery of CNVs might present for genetic counseling services (30). Here, we discuss other challenges raised by WGS arrays, describing some of the issues that require further consideration, as well as some potential solutions to these problems.
NEW TECHNOLOGIES, NEW CHALLENGES
The perspective of our group is drawn from whole genome analysis technology development, CNV disease-association research and databases, clinical diagnostics, genetic counseling, and ethical, legal, and social issues, as well as policy research. Although ethical, legal, and social issues related to genetic testing are not novel (9,11), our recent experiences with this technology indicate that these questions are gaining new momentum, as medicine turns increasingly toward whole genome applications. Clinical genetic services are always assessing new technologies, but the new genome-wide technologies are producing datasets that are magnitudes larger in size and complexity. The clinical significance of much of this genomic information is not yet certain, and all these issues are compounded by an accelerated pace of discovery. As their clinical utility increases, the use of whole genome analyses will expand, providing informative data on many more individuals than are detected by conventional genetic testing (31). However, in contrast to the massive investment in the technologies, there has been little attention paid to developing practical mechanisms for translating these advances to patient care and to evaluating and managing repercussions.
Here, we present three stories created from composite patient cases to illustrate some of the most pressing issues arising. We are using high-density WGS arrays in a research setting to examine the association of CNVs in autism spectrum disorder (ASD). We anticipate similar issues will also be experienced in the clinic setting, as the current predominant use of targeted and lower density arrays shifts to the adoption of genome-wide higher resolution arrays and ultimately to full genome sequencing. Anticipating all eventualities that could arise is challenging (9). Although it will be possible to address at least some of these concerns in the informed consent process provided by individual genetic service providers, others represent much more fundamental consideration. Here, we highlight the importance of beginning to consider these issues at this early point in the trajectory of whole genome analysis and to begin to restructure our healthcare system to prepare for advances in genomic medicine.
The etiology for ASDs includes multiple genetic and possibly environmental and epigenetic influences. Recent work is providing evidence for both monogenic and complex mechanisms sometimes involving CNVs, with 10–20% of clinically diagnosed cases having an identifiable genetic association (32–37). For clarity, we have discussed each of the issues below in the context of just one of the ASD cases; however, most of them could pertain to all three stories, as well as in hundreds of other medical situations.
Story 1: Diagnostic Uncertainty
This 6-year-old boy was diagnosed with ASD at 18 mo. Karyotyping and targeted genome testing revealed no chromosomal abnormalities. The family was eager to pursue higher- resolution analysis, such that a more definitive test result might open new avenues for his clinical management, as well as to identify possible recurrent reproductive risk. Both the child's and his non-ASD parents' DNA were examined using high-density 1.8 million marker arrays (Affymetrix 6.0), which can typically detect ∼100 CNVs per sample. This analysis revealed three rare de novo (not inherited) CNVs in the child's sample and another CNV that was also present in the DNA of his father, but none of these were observed in over a thousand ancestrally matched control samples. All four CNVs were therefore considered novel variants; however, beyond being absent in controls, there was no supporting evidence yet for their involvement in ASD. Which one, if any, was responsible or contributed to the child's ASD? In this case, we were not able to determine the variant responsible for the child's condition or to explain the clinical effects of any of the four CNVs. Additionally, the unknown significance of the novel CNV in the father's sample made it impossible to accurately determine reproductive recurrence risks for the parents.
This case exemplifies some of the diagnostic unpredictabilities and uncertainties that are expected to become increasingly common with the use of higher resolution WGS, in particular, when examining complex genetic disorders. Identification of etiology has enormous potential for benefit, but at what point is the possibility of providing more information for families outweighed by uncertainty about the meaning of detected variation? This question rests on a fine balance. On one hand, the collection of large datasets from patients contributes to our knowledge of the clinical significance of CNVs, which will ultimately lead to more accurate diagnosis. On the other hand, our current understanding of these variants is still rudimentary (17).
In the current clinical genetics paradigm, the disclosure of findings is guided by the stated needs, beliefs, and expectations of each patient/family together with bioethical tenets (see the National Society of Genetic Counselors Code of Ethics at http://www.nsgc.org/about/codeEthics.cfm). If, as in this case, a family pursues higher resolution testing, our approach would be to share all reported findings in accordance with their preferences while explaining the limitations of current knowledge. Under these circumstances, most families recognize that this testing is “on the cutting edge.” However, this does not preclude some measure of discomfort on the part of clinicians when they are unable to explain test results (23) and distress for families over the meaning of unexplained variants (38). In addition, for those hoping to base reproductive choices on these results, such findings may be particularly worrisome. As we move to higher resolution methods, we can expect more variants of unknown clinical significance to be revealed. Thus, instances of uncertainty may increase until enough case and control data are accumulated and interpreting the information in an individual patient's scan becomes more straight-forward (1,31,39).
The application of WGS arrays is already significantly increasing workloads for genetic services staff and, in our experience, shifting our approach to patient care. Newer clinical arrays are producing numerous CNV calls, or “hits,” resulting in more time spent synthesizing information from disparate websites to understand if, and how, these data could be incorporated into medical management (40). To assist in interpretation, samples from parents and other family members are now more often needed and, similarly, coordinating testing and analyzing these additional datasets is adding to workloads.
Overall, the amount of information that needs to be presented to families to facilitate informed decision making is also increasing. Although many clinicians are experienced in conveying nuanced data, this material is becoming increasingly intricate, especially regarding mechanisms of complex trait inheritance, variable expressivity, and penetrance (17,41,42). In addition, higher resolution arrays are more likely to produce multiple issues of uncertainty, incidental findings, or evolving knowledge (see stories 2 and 3) within an individual case placing greater demands on healthcare professionals with respect to interpretation, communication, and case management. Administrators will need to consider and strategize accordingly for the increasing time and staffing requirements that WGS tools will likely entail.
