Abstract
The aim of this study was to examine the influence of maternal intestinal and vaginal bifidobacteria on the establishment of bifidobacteria colonizing the gut in infants. Fecal samples from 110 healthy pregnant mothers within 1 mo before delivery and their babies at 1 mo of age and 100 vaginal swabs from the mother within 7 d before delivery were collected at a maternity hospital in Fukuoka city, Japan. The fecal and vaginal samples were assayed by PCR to detect Bifidobacterium species and by real-time PCR assays to estimate the bifidobacterial number. The detection of Bifidobacterium breve in the mothers' feces was significantly associated with increases in both the bifidobacterial counts and number of Bifidobacterium species in the babies' feces. In addition, a cesarean section was significantly associated with both a decrease in the counts and diversity of bifidobacteria in the babies' feces. The number of Bifidobacterium species detected in the vaginal swabs of mothers were not associated with either the bifidobacterial counts or the diversity of bifidobacteria in the babies' feces. The most important determinants of intestinal bifidobacteria in infants were the colonization of B. breve in the mothers' gut and vaginal delivery.
Similar content being viewed by others
Main
The intestinal microbiota plays an important role in human health by exerting metabolic activities that result in the salvage of energy and absorbable nutrients, by providing a barrier for the colonization of pathogens, and by stimulating the development of the immune system (1). The development of fecal microbiota in neonates is crucial because those bacteria first colonize the sterile intestine of the neonates and thus have a large effect on the host (2). Initial colonization is also relevant to the final composition of the permanent microbiota in adults.
Although the intestinal tract is first colonized by facultative aerobes, such as Enterobacteriaceae, after birth, these aerobes stay there transiently and are soon replaced by anaerobes such as bifidobacteria (3,4). Around 1 mo after birth, the genus Bifidobacterium becomes predominant (5) and remains so in the intestine during infancy. Furthermore, bifidobacteria are considered to be the most important health-beneficial bacteria for infants. First, bifidobacteria plays an important anti-inflammatory role on the mucosal surface (6) and can provide protection against enteric and systemic disorders caused by bacterial pathogens (7). Second, probiotics, including bifidobacteria might be related to the establishment of allergies (8,9), and thus, play a role in the prevention of allergic disease (10). Third, initial intestinal microbiota, in particular bifidobacterial colonization, has been suggested to be a prerequisite for the induction of oral tolerance on the basis of investigation of oral tolerance induction to the IgE response using germ-free mice (11). Finally, the administration of bifidobacteria might be associated with fewer abnormal abdominal signs, thereby stabilizing immature intestinal microbiota in preterm infants (12).
Several studies have reported that, during and after birth, microbes from both the mother and surrounding environment colonize the gastrointestinal tract of the infant (13–15). The intestinal microbiota of infants born through a cesarean section is highly different from that of the neonates born through a vaginal delivery, indicating the significant role of mother-to-infant transmission of intestinal and vaginal microbiota in the development of neonatal intestinal microbiota (16,17). Although the development of neonatal intestinal microbiota depends on the mother's microbiota, mode of delivery, and environment, it is not clear what factors of the mothers' microbiota before delivery influence the establishment of intestinal microbiota in infants. As reported earlier, bifidobacteria are predominant in the fecal microbiota of infants and play an important role; therefore, it is important to understand how maternal intestinal and vaginal bifidobacteria influence the establishment of intestinal bifidobacteria in infants, in particular, the total count of bifidobacteria and the number of Bifidobacterium species (i.e., the diversity of Bifidobacterium species). Moreover, although vaginal microbiota is thought to be transferred to the infant and to influence early microbiota development in the infant, only a few studies focusing on this phenomenon are available.
Therefore, in this study, we examined the influence of maternal bifidobacteria in the gut and vagina on the establishment of intestinal bifidobacteria in infants at 1 mo of age, when a stable intestinal microbiota has been established.
MATERIALS AND METHODS
Study population.
