Abstract
The human fetus and neonate swallow biologically significant quantities of IL-8/CXC ligand 8 (CXCL8) in amniotic fluid and breast milk, and this remains measurable through simulated neonatal gastric and proximal intestinal digestions. We sought to confirm the structural and functional integrity of IL-8/CXCL8 in digestates and determine the mechanisms underlying this protease resistance. We observed that in comparison with BSA, IL-8/CXCL8 is highly resistant to pepsin and can be detected intact in assays for structural, immunologic, and functional integrity. In a computational molecular docking simulation, IL-8/CXCL8 was observed to fit poorly in the pepsin active site. On the basis of simulated mutation analyses, we hypothesized that this protease resistance is due to disulfide bond-related tertiary folding in IL-8/CXCL8. This was confirmed on chemical reduction of these groups.
Similar content being viewed by others
Main
IL-8/CXC ligand 8 (CXCL8), the prototypal neutrophil-specific CXC chemokine in humans, is a key mediator of acute inflammation (1). In the developing intestine, elevated serum and tissue IL-8/CXCL8 concentrations are associated with inflammatory conditions such as necrotizing enterocolitis (2,3). It seems anomalous that the human fetal/neonatal gastrointestinal lumen is normally exposed to biologically significant IL-8/CXCL8 concentrations in swallowed amniotic fluid (4,5) and breast milk (6–9).
In an in vitro model of neonatal gastric and proximal intestinal conditions, IL-8/CXCL8 remains undigested, as tested by enzyme immunoassay (8). Recombinant human IL-8/CXCL8 promotes cellular migration, proliferation, and differentiation in intestinal epithelial cell (IEC) lines, besides protecting these cells from chemical injury (8). For assessing the physiologic relevance of these effects, the intact survival of IL-8/CXCL8 through gastric digestion needs confirmation in assays for structural and functional integrity. We hypothesized that IL-8/CXCL8 survives neonatal gastric digestion as a result of an intrinsic resistance to aspartyl proteinase activity. We treated recombinant human IL-8/CXCL8 (rhIL-8/CXCL8) with an established gastric fluid model (Simulated Gastric Fluid; USP, 10) (10) and measured IL-8/CXCL8 concentration and bioactivity in a variety of assays. IL-8/CXCL8-pepsin interaction was studied further in computational simulations involving molecular docking. The protease resistance of IL-8/CXCL8 was identified to be due to its disulfide bond-related tertiary folding and confirmed in actual chemical reduction experiments.
METHODS
Simulated gastric fluid (SGF) was prepared with 3.2 mg/mL of porcine pepsin (3200–4500 units/mg protein; Sigma Chemical Co., St. Louis, MO) in 0.2% NaCl/0.7% HCl (pH 1.2; US Pharmacopoeia) (10). Carrier-free rhIL-8/CXCL8 (R&D Systems, Minneapolis, MN) and BSA (control; Sigma Chemical Co.) were freshly reconstituted in PBS as 0.25-mg/mL solutions, which provided a pepsin:protein ratio of 50:1. Twenty microliters of the test/control protein solution was incubated with 80 μL of SGF at 37°C for 0, 5, 15, or 60 min. Reactions were terminated by addition of 35 μL of 0.32 M of sodium carbonate. The digestates from the four time points were tested for immunologic, structural, and functional integrity.
Immunologic analysis.
IL-8/CXCL8 was measured by the Human IL-8/CXCL8 Quantikine ELISA kit, second generation (R&D; sample required: 50 μL; range 10–2000 pg/mL). BSA was measured with the BSA immunoenzymetric assay (Cygnus Technologies, Southport, NC; sample 50 μL; range 0.25–32 ng/mL).
Peptide gel electrophoresis.
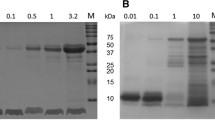
Peptide gel electrophoresis was performed on digestates from the SGF studies by a previously described standard method (11). rhIL-8/CXCL8 (0.5 μg) was used as positive control.
Western blot.
The digestate proteins (after electrophoresis) were transferred to a polyvinylidene difluoride membrane (BioRad). After blocking, the membrane was treated sequentially with monoclonal anti-human IL-8/CXCL8 IgG1 antibody (1 μg/mL; R&D), goat anti-mouse IgG-horseradish peroxidase (1:10,000; Upstate Biotechnology, Charlottesville, VA), and the ECL-plus staining reagents (Amersham Biosciences, Piscataway, NJ).
Image analysis.
The ELISA results from SGF digestates were validated by digital image densitometric analysis of the electrophoretic separations. We used the software application Scion Image version 4.0.2 (ported from the NIH Image for the Macintosh by Scion Corporation, Frederick, MD, and available on the Internet at http://www.scioncorp.com) (12). The band grayscale values were converted to percentages for comparison.
FHs-74-Int cell culture.