The need for increased genomics education for clinicians (43–45) and for the training of greater numbers of genetic counselors and clinical geneticists has been noted (46). However, in addition to increasing training and education within the current genetic service paradigms, the sheer volume of data generated by high-resolution genome-wide microarray analysis will require the development of new tools (see stories 2 and 3) and counseling paradigms to interpret the data and inform patients.
To inform decision making about the effective integration of whole genome analysis, we believe it is imperative to i) begin empirical investigation of the time, personnel, educational, and economic resources that will be required for the management, explanation, and counseling of genome-wide data; ii) initiate deliberation with those engaged in providing this testing, on what uses and in what contexts these diagnostics should be most appropriately applied, and what might comprise measurable outcomes; and iii) determine best practices for effective communication/genetic counseling of this data, especially with respect to incomplete knowledge and diagnostic uncertainty.
Importantly, the establishment of centralized, user-friendly databases archiving high-resolution genome-wide datasets and integrating evolving phenotypic findings would be a fundamental step toward improving the interpretation of scans, maximizing their clinical utility, improving patient care, and smoothing the path toward broad integration of personalized genomics. The genetics and healthcare communities need to attempt to consolidate currently disparate genotype-phenotype and clinical databases as a common public resource (47).
Story 2: An Unexpected Finding
The results of another ASD family trio present another key issue raised by WGS data—that of incidental findings. We identified one de novo CNV in the child's DNA leading to hemizygous deletion of SHANK3, a gene that has recently been shown to be a presumed monogenic cause of ASD (32,48). However, in both the mother and child's samples, we also detected a CNV deletion, which overlapped a large segment of the BRCA1 tumor suppressor gene, variants that are typically associated with a lifetime risk of developing breast cancer (49). The mother, aged 24, was currently not symptomatic and did not have a family history of cancer.
Although this case was studied in the research setting where there is typically no consensus on how incidental findings should be handled (50), for the purpose of this article, we consider the multiple ethical questions that might present if such a scenario arose in the clinic. Would there be an obligation on our part to inform the mother of this incidental finding? Complex disease risk factors like BRCA1 have variable penetrance and testing for mutations in this gene is usually only carried out for women with a family history of breast or ovarian cancer. Thus, in the absence of a clinical presentation or family history, the medical rationale to convey these findings would be problematic. Informing the mother and facilitating a referral to appropriate experts would allow her and her family members who might share the variant, the choice of taking precautionary measures. But the risk of inducing significant harms existed, ranging from undue anxiety to unwarranted medical interventions.
A second set of ethical concerns revolved around the question of the child's BRCA1 status. Should this be made known to the parents? Imparting information that has immediate medical relevance, such as the confirmation of a diagnosis in a symptomatic child, has strong justification. Conversely, revealing a predisposition to adult onset disorders is contentious and can have adverse psychosocial repercussions for both the child and the family (51,52). These findings also have several legal implications in addition to the ethical and clinical concerns. Should the child, the mother, or even her family members later develop cancer, would there be grounds for litigation based on the claim that the presence of the CNV should have been conveyed when it was detected? Another potential concern is the possibility that third-party access to such genomic information could result in insurance or employment discrimination, particularly in some countries (such as Canada) where genetic information nondiscrimination laws are unclear (53). A discussion of the privacy issues surrounding genetic information is outside the scope of this article, but addressing the question of who should have access to these data and under what circumstances will be a crucial debate going forward, especially as the ability to interpret the effects of genomic variation increases (8,10,11).
After such a variant has been found, how does one present the choice to families to know or not to know? The act of posing this question may itself cause distress implying that something significant has been detected. Incidental genetic findings have long been source of ethical dilemmas in the context of conventional testing (54), but the nature of WGS data brings this issue to the fore. In the course of any genome-wide analysis, thousands of points of information about the individual are captured. Although it is not clear yet how much of this will be clinically meaningful (39), it is likely that at least some of it will have implications for the individual's health and disease susceptibility and some will have the potential to cause harm. Equally, it seems impossible, in terms of time and human and financial resources, to interpret and communicate all of this genomic information.
Similar to the other stories, this case illustrates the importance of pretest counseling during which potential outcomes should be explicitly raised and considered. Although the issue of incidental findings would be discussed, it is not always possible to anticipate what these might specifically include. Nonetheless, a framework would be in place to inform the family of the BRCA1 variants and to provide genetic counseling on the limited predictive power of such probabilistic findings, and the lack of certainty about their significance in light of the clinical presentation. Current clinical paradigms may consider it paternalistic to choose which findings merit disclosure. However, as we move forward, new systems for filtering results and determining which should be reported will be crucial. Not only will this be necessary to manage the volume of data but also it will be important for preserving the well being of patients and for addressing the needs of clinicians who will be faced with an increasing number of difficult medical and ethical decisions.
During our diagnostic investigations, multiple questions are arising. How and who will decide which data to reveal and which to mask? Is a genomic variant discovered by happenstance in an apparently healthy individual a cause for concern? How should we proceed if variants are discovered indicating carrier status or increased risk of disease later in life? Should subjective boundaries for which additional information would be acceptable to reveal be negotiated with patients before scanning? Are there some situations in which this agreement should be breached, for example, the finding of a variant of immediate medical significance, such as one conferring the risk of sudden cardiac death? What would be the mechanism for transmitting such information? Healthcare providers ordering arrays will need to anticipate their responsibilities in reviewing all findings including those outside their area of specialization and facilitating referrals to the appropriate subspecialities.