Healthy pregnant women (n = 124; mean 33 ± 4 y), 1 mo before the expected delivery day, were recruited to a maternity hospital in Fukuoka city, Japan between September 2006 and January 2008. The feces collected from the mothers within 1 mo before delivery and those collected from the babies between the 3 wk of age and the 6 wk of age were included in the analysis. Samples of feces that were insufficient in amount as well as those from babies who had taken antibiotics after birth were excluded from the analysis. A total of 110 mother–infant pairs were enrolled in the study. The clinical characteristics and demographic data of the infants are summarized in Table 1. Mothers from whom vaginal swabs could not be collected within 7 d before delivery were excluded from the analysis. A total of 100 vaginal swabs were finally analyzed in this study.
Fecal and vaginal samples.
Fecal samples were collected in sterile plastic bags and immediately taken to the laboratory. All fecal samples were stored at −80°C until analysis. Vaginal swabs were collected using a cotton swab rolled over sections of the vaginal wall. The vaginal swabs were then sent to the laboratory within 24 h at 4°C and thereafter were processed for DNA extraction.
DNA extraction.
Bacterial DNA was extracted from the feces using an Ultra Clean Soli DNA Isolation Kit (Mo Bio Laboratories, Carlsbad, CA) according to the method described by Clement and Kitts (18) with some modifications (19). Briefly, the fecal sample (0.1 g) was added to a tube that contained lysozyme (25 mg/mL) and N-acetylmuramidase (0.3 mg/mL) and incubated for 30 min at 37°C for cell lysis. Next, the extracted DNA was purified using a High Pure PCR Template Preparation Kit (Roche Diagnostics GmbH, Mannheim, Germany) according to the manufacturer's instructions. The produced DNA was stored at −20°C until PCR amplification. The bacterial cells from the vagina were collected by suspending the cotton swab in 1 mL distilled water and centrifuging it at 15,000 rpm. Bacterial DNA was extracted from the collected cells using a Wizard Genomic DNA Purification Kit (Promega Corporation, Madison, WI) according to the manufacturer's instructions. The extracted DNA was stored at −20°C until PCR amplification.
Identification of Bifidobacterium species by PCR amplification.
DNA from the samples was analyzed for the presence of B. adolescentis, B. bifidum, B. breve, B. catenulatum group, B. dentium, B. infantis, or B. longum. The Bifidobacterium species-specific primer used in this study was described by Matsuki et al. (20). Each PCR mixture (50 μL) was composed of 10 μL of a 5× Taq Buffer (Promega Corporation, Madison, WI), 1.5 mM MgCl2, deoxynucleoside triphosphate at a concentration of 200 μM, species-specific primer at a concentration of 200 μM, and 1.25 U of Taq polymerase (Promega Corporation). The PCR was carried out with Mastercycler ep gradient S (Eppendorf, Hamburg, Germany). The following amplification program was used: one cycle at 94°C for 5 min, followed by 35 cycles of 94°C for 20 s, 55°C for 20 s, 72°C for 30 s, and, finally, one cycle of 72°C for 5 min. The amplified DNA was verified by electrophoresis of the PCR products in 1.5% agarose in a 1× TAE buffer. The prevalence of colonization was expressed as the percentage of positive samples in each Bifidobacterium species in the total samples.
Real-time PCR.
Real-time PCR amplification and detection were performed in an ABI PRISM 7700 sequence detection system (Applied Biosystems, Foster, CA). Bacterial DNA from the samples was subjected to real-time PCR assays on 16s rDNA gene sequences. The species-specific primer and PCR conditions for bifidobacteria used in this study were described by Matsuki et al. (21). For the detection of bifidobacteria, amplification was performed in a total volume of 25 μL, containing 12.5 μL SYBR GREEN Master Mix (Applied Biosystems, Foster, CA) and 0.25 μM of each specific primer. The amplification program had one cycle of 94°C for 5 min, 40 cycles of 94°C for 20 s, 55°C for 30 s, and 72°C for 1 min, and, finally, one step of each cycle of 94°C for 15 s. The values of log10 cells/g feces for the bifidobacteria were calculated for each stool sample from the threshold cycle values using constructed standard curves. Bacterial DNA from all vaginal samples was also subjected to a real-time assay for lactobacilli. According to the method described by Byun et al. (22), the number of bifidobacteria in the vaginal swabs was indicated semiquantitatively by the logarithmic ratio of the counts of bifidobacteria to those of lactobacilli (log10 [counts of bifidobacteria/counts of lactobacilli], B/L ratio) because the amount of vaginal swabs could not be measured.