The FHs-74-Int cell line (ATCC, Rockville, MD) was used to evaluate the effect of IL-8/CXCL8 digestates on cell proliferation. This epithelial cell line is derived from a 12- to 16-wk fetus, and we have previously shown that rhIL-8/CXCL8 increases cell proliferation in these cells (8). These cells were cultured (passage numbers 13–15) as described earlier (8).
Cell proliferation experiments.
Cells were plated in 96-well plates at a density of 1–2 × 104/well. After 24 h, media were changed to phenol-red free media that contained the SGF digestates diluted to estimated (per ELISA results) 10-nM concentrations and incubated for 3 d. These concentrations were chosen on the basis of earlier observations (8). Negative controls were run with PBS and with SGF + PBS + stop buffer in the volumes used for the SGF treatment studies, whereas rhIL-8/CXCL8 (R&D) was used as positive control. Each set was run in triplicate. The cells were then labeled with Bromodeoxyuridine for 24 h, and its incorporation was measured by ELISA (Roche Diagnostics, Indianapolis, IN) as described previously (8).
Peripheral blood neutrophil chemotaxis assay.
Peripheral venous blood was obtained from healthy adult volunteers (approved by the institutional review board). Neutrophils were isolated to 90–95% purity by sequential dextran sedimentation (Sigma Chemical Co.), erythrocyte lysis (PharMLyse; BD Biosciences, San Diego, CA), and differential centrifugation by Ficoll-Hypaque density gradient (Sigma Chemical Co.). These cells were resuspended in 5 mL of RPMI 1640 without Phenol red (Sigma Chemical Co.) that contained 10% FCS (HyClone, Logan, UT) and labeled with calcein AM (5 μg/mL; Molecular Probes, Eugene, OR). Chemotaxis was tested by an automated assay (13) using disposable 96-well chemotaxis chambers (ChemoTx, Neuro Probe). The wells were filled with 29 μL of the SGF digestates diluted to an estimated (per ELISA results) rhIL-8/CXCL8 concentration of 1 nM (based on pilot data). Negative and positive controls were run as in cell proliferation studies. The wells were covered with an 8.0-μm pore size polycarbonate filter, and 25 μL of the labeled neutrophil suspension (3 × 106 cells/mL) was placed on the filter sites. The chamber was incubated at 37°C, 5% CO2 for 60 min. The plates were read spectrophotometrically at 485/530 nm (Fusion Universal Microplate Analyzer; Perkin Elmer, Shelton, CT), and neutrophil migration was assessed as the total fluorescence for each well. Each sample was run in duplicate.
Molecular docking.
The protein data bank (PDB) structures for IL-8/CXCL8 (PDB ID: 1IKL) (14) and porcine pepsin (PDB ID: 3PEP) (15) were obtained from the Research Collaboratory for Structural Bioinformatics (http://www.rcsb.org/pdb) (16). The interaction of IL-8/CXCL8 with porcine pepsin was studied by computational docking using MacroModel/BatchMin version 7.0 (Schrödinger, Inc., Portland, OR) (17). The docking procedure involved rigid body translation/rotation (using the BatchMin MOLS command) followed by energy minimization using the Truncated Newton Conjugate Gradient procedure (18). Porcine pepsin atoms were fixed at crystallographic positions, whereas the IL-8/CXCL8 structure was allowed to translate and rotate during the docking and energy minimization procedures. Modeling was performed both in vacuo and with generalized born/surface area solvation (19). Energy minimization protocols were based on the AMBER* force field distance-dependent electrostatics (20), which were further attenuated by a factor of 4 (“4r” electrostatics). The energy cutoff for the saved docked structures was set at 50 kJ/mol.
On the basis of the data from Powers et al. (21) on pepsin specificity, we identified amino acid residues and peptide bonds in IL-8/CXCL8 that might have relatively higher probabilities of pepsin-mediated cleavage. The amino acid targets, in decreasing order of susceptibility, were phenylalanine (positions 17, 21, ad 65; cleavage probability, p = 0.51), leucine (5, 25, 43, 49, 51, and 67; p = 0.41), tryptophan (57; p = 0.40), aspartate (45 and 52; p = 0.26), glutamate (4, 24, 29, 38, 48, 55, 63, and 70; p = 0.24), tyrosine (13; p = 0.24), and alanine (35 and 69; p = 0.16). These 23 amino acid residues are subsequently referred to as the “pepsin target residues.” In addition, the peptide bonds between the residues 17phenylalanine-18histidine (cleavage probability, p > 0.8), 21phenylalanine-22isoleucine (p < 0.6), and 65phenylalanine-66leucine (p < 0.6) were marked. The position of these residues was studied vis-à-vis those of pepsin active site aspartates at positions 32 and 215, and the S1 active site subunit residues (30isoleucine, 75tyrosine, 77threonine, 111phenylalanine, 112leucine, 117phenylalanine, 120isoleucine, and 217glycine) (22).