In the future, variants influencing almost all disease and carrier statuses, behavioral traits, and IQ may be known with each one carrying a different, albeit somewhat subjective, valence on a continuum from neutral (e.g. hair color) to extreme (e.g. Huntington disease status). Frameworks by which to categorize the impacts of conventional genetic test results based on clinical relevance and ability to act have been proposed (55). Similar considerations might be incorporated in the development of algorithms for the filtering, sorting, and classification of WGS results before posttest counseling. Some clinical labs have already begun to reflect on these kinds of issues and have considered blocking data from genomic regions involving adult onset conditions when testing prenatal or pediatric samples. Others are addressing the potential for incidental findings in the informed consent processes (56). However, as the information content of testing radically increases, new solutions will be required, and the practical and ethical implications of managing these data must be explicitly considered.
Story 3: A Rapidly Changing Knowledge Base
This individual was a 5-year-old girl who had been diagnosed with ASD at age 2. Analysis revealed two de novo CNVs in her genome. One of these was likely causative for the child's ASD, because it was a microdeletion overlapping the recently described ASD-susceptibility locus at the chromosome 16p11.2 (35,37,57). The other appeared to be of unknown clinical significance. A year after we had completed the study and returned the results to the family, a new study revealed an association between the cryptic CNV we had found in the child's sample and a new neurogenetic syndrome. This new information could have implications for her clinical management.
This example raises numerous issues: how will healthcare providers manage cases in the face of such a rapidly changing knowledge base? Like conventional diagnostics in which the test itself might be updated, with WGS, it is not only the quantity of data yielded that is changing but also their meaning that needs constant reinterpretation. Once WGS reaches the resolution of genome saturation, rather than having to retest samples, updating results could be achieved by simply revisiting the scan raising the question: if the information is so readily available should it be continually accessed and, if so, how? Does a clinician have an ethical responsibility to recontact the family if they become aware of new clinically relevant information? Moreover, will they be able to keep abreast of relevant findings on their patients (58,59)?
In the context of traditional genetic testing, it is generally accepted that such recontacting is desirable (60,61). However, practical considerations to operationalize such a policy and its inherent ethical complexity have prevented this from becoming a legal obligation for genetics services staff (61–63). Although guidelines have not been specifically articulated, current paradigms in the United States, United Kingdom, and Canada place the ultimate responsibility for initiating recontact for reevaluation of findings on patients or their families (64). However, at the time of such patient-clinician contact, there is a legal precedent for primary care physicians to have integrated any new information for those under their provision of care (61,64).
With the understanding of CNVs currently so labile and the publication of new disease loci (see the Online Mendelian Inheritance of Man Database at http://www.nslij-genetics.org/search_omim.html) and control information [see Database of Genomic Variants at http://projects.tcag.ca/variation/ (25)] so rapid, the potential emergence of new interpretations of clinical relevance is high and often carries ethical considerations. As an example, a new finding that alters a reproductive risk assessment for a family might sometimes be welcomed, but alternatively could be the source of great distress if the opportunity to reproduce had been passed over by a family or a termination was performed on the basis of an incorrect risk estimate. Once new relevant findings are identified, similar dilemmas to those discussed in story 2 may arise regarding the selection and prioritization of which results should be communicated.
Current clinical consent forms at our institution allow families to indicate their desire for recontact in the event that updated molecular testing might significantly alter the information previously provided. Thus, in this case the family would be informed of the new observation, as well as the implications for the patient's clinical management. However, as we move forward, how such outcomes are handled, including who will bear the responsibility for updating, how often this should be performed, and what the boundaries for such reevaluations would be, need to be considered. Certainly a patient returning to a physician about a previously identified potential health risk would expect to get up-to-date information. In the future, this could be achieved by simply reinterpreting microarray or genome-sequence findings in the context of the available knowledge base; however, at this time, few clinicians would have the knowledge, tools, or the time to reinterpret such data.
Staying abreast of a rapidly expanding body of literature for the initial interpretation of scans also applies to service providers (58). Employing teams of geneticists to manually update databases is addressing the growing challenge. However, the need for more systematic solutions to assist at all levels of interpretation and reinterpretation is becoming increasingly apparent. Web-based interfaces linked to centrally curated dynamic genotype-phenotype resources (see story 1) would assist in this regard. Such mechanisms could allow clinicians to mine datasets, and identify, update, and prioritize emerging valid and clinically relevant information at the point of need and to submit phenotypic and clinical information. Equally, service providers and researchers could use such a system to share data and interpret findings.
To inform these developments, it will be vital to i) evaluate the time, economic, and human resources that would be needed to interpret and reinterpret genome-wide datasets and reassess the clinical validity and utility of the information in the context of new research findings; and ii) explore the potential benefits and harms of updating patients with new information, including understanding, from both a clinician and patient perspective, which findings patients need to know and have a right to know, and which would be of lesser priority. Empirical evidence on the experiences, perspectives, fears, and desires of patients and families regarding updating and recontact even in the context of conventional genetic testing (61) will be pivotal to establish a proper standard of care.
Commentators have emphasized the practicality and the social and ethical utility of putting individual family and patient preferences at the heart of decisions to update genetic information (61,62). Web-based tools could also assist, providing a means by which families could update their preferences about if, and how often, they might wish to have their records updated, or even to allow them to track changes in the interpretation of their genomic data. Some DTC personalized genomics services are already offering similar informational support for their clients (65). Empirical investigation of i) stakeholder perspectives on the attributes and norms that would be desirable in new systems to manage WGS data and ii) the interest in, and the economic and technical feasibility of developing such mechanisms should be undertaken. In addition to informing healthcare professionals about the issues we have highlighted, educational resources should be made available to the public in particular for pretest counseling.
CONCLUSION
Our discussion illustrates some of the challenges that will be encountered as the resolution of WGS diagnostics increases and their clinical application expands. Whole genome analysis is already cost effective and becoming routine in clinical practice. Higher-density arrays similar to the ones from these research studies will be imminently introduced into standard clinical service.