Culture of bifidobacteria in the presence of galacto-oligosaccharides.
Strains of B. breve (n = 29) and B. longum (n = 17) obtained from the feces of babies were suspended in an EG broth at 109 CFU/mL and 1 mL of the suspension was added to a culture flask, containing 100 mL fresh EG broth. One percent (wt/vol) galacto-oligosaccharides (Oligomate 55 NP, Yakult Pharmaceutical Industry, Tokyo, Japan) was then added to the culture flask just before the start of incubation. Next, the culture was anaerobically incubated at 37°C for 24 h. At 12 and 24 h after incubation, an aliquot was removed to determine the bacterial number spectrophotometrically at 550 nm.
Statistical analysis.
Continuous variables were reported as the medians and the interquartile range or the means and SD, according to their distribution. The Wilcoxon rank test was used to compare the bifidobacterial counts between the mothers and babies. The Spearman's correlation coefficient was calculated to determine the relationship between the fecal bifidobacterial counts of the mothers and those of the babies or between the bifidobacterial counts of the vaginal swabs (B/L ratio) and those of the babies' feces. The Mann-Whitney U test was used to determine the unadjusted maternal effects on the bifidobacterial counts and the number of Bifidobacterium species in the babies' feces. The distribution of the bifidobacterial counts was not normal because it had a detection limit. The lowest value (5.0) under measurable limits was inserted in the bifidobacterial counts. A proportional odds logistic regression was used to examine the influence of the maternal determinants on the bifidobacterial counts and diversity in the babies' feces. The proportional odds logistic regression is a model for not only ordinal categorical outcome variables but also skewed continuous outcome variables using ranks of data (23). The following independent variables were included in this model: the colonization of each Bifidobacterium species in the mothers' feces, the delivery mode, and the number of Bifidobacterium species in the vaginal swabs. The proportional odds assumption was assessed with a test of parallel lines. The values of p < 0.05 were considered statistically significant. SPSS (SPSS, Chicago, IL) version 16.0 for Windows was used for the data analysis.
Ethical considerations.
This study protocol was approved by the Committee on Ethical Practice in Tokai University School of Medicine. Written informed consent was obtained from all the mothers.
RESULTS
Counts of bifidobacteria in the feces and vaginal swabs.
The median counts of bifidobacteria in the feces and vaginal swabs are shown in Figure 1. The counts of bifidobacteria in the feces of the mothers were significantly higher than those of bifidobacteria in their babies' feces (p < 0.001, Wilcoxon rank test). The counts of lactobacilli were higher than those of bifidobacteria in the vaginal swabs of the mothers (Median B/L ratio = −1.1). There was no significant correlation between the fecal bifidobacterial counts of the mothers and those of the babies or between the bifidobacterial counts of the vaginal swabs (B/L ratio) and those of the babies' feces (R = 0.054, R = −0.113, respectively, Fig. 2).
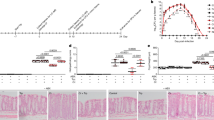
Bifidobacterial counts in the feces (n = 110) and vaginal swabs (n = 100). A, Bars represent the median of bifidobacterial counts in the feces of mothers and babies. The median (interquartile range) of mothers and babies was 9.6 (9.2–10.0) and 9.4 (7.7–9.8), respectively. *p < 0.001 (examined by the Wilcoxon rank test). B, The bar represents the median of the B/L ratio of vaginal swabs. The B/L ratio is the log10 [count of bifidobacteria/count of lactobacilli] (B/L ratio) in the vaginal swabs. The median (interquartile range) of the B/L ratio was −1.1 (−2.1 to 1.0).
Relationship between the bifidobacterial counts in the feces of babies and the bifidobacterial counts in the mothers' feces. A, Correlation between the fecal bifidobacterial counts of the mothers and those of the babies (R = 0.054, p = 0.572, Spearman's correlation coefficient). B, Correlation between the bifidobacterial counts of the vaginal swabs (B/L ratio) and those of the babies' feces (R = −0.133, p = 0.189, Spearman's correlation coefficient).
Bifidobacterium species composition in the mothers and babies.