The docking procedure was started after a manual attempt to position the N-terminal loop phenylalanine residues (which had the highest theoretical probability of cleavage by pepsin) at a distance <25 Å and was performed sequentially both in vacuo and with solvation. The pepsin active site shell for positioning the N-terminal loop of IL-8/CXCL-8 was based on the data from Sielecki et al. (22).
Flexibility predictions for IL-8/CXCL8.
The flexible and rigid segments of IL-8/CXCL8 were predicted by using the software algorithm FIRST (Floppy Inclusion and Rigid Substructure Topography) (23). IL-8/CXCL8 structure was optimized for calculation of hydrogen bonds by adding atomic hydrogens to the PDB structure by using the application WHATIF (24).
Computational simulation of removal of disulfide bonds in IL-8/CXCL8.
The cysteine residues at positions 7 and 9 in the PDB molecular structure of IL-8/CXCL8 were mutated to alanine by using the Swiss-pdb deep viewer software application (25). The “mutated” molecule was studied further by introducing torsion at the α-carbon φ-angles of the identified 29glutamic acid “hinge” residue. This torsional movement produced multiple hypothetical geometries, which were then subjected to global energy minimization (in vacuo) by the GROMOS 43B1 force field (26). Relative solvent accessibility maps were computed for these refined hypothetical structures, wherein maximum accessibility was defined as the accessible surface of an amino acid “X” in a glycine-glycine-X-glycine-glycine pentapeptide in extended conformation. This scale provides a measure that is approximate but sufficiently accurate to differentiate core amino acids from surface ones. The mean relative solvent accessibilities for the “pepsin target residues” (vide supra) were compared statistically. The product of relative accessibility of each of these amino acids with its individual cleavage probability (21) was defined as “accessibility index” and also used for comparisons.
IL-8/CXCL8 was reduced by treatment with Immobilized Tris
[2-carboxyethyl] phosphine hydrochloride (TCEP) Disulfide Reducing Gel slurry (Pierce Biotechnology, Rockford, IL), wherein TCEP is covalently conjugated with 4% cross-lined beaded agarose support (27). Equal volumes of TCEP and 0.25 mg/mL of carrier-free rhIL-8/CXCL8 (R&D) were incubated for 30 min (based on pilot data), and the reduced protein was finally recovered by centrifugation at 1000 rpm again for 1 min.
As native IL-8/CXCL8 is highly stable up to 80°C (28), the procedure was repeated at 75°C to increase the efficiency of disulfide reduction without chemical denaturation. The structural integrity of the reduced protein was re-confirmed by gel electrophoresis (vide supra).
The extent of disulfide bond reduction in rhIL-8/CXCL8 was measured as the free sulfhydryl concentration in solution (Ellman's method) (29). Ellman's reagent solution was prepared by dissolving 4 mg of 5,5′-Dithio-bis-(2-nitrobenzoic acid) (Pierce) in 1 mL of reaction buffer [0.1 M of sodium phosphate (pH 8.0) and 1 mM of EDTA). Standards were prepared with cysteine hydrochloride monohydrate (0–1.5 mM). The reaction was run by adding 250 μL of test/standard solution to 50 μL of Ellman's reagent and 2.5 mL of reaction buffer for 15 min and measured as absorbance at 412 nm.
The reduced rhIL-8/CXCL8 was subjected to the SGF experiments (vide supra) to determine the contribution of intramolecular disulfide bonds to pepsin resistance. BSA was run as control. IL-8/CXCL8 and BSA concentrations were measured at different time points by ELISA.
Statistical methods.
IL-8/CXCL8 concentrations were described as mean ± SD. Continuous, independently measured data points were identified as parametric or nonparametric by testing for normality (Kolmogorov-Smirnov/Shapiro-Wilk tests) and equivalence of variance (Levene's test). Results of the digestion studies were depicted as proportion (percentage) of protein detectable at different time points. Digestion time-point comparisons were performed by a one-tailed two-factor mixed factorial ANOVA. Sphericity was tested by Mauchly's test, and the Greenhouse-Geisser test was used for analysis when homogeneity of covariance could not be demonstrated. Bivariate correlations were sought by the Pearson method. For all statistical procedures, an α error up to 0.05 was accepted.
RESULTS
Immunologic Analyses
The IL-8/CXCL8 concentrations detectable by ELISA at 0, 5, 15, and 60 min of treatment with SGF were 100, 63.30 ± 6.02, 13.66 ± 4.52, and 5.61 ± 2.88%, respectively. In contrast, BSA was undetectable at 5 min onward. These two patterns were significantly different (p = 0.013; (Fig. 1).
IL-8/CXCL8 digestion with simulated gastric fluid. Results are depicted as means ± SEM in a clustered line diagram. SGF was prepared with 3.2 mg/mL of porcine pepsin in 0.2% NaCl/0.7% HCl (pH 1.2). The figure shows concentrations of BSA (control; thick dotted line), rhIL-8/CXCL8 concentrations measured by ELISA (solid line), and IL-8/CXCL8 band intensities (image analysis) on gel electrophoresis of the digestates (thin dotted line). Each experiment (four replicates) was performed in two runs, and all ELISA measurements were in duplicate; n = 4 for statistical analyses; *p < 0.05; **p < 0.01; ***p < 0.001.