To best prepare for the onslaught of whole genome data, there is need to i) raise awareness of the issues and establish interim guidelines to address the challenges discussed in this piece through meetings of multidisciplinary stakeholders (e.g. clinical geneticists, clinicians, scientists, genetic counselors, potentially DTC entities, and parent support groups); ii) further define unexplored clinical, ethical, legal, social, economic, and educational issues arising from advances in genomics; iii) develop algorithms and new IT resources for efficient and cost-effective management, interpretation, and reinterpretation of massive datasets; iv) establish deliberative processes, for example, by creating groups that might meet at regular intervals to discuss emerging empirical data and output action steps; and v) adapt and simplify current clinical genetic services paradigms, including consent processes. Increased understanding in each of these areas will ultimately contribute to resolving informed clinical diagnosis amid complex genomic data.
Abbreviations
- ASD:
-
Autism Spectrum Disorder
- CNV:
-
copy number variation
- DTC:
-
direct-to-consumer
- WGS:
-
whole genome scanning
References
Levy S, Sutton G, Ng PC, Feuk L, Halpern AL, Walenz BP, Axelrod N, Huang J, Kirkness EF, Denisov G, Lin Y, Macdonald JR, Pang AW, Shago M, Stockwell TB, Tsiamouri A, Bafna V, Bansal V, Kravitz SA, Busam DA, Beeson KY, McIntosh TC, Remington KA, Abril JF, Gill J, Borman J, Rogers YH, Frazier ME, Scherer SW, Strausberg RL, Venter JC 2007 The diploid genome sequence of an individual human. PLoS Biol 5: e254
Wheeler DA, Srinivasan M, Egholm M, Shen Y, Chen L, McGuire A, He W, Chen YJ, Makhijani V, Roth GT, Gomes X, Tartaro K, Niazi F, Turcotte CL, Irzyk GP, Lupski JR, Chinault C, Song XZ, Liu Y, Yuan Y, Nazareth L, Qin X, Muzny DM, Margulies M, Weinstock GM, Gibbs RA, Rothberg JM 2008 The complete genome of an individual by massively parallel DNA sequencing. Nature 452: 872–876
Bentley DR, Balasubramanian S, Swerdlow HP, Smith GP, Milton J, Brown CG, Hall KP, Evers DJ, Barnes CL, Bignell HR, Boutell JM, Bryant J, Carter RJ, Keira Cheetham R, Cox AJ, Ellis DJ, Flatbush MR, Gormley NA, Humphray SJ, Irving LJ, Karbelashvili MS, Kirk SM, Li H, Liu X, Maisinger KS, Murray LJ, Obradovic B, Ost T, Parkinson ML, Pratt MR, Rasolonjatovo IM, Reed MT, Rigatti R, Rodighiero C, Ross MT, Sabot A, Sankar SV, Scally A, Schroth GP, Smith ME, Smith VP, Spiridou A, Torrance PE, Tzonev SS, Vermaas EH, Walter K, Wu X, Zhang L, Alam MD, Anastasi C, Aniebo IC, Bailey DM, Bancarz IR, Banerjee S, Barbour SG, Baybayan PA, Benoit VA, Benson KF, Bevis C, Black PJ, Boodhun A, Brennan JS, Bridgham JA, Brown RC, Brown AA, Buermann DH, Bundu AA, Burrows JC, Carter NP, Castillo N, Chiara E, Catenazzi M, Chang S, Neil Cooley R, Crake NR, Dada OO, Diakoumakos KD, Dominguez-Fernandez B, Earnshaw DJ, Egbujor UC, Elmore DW, Etchin SS, Ewan MR, Fedurco M, Fraser LJ, Fuentes Fajardo KV, Scott Furey W, George D, Gietzen KJ, Goddard CP, Golda GS, Granieri PA, Green DE, Gustafson DL, Hansen NF, Harnish K, Haudenschild CD, Heyer NI, Hims MM, Ho JT, Horgan AM, Hoschler K, Hurwitz S, Ivanov DV, Johnson MQ, James T, Huw Jones TA, Kang GD, Kerelska TH, Kersey AD, Khrebtukova I, Kindwall AP, Kingsbury Z, Kokko-Gonzales PI, Kumar A, Laurent MA, Lawley CT, Lee SE, Lee X, Liao AK, Loch JA, Lok M, Luo S, Mammen RM, Martin JW, McCauley PG, McNitt P, Mehta P, Moon KW, Mullens JW, Newington T, Ning Z, Ling Ng B, Novo SM, O'Neill MJ, Osborne MA, Osnowski A, Ostadan O, Paraschos LL, Pickering L, Pike AC, Pike AC, Chris Pinkard D, Pliskin DP, Podhasky J, Quijano VJ, Raczy C, Rae VH, Rawlings SR, Chiva Rodriguez A, Roe PM, Rogers J, Rogert Bacigalupo MC, Romanov N, Romieu A, Roth RK, Rourke NJ, Ruediger ST, Rusman E, Sanches-Kuiper RM, Schenker MR, Seoane JM, Shaw RJ, Shiver MK, Short SW, Sizto NL, Sluis JP, Smith MA, Ernest Sohna Sohna J, Spence EJ, Stevens K, Sutton N, Szajkowski L, Tregidgo CL, Turcatti G, Vandevondele S, Verhovsky Y, Virk SM, Wakelin S, Walcott GC, Wang J, Worsley GJ, Yan J, Yau L, Zuerlein M, Rogers J, Mullikin JC, Hurles ME, McCooke NJ, West JS, Oaks FL, Lundberg PL, Klenerman D, Durbin R, Smith AJ 2008 Accurate whole human genome sequencing using reversible terminator chemistry. Nature 456: 53–59