The prevalence of the detection of each Bifidobacterium species in the samples of both the feces and vaginal swabs is shown in Table 2. In the feces of mothers, B. longum was the most common species (91.8%). In the feces of babies, on the other hand, B. breve was the most frequent species (43.6%). In the vaginal swabs of mothers, B. breve, which was the most common species found in the feces of babies, was also the most prevalent species (56.0%). To investigate why B. breve was the most frequent Bifidobacterium species in the gut of babies, the growth of B. breve strains was examined in vitro using B. longum as the control in the presence of galacto-oligosaccharide, which is a major component in mother's milk (Fig. 3). As a result, strains of B. breve isolated from the feces of babies showed a significantly higher growth than those of B. longum.
Growth of Bifidobacterium strains in the presence of galacto-oligosaccharide. Bars represent the median of O.D. strains of B. breve (n = 29) and B. longum (n = 17) obtained from the feces of babies were incubated for 12 (A) and 24 (B) h, as described in Materials and Methods section. A, The median (interquartile range) of O.D. for B. breve and B. longum was 0.53 (0.45–0.61) and 0.44 (0.35–0.49), respectively. B, The median (interquartile range) of O.D. for B. breve and B. longum was 0.51 (0.43–0.57) and 0.35 (0.27–0.43), respectively. *p = 0.008 (examined by the Mann-Whitney U test), †p < 0.001 (examined by the Mann-Whitney U test).
Maternal fecal bifidobacterial species influencing the count and diversity of fecal bifidobacteria in babies.
Table 3 shows the relationship between the Bifidobacterium species in the feces of mothers and the counts/diversity of bifidobacteria in their babies' feces. The detection of B. breve in the mothers' feces was significantly associated with increases in both the counts and diversity of bifidobacteria in the babies' feces (p = 0.018 and p < 0.001, respectively). Such significant increases of bifidobacteria in the babies' feces were also found when B. infantis was detected in the feces of mothers (p = 0.005), although the prevalence of this species was low in their feces (five of 110). Using the vaginal swabs of mothers within 7 d before delivery, the relationship between the Bifidobacterium species of the vaginal swabs and the bifidobacterial status in the feces of babies was then analyzed in the same way. However, no particular Bifidobacterium species colonizing the vagina affected the bifidobacteria status in the gut of babies (data not shown). Furthermore, the effects of the delivery mode on the bifidobacterial status in the feces of babies were analyzed (Table 3). In comparison to vaginal delivery, a cesarean section decreased the diversity in the feces of babies (p = 0.006).
Next, a proportional odds logistic regression analysis was performed to examine the proper effect of these different maternal determinants on the status of bifidobacteria in the babies' feces (Table 4). The detection of B. breve in the mothers' feces was still significantly associated with the increases in both the bifidobacterial counts and the number of Bifidobacterium species in the babies' feces (p = 0.038 and p < 0.001, respectively). A cesarean section was significantly associated with both decreases in the counts and diversity of bifidobacteria in the feces of babies (p = 0.033 and p = 0.016, respectively). In addition, the detection of B. infanits in the mothers' feces, which was a significant factor in the univariate analysis, was associated with the counts of bifidobacteria in the feces of babies (p = 0.001).
DISCUSSION
This study investigated the influence of bifidobacteria in the gut and vagina of the mothers on the establishment of intestinal bifidobacteria in their babies at the age of 1 mo. Regarding the analysis of the bifidobacterial counts, the fecal counts of bifidobacteria were significantly lower in the babies than those of the bifidobacteria in the mothers in this study. However, no such difference in the counts was found between them in the report by Grönlund et al. (24), in which one half of the mothers had received probiotic supplements during pregnancy. Considering that no probiotic supplements were administered to the mothers in this study, we can assume that the intestinal ecosystem of the baby had intrinsically less capacity to colonize bifidobacteria than that of the mother. There was no significant correlation between the counts of bifidobacteria in the mothers' feces/vaginal swabs and those in the babies' feces. This indicated that the total counts of bifidobacteria per se in the mother did not affect the bifidobacterial status of the baby, as has been reported previously (25).