Structural Analyses
Peptide gel electrophoresis/Western blot.
The gel is shown in (Figure 2A. The band for rhIL-8/CXCL8 could be identified at 60 min of SGF treatment, in contrast to BSA that became nondetectable at 5 min (data not shown). This persistent band was recognized by monoclonal anti-human IL-8/CXCL8 antibody in Western blot (Fig. 2B).
Image analysis.
The SGF digestates at 0, 5, 15, and 60 min of treatment showed IL-8/CXCL8 electrophoretic bands with grayscale intensities of 100, 73.75 ± 7.25, 55.94 ± 4.76, and 36.75 ± 3.74%, respectively. These values correlated significantly with the values measured by ELISA (Pearson's r = 0.97, p < 0.001; (Fig. 1).
Functional Analyses
Cell proliferation experiments.
The SGF digestates at 0, 5, 15, and 60 min of treatment, when added in dilutions corresponding to 100-nM concentration, increased IEC proliferation as measured in a Bromodeoxyuridine cell ELISA. The optical densities for control (medium alone) were 1.54 ± 0.06, and with addition of SGF digestates at 0, 5, 15, and 60 min increased to 1.83 ± 0.04 (p = 0.001), 1.73 ± 0.05 (p = 0.03), 1.82 ± 0.03 (p = 0.001), and 1.86 ± 0.17 (p < 0.001). These results were similar to positive control (100 nM of rhIL-8/CXCL8; OD = 1.85 ± 0.05; p = 0.001). This effect was neutralized when the 60-min SGF digestate was added along with anti-human IL-8/CXCL8 antibody (1.51 ± 0.03; p = 0.96). The cell proliferation ELISA was also run with PBS (1.58 ± 0.10; p = 0.80) and SGF (1.59 ± 0.10; p = 0.71) controls, which were not statistically different from the culture medium alone (Fig. 3).
Bar diagram showing the effect of SGF-digested IL-8/CXCL8 on human fetal intestinal cell proliferation, indicating functional integrity of “digested” IL-8/CXCL8. Cellular proliferation was estimated by a cell ELISA for Bromodeoxyuridine incorporation. The diagram depicts mean ± SEM optical densities. The experiment (four replicates) included two runs, and each sample was tested in duplicate; n = 4 for statistical analyses; *p < 0.05; **p < 0.01; ***p < 0.001.
Neutrophil chemotaxis assay.
The digested rhIL-8/CXCL8 samples increased neutrophil chemotaxis at 0, 5, 15, and 60 min of treatment with SGF when compared with PBS and SGF controls (p < 0.001 for each time point). These results were not significantly different between the individual time points (p = 0.13–0.31) or from positive control (1 nM of rhIL-8/CXCL8; p = 0.06–0.7). SGF did not increase neutrophil chemotaxis when compared with PBS (p = 0.65; (Fig. 4).
Bar diagram showing the effect of SGF-digested IL-8/CXCL8 on neutrophil chemotaxis. Neutrophil chemotaxis was measured by an automated assay measuring movement of calcein-labeled neutrophils across 8-μm polycarbonate filters. The diagram depicts mean ± SEM fluorescence intensities. The experiment (four replicates) included two runs, and each sample was tested in duplicate; n = 4 for statistical analyses; *p < 0.05; **p < 0.01; ***p < 0.001.
Computational Simulations
Molecular docking.
The in vacuo docking procedure yielded only one docked geometry after 5000 steps. The potential target residues for enzymatic cleavage were separated from the pepsin active site aspartates by >25 Å, indicating a poor enzyme-substrate fit. Similarly, on repeating the procedure using the generalized born/surface area solvation protocol, only a single structure within the predefined energy window of 50 kJ/mol was obtained repeatedly after 1800 steps. The docked geometry continued to show a poor fit; the distance of the high cleavage probability amino acid residues actually increased to ≥35 Å, with the N-terminal loop now facing away from the enzyme active site (Fig. 5).
Lowest energy dock structure between porcine pepsin and IL-8/CXCL8. Rigid body docking was performed with the software application MACROMODEL version 7.0 with generalized born/surface area solvation. The above structure showed a poor enzyme-substrate fit, with all potential target residues for peptic cleavage separated from the active site residues in pepsin by >25 Å.
Flexibility predictions for IL-8/CXCL8.
The software algorithm FIRST identified a highly flexible hinge in IL-8/CXCL8 at the 29glutamate position (Fig. 6).
Computational simulation of removal of disulfide bonds in IL-8/CXCL8. The molecular structure of IL-8/CXCL8 was studied after cysteine→alanine “mutations” at positions 7 and 9. Multiple hypothetical versions of the mutated IL-8/CXCL8 structure were obtained with serial increments of torsional strain at 29glutamate (Fig. 7).