Ley TJ, Mardis ER, Ding L, Fulton B, McLellan MD, Chen K, Dooling D, Dunford-Shore BH, McGrath S, Hickenbotham M, Cook L, Abbott R, Larson DE, Koboldt DC, Pohl C, Smith S, Hawkins A, Abbott S, Locke D, Hillier LW, Miner T, Fulton L, Magrini V, Wylie T, Glasscock J, Conyers J, Sander N, Shi X, Osborne JR, Minx P, Gordon D, Chinwalla A, Zhao Y, Ries RE, Payton JE, Westervelt P, Tomasson MH, Watson M, Baty J, Ivanovich J, Heath S, Shannon WD, Nagarajan R, Walter MJ, Link DC, Graubert TA, DiPersio JF, Wilson RK 2008 DNA sequencing of a cytogenetically normal acute myeloid leukaemia genome. Nature 456: 66–72
1000 Genomes: a deep catalog of Human Genetic Variation Project overview. Available at: http://www.1000genomes.org/page.php?page=about"> Accessed March 20, 2009
Fox JL 2008 What price personal genome exploration?. Nat Biotechnol 26: 1105–1108
Obama B 2008 US election: questioning the candidates. Interview by Alexandra Witze. Nature 455: 446–449
McGuire AL, Caulfield T, Cho MK 2008 Research ethics and the challenge of whole-genome sequencing. Nat Rev Genet 9: 152–156
Tabor HK, Cho MK 2007 Ethical implications of array comparative genomic hybridization in complex phenotypes: points to consider in research. Genet Med 9: 626–631
Robertson JA 2003 The $1000 genome: ethical and legal issues in whole genome sequencing of individuals. Am J Bioeth 3: W-IF1
Caulfield T, McGuire AL, Cho M, Buchanan JA, Burgess MM, Danilczyk U, Diaz CM, Fryer-Edwards K, Green SK, Hodosh MA, Juengst ET, Kaye J, Kedes L, Knoppers BM, Lemmens T, Meslin EM, Murphy J, Nussbaum RL, Otlowski M, Pullman D, Ray PN, Sugarman J, Timmons M 2008 Research ethics recommendations for whole-genome research: consensus statement. PLoS Biol 6: e73
Carter NP 2007 Methods and strategies for analyzing copy number variation using DNA microarrays. Nat Genet 39: S16–S21
Beaudet AL, Belmont JW 2008 Array-based DNA diagnostics: let the revolution begin. Annu Rev Med 59: 113–129
Feuk L, Carson AR, Scherer SW 2006 Structural variation in the human genome. Nat Rev Genet 7: 85–97
Bejjani BA, Shaffer LG 2006 Application of array-based comparative genomic hybridization to clinical diagnostics. J Mol Diagn 8: 528–533
Veltman JA, de Vries BB 2006 Diagnostic genome profiling: unbiased whole genome or targeted analysis? J Mol Diagn 8: 534–537; discussion 537–539.
Buchanan JA, Scherer SW 2008 Contemplating effects of genomic structural variation. Genet Med 10: 639–647
Shlien A, Tabori U, Marshall CR, Pienkowska M, Feuk L, Novokmet A, Nanda S, Druker H, Scherer SW, Malkin D 2008 Excessive genomic DNA copy number variation in the Li-Fraumeni cancer predisposition syndrome. Proc Natl Acad Sci USA 105: 11264–11269
Vissers LE, de Vries BB, Osoegawa K, Janssen IM, Feuth T, Choy CO, Straatman H, van der Vliet W, Huys EH, van Rijk A, Smeets D, van Ravenswaaij-Arts CM, Knoers NV, van der Burgt I, de Jong PJ, Brunner HG, van Kessel AG, Schoenmakers EF, Veltman JA 2003 Array-based comparative genomic hybridization for the genomewide detection of submicroscopic chromosomal abnormalities. Am J Hum Genet 73: 1261–1270
de Vries BB, Pfundt R, Leisink M, Koolen DA, Vissers LE, Janssen IM, Reijmersdal S, Nillesen WM, Huys EH, Leeuw N, Smeets D, Sistermans EA, Feuth T, van Ravenswaaij-Arts CM, van Kessel AG, Schoenmakers EF, Brunner HG, Veltman JA 2005 Diagnostic genome profiling in mental retardation. Am J Hum Genet 77: 606–616
Sahoo T, Cheung SW, Ward P, Darilek S, Patel A, del Gaudio D, Kang SH, Lalani SR, Li J, McAdoo S, Burke A, Shaw CA, Stankiewicz P, Chinault AC, Van den Veyver IB, Roa BB, Beaudet AL, Eng CM 2006 Prenatal diagnosis of chromosomal abnormalities using array-based comparative genomic hybridization. Genet Med 8: 719–727
GeneTests Home Page. GeneTests. Available at: http://www.genetests.com/. Accessed March 20, 2009
Lee C, Iafrate AJ, Brothman AR 2007 Copy number variations and clinical cytogenetic diagnosis of constitutional disorders. Nat Genet 39: S48–S54
Wang J, Wang W, Li R, Li Y, Tian G, Goodman L, Fan W, Zhang J, Li J, Zhang J, Guo Y, Feng B, Li H, Lu Y, Fang X, Liang H, Du Z, Li D, Zhao Y, Hu Y, Yang Z, Zheng H, Hellmann I, Inouye M, Pool J, Yi X, Zhao J, Duan J, Zhou Y, Qin J, Ma L, Li G, Yang Z, Zhang G, Yang B, Yu C, Liang F, Li W, Li S, Li D, Ni P, Ruan J, Li Q, Zhu H, Liu D, Lu Z, Li N, Guo G, Zhang J, Ye J, Fang L, Hao Q, Chen Q, Liang Y, Su Y, San A, Ping C, Yang S, Chen F, Li L, Zhou K, Zheng H, Ren Y, Yang L, Gao Y, Yang G, Li Z, Feng X, Kristiansen K, Wong GK, Nielsen R, Durbin R, Bolund L, Zhang X, Li S, Yang H, Wang J 2008 The diploid genome sequence of an Asian individual. Nature 456: 60–65