In this study using PCR methods, bifidobacteria were detected in more than 80% of the vaginal swab samples, and their counts were almost comparable to those of lactobacilli, whereas just a presence of bifidobacteria in the vagina had been previously reported using a culture-dependent method (26). The vaginal epithelium contains glycogen, a multichained polysaccharide. The amount of glycogen is regulated by the secretion level of both estrogens and progesterones (26,27). Because polysaccharides selectively increase the growth of bifidobacteria (28–30), it is suggested that the vagina during pregnancy, in which female hormones are present at high levels, may therefore sustain the colonization of high counts of bifidobacteria.
Regarding the analysis of diversity of Bifidobacterium species, B. breve was the most prevalent species in both the feces of babies and the vaginal swabs of mothers, although this species was far less prevalent in the feces of mothers. Ninety-five percent of the babies in this study received breast or mixed milk containing a high concentration of galacto-oligosaccharide. Moreover, B. breve was able to grow well in the presence of galacto-oligosaccharide (Fig. 3). Therefore, the high level of galacto-oligosaccharide in the gut of babies resulted in B. breve being the most prevalent Bifidobacterium species in their guts. Glycogen and/or its degradation products may support the growth of B. breve in the vagina. The difference in the prevalence of the colonization of Bifidobacterium species in feces was reported in a previous study using German samples (30). In that study, the dominant Bifidobacterium species was B. infantis; however, this species was the third in our study. This discordance may be caused by differences in environmental and genetic backgrounds.
In the analysis of the relationship between the bifidobacterial status of the mothers and babies, the detection of B. breve in the feces of mothers was significantly associated with the increases in both the counts and diversity of bifidobacteria in the feces of babies. Because B. breve was assumed to be an appropriate species in the gut of babies, it is likely that it would rapidly increase in number if it had been colonizing the gut of mothers and had moved to that of babies during delivery or after birth. On the other hand, the colonization of B. breve and other species in the vaginal swabs did not statistically influence the counts or diversity of fecal bifidobacteria in the babies. The bifidobacterial counts in the vagina were much smaller than those in the intestine (27). Such a low number may not be able to affect the status of babies' bifidobacteria.
In this study, delivery by cesarean section had a significantly negative effect on the counts and diversity of the bifidobacteria in the babies' feces, as reported previously (16,31). Because the route of a cesarean delivery is away from the anus, it is likely that B. breve in the feces of mothers cannot gain access to the babies during delivery.
Although we analyzed the difference in the bifidobacterial counts and diversity in the gut between the newborns who were exclusively breast-fed and those who were partially breast-fed, no significant difference was found between the two groups (p = 0.992 and 0.731, respectively). We did not include these data in the analyses of this study because the data of feeding were not maternal factors regarding the time before and during delivery. In this study, our analysis was focused on the maternal determinants that affect the intestinal bifidobacterial colonization of infants.
In previous studies, the administration of a probiotic to the mothers during pregnancy had a tendency to increase the bifidobacterial diversity in infants (19). In addition, a probiotic administered to mothers during pregnancy has been recovered from the feces of infants (32). These studies indicated that probiotic consumption might render an ecosystem favorable for the growth of bifidobacteria in the gut of the mother and that such an ecosystem could be transmitted from mother to infant. Bifidobacterium species, in particular B. breve, is considered to be the most important health-beneficial bacteria for infants (6,7,12). These reports, when considered together, suggest that the administration of B. breve to mothers might increase the transmission of this species to infants.
There are some limitations associated with this study. First, only the effect of the mothers' bifidobacteria on the babies' bifidobacteria was examined. Further research is needed to clarify the effects of bacterial species other than Bifidobacterium. Second, the strain of bifidobacteria in the feces or vaginal swabs was not determined. Further study on the strain of bifidobacteria will be needed to elucidate the transmission route from mothers to infants. In conclusion, the significant determinants affecting the intestinal bifidobacterial status of infants were the colonization of B. breve in the mothers' gut and vaginal delivery.