(A) Native IL-8/CXCL8. (B) One of the hypothetical versions of IL-8/CXCL8 structure following computational substitution of cysteine (residues 7 and 9), followed by simulated application of torsional strain at a high flexibility hinge at 29glutamate position. The above computational simulations were performed by using the software application Swiss PDB viewer and its associated algorithms.
This torsional strain led to an opening of IL-8/CXCL8 structure and increased the solvent accessibility of the N-loop amino acid residues. The mean relative accessibility of the “pepsin target residues” increased significantly from 29.26 ± 17.38% in native IL-8/CXCL8 structure to 37.36 ± 14.54% (p = 0.002) in the hypothetical “mutated” structure shown in (Figure 7. When the accessibility indices (relative accessibility × cleavage probability) for each of these 23 amino acids were compared, the difference again was significantly different (10.04 ± 8.02 versus 12.59 ± 7.94%, respectively; p = 0.001).
Chemical Reduction of Intramolecular Disulfide Bonds in IL-8/CXCL8
The degree of disulfide bond reduction was measured as the concentration of free sulfhydryl groups in solution. The free sulfhydryl concentrations in native rhIL-8/CXCL8, after reduction with Immobilized TCEP at room temperature and after reduction with immobilized TCEP at 75°C, were equivalent to 0, 0.202, and 0.301 mM of cysteine.
rhIL-8/CXCL8 migrated as a single electrophoretic band similar to native rhIL-8/CXCL8 after treatment with TCEP at room temperature. The results were similar when the reduction was performed at 75°C.
After immobilized TCEP reduction at room temperature, IL-8/CXCL8 concentrations at 0, 5, 15, and 60 min of treatment with SGF were 100%, 4.17 ± 0.72%, 2.01 ± 0.34%, and nondetectable, respectively (p = 0.002). When IL-8/CXCL8 was reduced at 75°C, the concentrations were 100%, 0.06 ± 0.04%, 0.03 ± 0.01%, and nondetectable, respectively (p = 0.002). These results are shown in (Figure 8.
Effect of reduction of intramolecular disulfide bonds on IL-8/CXCL8 resistance to digestion with SGF. IL-8 concentrations are depicted as the detectable proportion (%) of the initial concentration. Disulfide bonds were reduced by chemical treatment with TCEP immobilized by covalent conjugation to 4% cross-lined beaded agarose support. The reactions were carried out at room temperature and at 75°C. Reduction was confirmed by measuring free sulfhydryl concentrations by Ellman's method. The experiment (four replicates) was run twice, and all ELISA measurements were in duplicate; n = 4 for statistical analyses; *p < 0.05; **p < 0.01; ***p < 0.001.
DISCUSSION
Several human milk-borne cytokines, including erythropoietin, granulocyte-colony stimulating factor, and epidermal growth factor, survive neonatal gastric digestion (30–33). Factors that have been postulated to explain this phenomenon include antiproteases in milk, relative lack of aspartyl proteinase activity in the neonatal stomach as a result of hypochlorhydria or paucity of the enzyme, and the protective effect of binding to cognate receptors in the gastrointestinal mucosa (30–33). However, several lines of evidence now suggest that aspartyl proteinase activity might actually be detectable in the fetal stomach from midgestation, thereby suggesting that additional factors might be at work in preservation of activity of these cytokines (34). It also remains to be seen whether these cytokines would retain bioactivity even after gastric acid and pepsin production is well established during infancy, as peptides such as epidermal growth factor seem to be digested into smaller, less active forms in acidic gastric secretions (34–36). In this study, we examined this issue in relation to IL-8/CXCL8.
Milk-borne IL-8/CXCL8 is highly resistant to simulated neonatal gastric digestion. Our previous demonstration of high concentrations in neonatal gastric contents provides further indirect evidence of its stability (37). Because the gastric contents were collected from neonates who had never received breast milk feeding, it is clear that human milk-borne antiproteases are not essential for the intact survival of IL-8/CXCL8. Similarly, the presence of high concentrations of IL-8/CXCL8 in the gastric luminal contents excludes a major protective role of binding to the mucosal receptors.
We also reported previously that IL-8/CXCL8 concentrations were lower in term gastric contents than in those of preterm neonates (37), which, arguably, could be due to partial digestion by higher proteolytic activity of the term neonatal stomach (38,39). However, in vitro, IL-8/CXCL8 survives simulated digestions performed with gastric aspirates obtained from term as well as preterm neonates (6–9). Our gastric aspirates were collected after 24 h of life; therefore, it is unlikely that amniotic fluid present from birth would be a major constituent of these samples. It is more likely that the origin of this IL-8/CXCL8 is from mucosal expression (37). This expression decreases with maturation and can provide an explanation for gestational differences in gastric content IL-8/CXCL8 levels.