Iafrate AJ, Feuk L, Rivera MN, Listewnik ML, Donahoe PK, Qi Y, Scherer SW, Lee C 2004 Detection of large-scale variation in the human genome. Nat Genet 36: 949–951
Sebat J, Lakshmi B, Troge J, Alexander J, Young J, Lundin P, Maner S, Massa H, Walker M, Chi M, Navin N, Lucito R, Healy J, Hicks J, Ye K, Reiner A, Gilliam TC, Trask B, Patterson N, Zetterberg A, Wigler M 2004 Large-scale copy number polymorphism in the human genome. Science 305: 525–528
Redon R, Ishikawa S, Fitch KR, Feuk L, Perry GH, Andrews TD, Fiegler H, Shapero MH, Carson AR, Chen W, Cho EK, Dallaire S, Freeman JL, Gonzalez JR, Gratacos M, Huang J, Kalaitzopoulos D, Komura D, MacDonald JR, Marshall CR, Mei R, Montgomery L, Nishimura K, Okamura K, Shen F, Somerville MJ, Tchinda J, Valsesia A, Woodwark C, Yang F, Zhang J, Zerjal T, Zhang J, Armengol L, Conrad DF, Estivill X, Tyler-Smith C, Carter NP, Aburatani H, Lee C, Jones KW, Scherer SW, Hurles ME 2006 Global variation in copy number in the human genome. Nature 444: 444–454
Kidd JM, Cooper GM, Donahue WF, Hayden HS, Sampas N, Graves T, Hansen N, Teague B, Alkan C, Antonacci F, Haugen E, Zerr T, Yamada NA, Tsang P, Newman TL, Tuzun E, Cheng Z, Ebling HM, Tusneem N, David R, Gillett W, Phelps KA, Weaver M, Saranga D, Brand A, Tao W, Gustafson E, McKernan K, Chen L, Malig M, Smith JD, Korn JM, McCarroll SA, Altshuler DA, Peiffer DA, Dorschner M, Stamatoyannopoulos J, Schwartz D, Nickerson DA, Mullikin JC, Wilson RK, Bruhn L, Olson MV, Kaul R, Smith DR, Eichler EE 2008 Mapping and sequencing of structural variation from eight human genomes. Nature 453: 56–64
Rodriguez-Revenga L, Mila M, Rosenberg C, Lamb A, Lee C 2007 Structural variation in the human genome: the impact of copy number variants on clinical diagnosis. Genet Med 9: 600–606
Daar AS, Scherer SW, Hegele RA 2006 Implications of copy number variation in the human genome: a time for questions. Nat Rev Genet 7: 414
Buchanan JA, Carson AR, Chitayat D, Malkin D, Meyn MS, Ray PN, Shuman C, Weksberg R, Scherer SW 2009 The cycle of genome-directed medicine. Genome Med 1: 16–23
Durand CM, Betancur C, Boeckers TM, Bockmann J, Chaste P, Fauchereau F, Nygren G, Rastam M, Gillberg IC, Anckarsater H, Sponheim E, Goubran-Botros H, Delorme R, Chabane N, Mouren-Simeoni MC, de Mas P, Bieth E, Roge B, Heron D, Burglen L, Gillberg C, Leboyer M, Bourgeron T 2007 Mutations in the gene encoding the synaptic scaffolding protein SHANK3 are associated with autism spectrum disorders. Nat Genet 39: 25–27
Autism Genome Project Consortium Szatmari P, Paterson AD, Zwaigenbaum L, Roberts W, Brian J, Liu XQ, Vincent JB, Skaug JL, Thompson AP, Senman L, Feuk L, Qian C, Bryson SE, Jones MB, Marshall CR, Scherer SW, Vieland VJ, Bartlett C, Mangin LV, Goedken R, Segre A, Pericak-Vance MA, Cuccaro ML, Gilbert JR, Wright HH, Abramson RK, Betancur C, Bourgeron T, Gillberg C, Leboyer M, Buxbaum JD, Davis KL, Hollander E, Silverman JM, Hallmayer J, Lotspeich L, Sutcliffe JS, Haines JL, Folstein SE, Piven J, Wassink TH, Sheffield V, Geschwind DH, Bucan M, Brown WT, Cantor RM, Constantino JN, Gilliam TC, Herbert M, Lajonchere C, Ledbetter DH, Lese-Martin C, Miller J, Nelson S, Samango-Sprouse CA, Spence S, State M, Tanzi RE, Coon H, Dawson G, Devlin B, Estes A, Flodman P, Klei L, McMahon WM, Minshew N, Munson J, Korvatska E, Rodier PM, Schellenberg GD, Smith M, Spence MA, Stodgell C, Tepper PG, Wijsman EM, Yu CE, Roge B, Mantoulan C, Wittemeyer K, Poustka A, Felder B, Klauck SM, Schuster C, Poustka F, Bolte S, Feineis-Matthews S, Herbrecht E, Schmotzer G, Tsiantis J, Papanikolaou K, Maestrini E, Bacchelli E, Blasi F, Carone S, Toma C, Van Engeland H, de Jonge M, Kemner C, Koop F, Langemeijer M, Hijimans C, Staal WG, Baird G, Bolton PF, Rutter ML, Weisblatt E, Green J, Aldred C, Wilkinson JA, Pickles A, Le Couteur A, Berney T, McConachie H, Bailey AJ, Francis K, Honeyman G, Hutchinson A, Parr JR, Wallace S, Monaco AP, Barnby G, Kobayashi K, Lamb JA, Sousa I, Sykes N, Cook EH, Guter SJ, Leventhal BL, Salt J, Lord C, Corsello C, Hus V, Weeks DE, Volkmar F, Tauber M, Fombonne E, Shih A 2007 Mapping autism risk loci using genetic linkage and chromosomal rearrangements. Nat Genet 39: 319–328