Abbreviations
- B/L ratio:
-
Log10 [count of bifidobacteria/count of lactobacilli]
References
Guarner F, Malagelada JR 2003 Gut flora in health and disease. Lancet 361: 512–519
Caicedo RA, Schanler RJ, Li N, Neu J 2005 The developing intestinal ecosystem: implications for the neonate. Pediatr Res 58: 625–628
Rotimi VO, Duerden BI 1981 The development of the bacterial flora in normal neonates. J Med Microbiol 14: 51–62
Mackie RI, Sghir A, Gaskins HR 1999 Developmental microbial ecology of the neonatal gastrointestinal tract. Am J Clin Nutr 69: 1035S–1045S
Benno Y, Sawada K, Mitsuoka T 1984 The intestinal microflora of infants: composition of fecal flora in breast-fed and bottle-fed infants. Microbiol Immunol 28: 975–986
Fujii T, Ohtsuka Y, Lee T, Kudo T, Shoji H, Sato H, Nagata S, Shimizu T, Yamashiro Y 2006 Bifidobacterium breve enhances transforming growth factor beta1 signaling by regulating Smad7 expression in preterm infants. J Pediatr Gastroenterol Nutr 43: 83–88
Thibault H, Aubert-Jacquin C, Goulet O 2004 Effects of long-term consumption of a fermented infant formula (with Bifidobacterium breve c50 and Streptococcus thermophilus 065) on acute diarrhea in healthy infants. J Pediatr Gastroenterol Nutr 39: 147–152
Björkstén B, Sepp E, Julge K, Voor T, Mikelsaar M 2001 Allergy development and the intestinal microflora during the first year of life. J Allergy Clin Immunol 108: 516–520
He F, Ouwehand AC, Isolauri E, Hashimoto H, Benno Y, Salminen S 2001 Comparison of mucosal adhesion and species identification of bifidobacteria isolated from healthy and allergic infants. FEMS Immunol Med Microbiol 30: 43–47
Kukkonen K, Savilahti E, Haahtela T, Juntunen-Backman K, Korpela R, Poussa T, Tuure T, Kuitunen M 2007 Probiotics and prebiotic galacto-oligosaccharides in the prevention of allergic diseases: a randomized, double-blind, placebo-controlled trial. J Allergy Clin Immunol 119: 192–198
Sudo N, Sawamura S, Tanaka K, Aiba Y, Kubo C, Koga Y 1997 The requirement of intestinal bacterial flora for the development of an IgE production system fully susceptible to oral tolerance induction. J Immunol 159: 1739–1745
Kitajima H, Sumida Y, Tanaka R, Yuki N, Takayama H, Fujimura M 1997 Early administration of Bifidobacterium breve to preterm infants: randomised controlled trial. Arch Dis Child Fetal Neonatal Ed 76: F101–F107
Fryklund B, Tullus K, Berglund B, Burman LG 1992 Importance of the environment and the faecal flora of infants, nursing staff and parents as sources of gram-negative bacteria colonizing newborns in three neonatal wards. Infection 20: 253–257
Murono K, Fujita K, Yoshikawa M, Saijo M, Inyaku F, Kakehashi H, Tsukamoto T 1993 Acquisition of nonmaternal enterobacteriaceae by infants delivered in hospitals. J Pediatr 122: 120–125
Tannock GW, Fuller R, Smith SL, Hall MA 1990 Plasmid profiling of members of the family enterobacteriaceae, lactobacilli, and bifidobacteria to study the transmission of bacteria from mother to infant. J Clin Microbiol 28: 1225–1228
Grönlund MM, Lehtonen OP, Eerola E, Kero P 1999 Fecal microflora in healthy infants born by different methods of delivery: permanent changes in intestinal flora after cesarean delivery. J Pediatr Gastroenterol Nutr 28: 19–25
Chen J, Cai W, Feng Y 2007 Development of intestinal bifidobacteria and lactobacilli in breast-fed neonates. Clin Nutr 26: 559–566
Clement BG, Kitts CL 2000 Isolating PCR-quality DNA from human feces with a soil DNA kit. Biotechniques 28: 640–642, 644, 646
Gueimonde M, Sakata S, Kalliomäki M, Isolauri E, Benno Y, Salminen S 2006 Effect of maternal consumption of lactobacillus GG on transfer and establishment of fecal bifidobacterial microbiota in neonates. J Pediatr Gastroenterol Nutr 42: 166–170