Gastric proteolytic activity is primarily due to the presence of aspartyl proteinase enzymes. These proteins are remarkably conserved through evolution and fetal development, as is evident from the work of Kageyama (34), and also from information available at the Conserved domain database, National Center for Biotechnology Information (http://www.ncbi.nlm.nih.gov/Structure/cdd/cddsrv.cgi?uid = pfam00026). The human fetus and neonate express several of these enzymes, including prochymosin, pepsinogens, and progastricsin, in a developmentally regulated sequence (34,40). The enzymes expressed earlier during development have higher pH optima than Pepsin A, which makes those better suited to the relatively hypochlorhydric environment of the fetus and the young infant (34,40). However, their tertiary structure, catalytic activity, and substrate specificity are still markedly similar to each other and to other known aspartyl proteinases (34,40). To study further the interaction of IL-8/CXCL8 with these enzymes, we chose an established model based on porcine pepsin, which has been used extensively to study protein stability and oral bioavailability of drugs (10,41). The porcine enzyme, besides being the best studied member of this family, is one of the closest structural homologs of the human pepsin A: a pairwise BLAST (Basic Local Alignment Search Tool; http://www.ncbi.nlm.nih.gov/blast/) analysis (15,42,43) shows a high degree of sequence alignment; a bit score of 587 (1514), 85% identities, 95% positives, no gaps, and an expectation score of e-166 against finding a better possible match, whereas flexible three-dimensional structural alignment using computational tools (44) gives root mean squared distances as low as 0.94 Å. If only the active site residues are aligned, then sequences are 95% identical and the root mean squared distance is only 0.69 Å, thus further confirming the similarity. Proteins that are considered “resistant” to gastric proteolysis are often stable in SGF for 5–60 min, in contrast to others, such as serum albumin, that are digested within seconds (45). The choice of a model with acidic pH and excess proteinase activity is also relevant as it helps in demonstrating that IL-8/CXCL8 is likely to retain its potential biologic significance during infancy as gastric acid and pepsin output increase (37,44).
In the present study, rhIL-8/CXCL8 retained immunologic (by ELISA and Western blot), structural (in gel electrophoresis), and functional (enterocyte proliferation and neutrophil chemotaxis) integrity even at 60 min of SGF treatment. The protease resistance can be explained on the basis of the poor docking fit observed in the computational simulation. The low predicted solvent accessibility of the target aromatic/hydrophobic amino acids in native IL-8/CXCL8 also suggests that the protease resistance of IL-8/CXCL8 is primarily due to its tertiary structure. Dimerization of IL-8/CXCL8 occurs mainly at concentrations higher than those seen in vivo or in our experimental conditions and therefore was not considered to be a major factor in protease resistance (46). IL-8 is also not believed to be significantly glycosylated, and its sequence lacks the tripeptide asparagine-arginine-threonine that is usually associated with N-glycosylation of asparagine (47). Therefore, we did not consider glycosylation as a likely factor in its resistance to aspartyl proteinases.
IL-8/CXCL8 is a 72–amino acid residue protein with two disulfide bridges, from Cys-7 to Cys-34 and from Cys-9 to Cys-50 (48). These disulfide bridges link the functionally important but mobile N-terminal region to the core of the protein and therefore form a critical part of the protein scaffold. Both disulfide bridges are essential for maintenance of the native tertiary fold and stability (49). In view of the poor docking fit between IL-8/CXCL8 and the porcine pepsin active site, we hypothesized that the tertiary folding and, hence, indirectly, the disulfide bridges in the IL-8/CXCL8 are central to this protease resistance.
Simulated removal of these disulfide bonds by computational “mutation” of the cysteine (to alanine) residues allowed, as expected (28), increased freedom of movement for the N-terminal residues. This also allowed torsional movement at the 29glutamate residue, which had been identified as a flexible “hinge” region on flexibility analysis. In these computational simulations, application of torsional strain at this site opens up the IL-8/CXCL8 structure, exposing the aromatic/hydrophobic residues and allowing for an improved docking fit. These predictions are very close to the experimental results reported previously by Clark-Lewis et al. (49). In their study, substitution of the cysteine residues with α-aminobutyric acid led to the loss of the compact molecular structure, with displacement and increased mobility of the 29– to 37–amino acid region.
Chemical reduction of the disulfide bonds in IL-8/CXCL8 confirmed our hypothesis that the resistance of IL-8/CXCL8 to gastric proteolysis is due to its disulfide structure. The validation of these computational tools suggests that this may be an accurate method to identify various other bioactive factors in amniotic fluid and human milk that survive digestion. The interaction of various bioactive molecules with the developing gastric and pancreatic enzyme systems is a highly relevant issue in the study of intestinal development (50). There is evidence from animal models that certain agents not only may interact with the gastrointestinal tract but also may be absorbed into the systemic circulation of the recipient (51). In human neonates, whereas some ingested peptides such as epidermal growth factor are absorbed systemically, other molecules, including erythropoietin and granulocyte-colony stimulating factor, are not (52–54). Further work is needed in this area, as improved understanding of the fate of various luminal peptides and other bioactive factors is likely to be of immense importance in neonatal nutrition.