Sebat J, Lakshmi B, Malhotra D, Troge J, Lese-Martin C, Walsh T, Yamrom B, Yoon S, Krasnitz A, Kendall J, Leotta A, Pai D, Zhang R, Lee YH, Hicks J, Spence SJ, Lee AT, Puura K, Lehtimaki T, Ledbetter D, Gregersen PK, Bregman J, Sutcliffe JS, Jobanputra V, Chung W, Warburton D, King MC, Skuse D, Geschwind DH, Gilliam TC, Ye K, Wigler M 2007 Strong association of de novo copy number mutations with autism. Science 316: 445–449
Marshall CR, Noor A, Vincent JB, Lionel AC, Feuk L, Skaug J, Shago M, Moessner R, Pinto D, Ren Y, Thiruvahindrapduram B, Fiebig A, Schreiber S, Friedman J, Ketelaars CE, Vos YJ, Ficicioglu C, Kirkpatrick S, Nicolson R, Sloman L, Summers A, Gibbons CA, Teebi A, Chitayat D, Weksberg R, Thompson A, Vardy C, Crosbie V, Luscombe S, Baatjes R, Zwaigenbaum L, Roberts W, Fernandez B, Szatmari P, Scherer SW 2008 Structural variation of chromosomes in autism spectrum disorder. Am J Hum Genet 82: 477–488
Abrahams BS, Geschwind DH 2008 Advances in autism genetics: on the threshold of a new neurobiology. Nat Rev Genet 9: 341–355
Weiss LA, Shen Y, Korn JM, Arking DE, Miller DT, Fossdal R, Saemundsen E, Stefansson H, Ferreira MA, Green T, Platt OS, Ruderfer DM, Walsh CA, Altshuler D, Chakravarti A, Tanzi RE, Stefansson K, Santangelo SL, Gusella JF, Sklar P, Wu BL, Daly MJ, Autism Consortium 2008 Association between microdeletion and microduplication at 16p11.2 and autism. N Engl J Med 358: 667–675
Darilek S, Ward P, Pursley A, Plunkett K, Furman P, Magoulas P, Patel A, Cheung SW, Eng CM 2008 Pre- and postnatal genetic testing by array-comparative genomic hybridization: genetic counseling perspectives. Genet Med 10: 13–18
Olson MV 2008 Human genetics: Dr Watson's base pairs. Nature 452: 819–820
Scherer SW, Lee C, Birney E, Altshuler DM, Eichler EE, Carter NP, Hurles ME, Feuk L 2007 Challenges and standards in integrating surveys of structural variation. Nat Genet 39: S7–S15
Mefford HC, Sharp AJ, Baker C, Itsara A, Jiang Z, Buysse K, Huang S, Maloney VK, Crolla JA, Baralle D, Collins A, Mercer C, Norga K, de Ravel T, Devriendt K, Bongers EM, de Leeuw N, Reardon W, Gimelli S, Bena F, Hennekam RC, Male A, Gaunt L, Clayton-Smith J, Simonic I, Park SM, Mehta SG, Nik-Zainal S, Woods CG, Firth HV, Parkin G, Fichera M, Reitano S, Lo Giudice M, Li KE, Casuga I, Broomer A, Conrad B, Schwerzmann M, Raber L, Gallati S, Striano P, Coppola A, Tolmie JL, Tobias ES, Lilley C, Armengol L, Spysschaert Y, Verloo P, De Coene A, Goossens L, Mortier G, Speleman F, van Binsbergen E, Nelen MR, Hochstenbach R, Poot M, Gallagher L, Gill M, McClellan J, King MC, Regan R, Skinner C, Stevenson RE, Antonarakis SE, Chen C, Estivill X, Menten B, Gimelli G, Gribble S, Schwartz S, Sutcliffe JS, Walsh T, Knight SJ, Sebat J, Romano C, Schwartz CE, Veltman JA, de Vries BB, Vermeesch JR, Barber JC, Willatt L, Tassabehji M, Eichler EE 2008 Recurrent rearrangements of chromosome 1q21.1 and variable pediatric phenotypes. N Engl J Med 359: 1685–1699
Ballif BC, Theisen A, Coppinger J, Gowans GC, Hersh JH, Madan-Khetarpal S, Schmidt KR, Tervo R, Escobar LF, Friedrich CA, McDonald M, Campbell L, Ming JE, Zackai EH, Bejjani BA, Shaffer LG 2008 Expanding the clinical phenotype of the 3q29 microdeletion syndrome and characterization of the reciprocal microduplication. Mol Cytogenet 1: 8
Begleiter ML 2002 Training for genetic counsellors. Nat Rev Genet 3: 557–561
Burke W, Emery J 2002 Genetics education for primary-care providers. Nat Rev Genet 3: 561–566
Guttmacher AE, Porteous ME, McInerney JD 2007 Educating health-care professionals about genetics and genomics. Nat Rev Genet 8: 151–157
Silversides A 2007 The wide gap between genetic research and clinical needs. CMAJ 176: 315–316
Brenner SE 2007 Common sense for our genomes. Nature 449: 783–784
Moessner R, Marshall CR, Sutcliffe JS, Skaug J, Pinto D, Vincent J, Zwaigenbaum L, Fernandez B, Roberts W, Szatmari P, Scherer SW 2007 Contribution of SHANK3 mutations to autism spectrum disorder. Am J Hum Genet 81: 1289–1297
Bradbury AR, Olopade OI 2007 Genetic susceptibility to breast cancer. Rev Endocr Metab Disord 8: 255–267
Wolf SM, Lawrenz FP, Nelson CA, Kahn JP, Cho MK, Clayton EW, Fletcher JG, Georgieff MK, Hammerschmidt D, Hudson K, Illes J, Kapur V, Keane MA, Koenig BA, LeRoy BS, McFarland EG, Paradise J, Parker LS, Terry SF, Van Ness B, Wilfond BS 2008 Managing incidental findings in human subjects research: analysis and recommendations. J Law Med Ethics 36: 219–248
Borry P, Fryns JP, Schotsmans P, Dierickx K 2006 Carrier testing in minors: a systematic review of guidelines and position papers. Eur J Hum Genet 14: 133–138