Matsuki T, Watanabe K, Tanaka R, Fukuda M, Oyaizu H 1999 Distribution of bifidobacterial species in human intestinal microflora examined with 16S rRNA-gene-targeted species-specific primers. Appl Environ Microbiol 65: 4506–4512
Matsuki T, Watanabe K, Fujimoto J, Kado Y, Takada T, Matsumoto K, Tanaka R 2004 Quantitative PCR with 16S rRNA-gene-targeted species-specific primers for analysis of human intestinal bifidobacteria. Appl Environ Microbiol 70: 167–173
Byun R, Nadkarni MA, Chhour KL, Martin FE, Jacques NA, Hunter N 2004 Quantitative analysis of diverse Lactobacillus species present in advanced dental caries. J Clin Microbiol 42: 3128–3136
Ely EW, Girard TD, Shintani AK, Jackson JC, Gordon SM, Thomason JW, Pun BT, Canonico AE, Light RW, Pandharipande P, Laskowitz DT 2007 Apolipoprotein E4 polymorphism as a genetic predisposition to delirium in critically ill patients. Crit Care Med 35: 112–117
Grönlund MM, Gueimonde M, Laitinen K, Kociubinski G, Grönroos T, Salminen S, Isolauri E 2007 Maternal breast-milk and intestinal bifidobacteria guide the compositional development of the Bifidobacterium microbiota in infants at risk of allergic disease. Clin Exp Allergy 37: 1764–1772
Shadid R, Haarman M, Knol J, Theis W, Beermann C, Rjosk-Dendorfer D, Schendel DJ, Koletzko BV, Krauss-Etschmann S 2007 Effects of galactooligosaccharide and long-chain fructooligosaccharide supplementation during pregnancy on maternal and neonatal microbiota and immunity–a randomized, double-blind, placebo-controlled study. Am J Clin Nutr 86: 1426–1437
Levison ME, Corman LC, Carrington ER, Kaye D 1977 Quantitative microflora of the vagina. Am J Obstet Gynecol 127: 80–85
Redondo-Lopez V, Cook RL, Sobel JD 1990 Emerging role of lactobacilli in the control and maintenance of the vaginal bacterial microflora. Rev Infect Dis 12: 856–872
Newburg DS 2000 Oligosaccharides in human milk and bacterial colonization. J Pediatr Gastroenterol Nutr 30: S8–S17
Kunz C, Rudloff S, Baier W, Klein N, Strobel S 2000 Oligosaccharides in human milk: structural, functional, and metabolic aspects. Annu Rev Nutr 20: 699–722
Haarman M, Knol J 2005 Quantitative real-time PCR assays to identify and quantify fecal Bifidobacterium species in infants receiving a prebiotic infant formula. Appl Environ Microbiol 71: 2318–2324
Penders J, Thijs C, Vink C, Stelma FF, Snijders B, Kummeling I, van den Brandt PA, Stobberingh EE 2006 Factors influencing the composition of the intestinal microbiota in early infancy. Pediatrics 118: 511–521
Schultz M, Göttl C, Young RJ, Iwen P, Vanderhoof JA 2004 Administration of oral probiotic bacteria to pregnant women causes temporary infantile colonization. J Pediatr Gastroenterol Nutr 38: 293–297
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Mikami, K., Takahashi, H., Kimura, M. et al. Influence of Maternal Bifidobacteria on the Establishment of Bifidobacteria Colonizing the Gut in Infants. Pediatr Res 65, 669–674 (2009). https://doi.org/10.1203/PDR.0b013e31819ed7a8
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1203/PDR.0b013e31819ed7a8
This article is cited by
-
A synbiotic intervention modulates meta-omics signatures of gut redox potential and acidity in elective caesarean born infants
BMC Microbiology (2021)
-
Maternal antimicrobial use at delivery has a stronger impact than mode of delivery on bifidobacterial colonization in infants: a pilot study
Journal of Perinatology (2018)
-
Genomic diversity and distribution of Bifidobacterium longum subsp. longum across the human lifespan
Scientific Reports (2018)
-
Evolution of gut Bifidobacterium population in healthy Japanese infants over the first three years of life: a quantitative assessment
Scientific Reports (2017)
-
Diet-induced changes in maternal gut microbiota and metabolomic profiles influence programming of offspring obesity risk in rats
Scientific Reports (2016)