Abbreviations
- CXCL8:
-
CXC ligand 8
- FIRST:
-
Floppy Inclusion and Rigid Substructure Topography
- IEC:
-
intestinal epithelial cell
- PDB:
-
protein data bank
- rhIL-8/CXCL8:
-
recombinant human IL-8/CXCL8
- SGF:
-
simulated gastric fluid
- TCEP:
-
Tris [2-carboxyethyl] phosphine hydrochloride
References
Baggiolini M, Clark-Lewis I 1992 Interleukin-8, a chemotactic and inflammatory cytokine. FEBS Lett 307: 97–101
Edelson MB, Bagwell CE, Rozycki HJ 1999 Circulating pro- and counterinflammatory cytokine levels and severity in necrotizing enterocolitis. Pediatrics 103: 766–771
Viscardi RM, Lyon NH, Sun CC, Hebel JR, Hasday JD 1997 Inflammatory cytokine mRNAs in surgical specimens of necrotizing enterocolitis and normal newborn intestine. Pediatr Pathol Lab Med 17: 547–559
Denison FC, Riley SC, Wathen NC, Chard T, Calder AA, Kelly RW 1998 Differential concentrations of monocyte chemotactic protein-1 and interleukin-8 within the fluid compartments present during the first trimester of pregnancy. Hum Reprod 13: 2292–2295
Kemp B, Winkler M, Maas A, Maul H, Ruck P, Reineke T, Rath W 2002 Cytokine concentrations in the amniotic fluid during parturition at term: correlation to lower uterine segment values and to labor. Acta Obstet Gynecol Scand 81: 938–942
Srivastava MD, Srivastava A, Brouhard B, Saneto R, Groh-Wargo S, Kubit J 1996 Cytokines in human milk. Res Commun Mol Pathol Pharmacol 93: 263–287
Michie CA, Tantscher E, Schall T, Rot A 1998 Physiological secretion of chemokines in human breast milk. Eur Cytokine Netw 9: 123–129
Maheshwari A, Lu W, Lacson A, Barleycorn AA, Nolan S, Christensen RD, Calhoun DA 2002 Effects of interleukin-8 on the developing human intestine. Cytokine 20: 256–267
Maheshwari A, Christensen RD, Calhoun DA 2003 ELR+ CXC chemokines in human milk. Cytokine 24: 91–102
Simulated Gastric Fluid 2000 The National Formulary 18. In: The National Formulary 18. Board of Trustees, United States Pharmacopeial Convention, Rockville, MD
Schagger H, Aquila H, Von Jagow G 1988 Coomassie blue-sodium dodecyl sulfate-polyacrylamide gel electrophoresis for direct visualization of polypeptides during electrophoresis. Anal Biochem 173: 201–205
O'Neill RR, Mitchell LG, Merril CR, Rasband WS 1989 Use of image analysis to quantitate changes in form of mitochondrial DNA after x-irradiation. Appl Theor Electrophor 1: 163–167
Frevert CW, Wong VA, Goodman RB, Goodwin R, Martin TR 1998 Rapid fluorescence-based measurement of neutrophil migration in vitro. J Immunol Methods 213: 41–52
Rajarathnam K, Clark-Lewis I, Sykes BD 1995 1H NMR solution structure of an active monomeric interleukin-8. Biochemistry 34: 12983–12990
Abad-Zapatero C, Rydel TJ, Erickson J 1990 Revised 2.3 angstroms structure of porcine pepsin. Evidence for a flexible subdomain. Proteins 8: 62–81
Berman HM, Westbrook J, Feng Z, Gilliland G, Bhat TN, Weissig H, Shindyalov IN, Bourne PE 2000 The Protein Data Bank. Nucleic Acids Res 28: 235–242
Mohamadi F, Richards NGJ, Guida WC, Liskamp R, Lipton M, Caufield C, Chang G, Hendrickson T, Still WC 1990 Macromodel—an integrated software system for modeling organic and bioorganic molecules using molecular mechanics. J Comput Chem 11: 440–467
Ponder JW, Richards FM 1987 An efficient newton-like method for molecular mechanics energy minimization of large molecules. J Comput Chem 8: 1016–1026
Hasel W, Hendrickson TF, Still WC 1988 A rapid approximation to the solvent accessible surface areas of atoms. Tetrahedron Comput Method 1: 103–108
McDonald DQ, Still WC 1992 AMBER* torsional parameters for the peptide backbone. Tetrahedron Lett 33: 7743–7748
Powers JC, Harley AD, Myers DV 1977 Subsite specificity of porcine pepsin. Adv Exp Med Biol 95: 141–157
Sielecki AR, Fedorov AA, Boodhoo A, Andreeva NS, James MN 1990 Molecular and crystal structures of monoclinic porcine pepsin refined at 1.8 A resolution. J Mol Biol 214: 143–170
Jacobs DJ, Rader AJ, Kuhn LA, Thorpe MF 2001 Protein flexibility predictions using graph theory. Proteins 44: 150–165
Vriend G 1990 WHAT IF: a molecular modeling and drug design program. J Mol Graph 8: 52–56