Canadian Paediatric Society 2003 Guidelines for the genetic testing of healthy children. Paediatr Child Health 8: 42–45
Orr S 2004 Privacy of genetic information in Canada: a brief examination of the legal and ethical tools that should frame Canada's regulatory response. Can J Law Technol 3: 127–140
Wertz DC 1992 Ethical and legal implications of the new genetics: issues for discussion. Soc Sci Med 35: 495–505
Burke W, Pinsky LE, Press NA 2001 Categorizing genetic tests to identify their ethical, legal, and social implications. Am J Med Genet 106: 233–240
GeneDx DNA diagnostic experts Sample submission form. Available at: http://www.genedx.com/pdf_files/genomedx_submission_form.pdf. Accessed March 20, 2009
Christian SL, Brune CW, Sudi J, Kumar RA, Liu S, Karamohamed S, Badner JA, Matsui S, Conroy J, McQuaid D, Gergel J, Hatchwell E, Gilliam TC, Gershon ES, Nowak NJ, Dobyns WB, Cook EH Jr 2008 Novel submicroscopic chromosomal abnormalities detected in autism spectrum disorder. Biol Psychiatry 63: 1111–1117
Shaffer LG, Beaudet AL, Brothman AR, Hirsch B, Levy B, Martin CL, Mascarello JT, Rao KW, Working Group of the Laboratory Quality Assurance Committee of the American College of Medical Genetics 2007 Microarray analysis for constitutional cytogenetic abnormalities. Genet Med 9: 654–662
Lintas C, Persico AM 2009 Autistic phenotypes and genetic testing: state-of-the-art for the clinical geneticist. J Med Genet 46: 1–8
Fitzpatrick JL, Hahn C, Costa T, Huggins MJ 1999 The duty to recontact: attitudes of genetics service providers. Am J Hum Genet 64: 852–860
Hunter AG, Sharpe N, Mullen M, Meschino WS 2001 Ethical, legal, and practical concerns about recontacting patients to inform them of new information: the case in medical genetics. Am J Med Genet 103: 265–276
Knoppers BM 2001 Duty to recontact: a legal harbinger?. Am J Med Genet 103: 277
Letendre M, Godard B 2004 Expanding the physician's duty of care: a duty to recontact?. Med Law 23: 531–539
Clarke JT, Ray PN 1996 The duty to update genetic counselling. Knoppers BM, Laberge C DNA Sampling and Human Genetic Research: Ethical, Legal and Policy Aspects. Kluwer Law International, Netherlands 121–132
Navigenics. Informed Consent Health Compass: Conditions included in the service. Available at: https://www.navigenics.com/visitor/what_we_offer/our_policies/informed_consent/health_compass/#a3. Accessed March 20, 2009
Acknowledgements
We gratefully acknowledge Jocalyn Clark for her comments on the manuscript.
Author information
Authors and Affiliations
Corresponding author
Additional information
Supported by The McLaughlin-Rotman Centre for Global Health, Program on Ethics and Commercialization (MRC), and The Centre for Applied Genomics (TCAG), both funded by Genome Canada through the Ontario Genomics Institute. TCAG is also funded by the Canadian Institutes for Health Research, the Canadian Institutes for Advanced Research, the McLaughlin Centre for Molecular Medicine, the Canadian Foundation for Innovation, the Ontario Ministry of Research and Innovation, and the Hospital for Sick Children Foundation. Matching funding partners for the MRC are listed at http://www.mrcglobal.org. A.S.D. and S.W.S. are supported by the McLaughlin Centre for Molecular Medicine. S.W.S. holds the GlaxoSmithKline Pathfinder Chair in Genetics and Genomics at the University of Toronto and the Hospital for Sick Children.
Rights and permissions
About this article
Cite this article
Ali-Khan, S., Daar, A., Shuman, C. et al. Whole Genome Scanning: Resolving Clinical Diagnosis and Management Amidst Complex Data. Pediatr Res 66, 357–363 (2009). https://doi.org/10.1203/PDR.0b013e3181b0cbd8
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1203/PDR.0b013e3181b0cbd8
This article is cited by
-
The impact of unsolicited findings in clinical exome sequencing, a qualitative interview study
European Journal of Human Genetics (2021)
-
Community oncologists’ perceptions and utilization of large-panel genomic tumor testing
BMC Cancer (2021)
-
Genomic medicine goes mainstream
npj Genomic Medicine (2016)
-
Parents’ Perspectives on Variants of Uncertain Significance from Chromosome Microarray Analysis
Journal of Genetic Counseling (2016)
-
Whole-genome sequencing expands diagnostic utility and improves clinical management in paediatric medicine
npj Genomic Medicine (2016)