Guex N, Peitsch MC 1997 SWISS-MODEL and the Swiss-PdbViewer: an environment for comparative protein modeling. Electrophoresis 18: 2714–2723
van Gunsteren WF, Hünenberger PH, Mark AE, Smith PE, Tironi IG 1995 Computer simulation of protein motion. Computer Phys Commun 91: 305–319
Han JC, Han GY 1994 A procedure for quantitative determination of tris(2-carboxyethyl)phosphine, an odorless reducing agent more stable and effective than dithiothreitol. Anal Biochem 220: 5–10
Rajarathnam K, Sykes BD, Dewald B, Baggiolini M, Clark-Lewis I 1999 Disulfide bridges in interleukin-8 probed using non-natural disulfide analogues: dissociation of roles in structure from function. Biochemistry 38: 7653–7658
Riddles PW, Blakeley RL, Zerner B 1983 Reassessment of Ellman's reagent. Methods Enzymol 91: 49–60
Juul SE, Zhao Y, Dame JB, Du Y, Hutson AD, Christensen RD 2000 Origin and fate of erythropoietin in human milk. Pediatr Res 48: 660–667
Calhoun DA, Lunøe M, Du Y, Staba SL, Christensen RD 1999 Concentrations of granulocyte colony-stimulating factor in human milk after in vitro simulations of digestion. Pediatr Res 46: 767–771
Britton JR, George-Nascimento C, Udall JN, Koldovsky O 1989 Minimal hydrolysis of epidermal growth factor by gastric fluid of preterm infants. Gut 30: 327–332
Kling PJ, Sullivan TM, Roberts RA, Philipps AF, Koldovsky O 1998 Human milk as a potential enteral source of erythropoietin. Pediatr Res 43: 216–221
Kageyama T 2002 Pepsinogens, progastricsins, and prochymosins: structure, function, evolution, and development. Cell Mol Life Sci 59: 288–306
Boyle JT 2003 Acid secretion from birth to adulthood. J Pediatr Gastroenterol Nutr 37: S12–S16
Playford RJ, Marchbank T, Calnan DP, Calam J, Royston P, Batten JJ, Hansen HF 1995 Epidermal growth factor is digested to smaller, less active forms in acidic gastric juice. Gastroenterology 108: 92–101
Maheshwari A, Lacson A, Lu W, Fox SE, Barleycorn AA, Christensen RD, Calhoun DA 2004 Interleukin-8/CXCL8 forms an autocrine loop in fetal intestinal mucosa. Pediatr Res 56: 240–249
Henderson TR, Hamosh M, Armand M, Mehta NR, Hamosh P 2001 Gastric proteolysis in preterm infants fed mother's milk or formula. Adv Exp Med Biol 501: 403–408
Koldovsky O 1994 Hormonally active peptides in human milk. Acta Paediatr Suppl 402: 89–93
Dunn BM, Hung S 2000 The two sides of enzyme-substrate specificity: lessons from the aspartic proteinases. Biochim Biophys Acta 1477: 231–240
Dressman JB, Reppas C 2000 In vitro-in vivo correlations for lipophilic, poorly water-soluble drugs. Eur J Pharm Sci 11: S73–S80
Altschul SF, Madden TL, Schaffer AA, Zhang J, Zhang Z, Miller W, Lipman DJ 1997 Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res 25: 3389–3402
Fujinaga M, Chernaia MM, Tarasova NI, Mosimann SC, James MN 1995 Crystal structure of human pepsin and its complex with pepstatin. Protein Sci 4: 960–972
Shatsky M, Fligelman ZY, Nussinov R, Wolfson HJ 2000 Alignment of flexible protein structures. Proc Int Conf Intell Syst Mol Biol 8: 329–343
Astwood JD, Leach JN, Fuchs RL 1996 Stability of food allergens to digestion in vitro. Nat Biotechnol 14: 1269–1273
Burrows SD, Doyle ML, Murphy KP, Franklin SG, White JR, Brooks I, McNulty DE, Scott MO, Knutson JR, Porter D et al. 1994 Determination of the monomer-dimer equilibrium of interleukin-8 reveals it is a monomer at physiological concentrations. Biochemistry 33: 12741–12745
Thim L, Norris K, Norris F, Nielsen PF, Bjorn SE, Christensen M, Petersen J 1993 Purification and characterization of the trefoil peptide human spasmolytic polypeptide (hSP) produced in yeast. FEBS Lett 318: 345–352
Clore GM, Gronenborn AM 1991 Comparison of the solution nuclear magnetic resonance and crystal structures of interleukin-8. Possible implications for the mechanism of receptor binding. J Mol Biol 217: 611–620
Clark-Lewis I, Dewald B, Loetscher M, Moser B, Baggiolini M 1994 Structural requirements for interleukin-8 function identified by design of analogs and CXC chemokine hybrids. J Biol Chem 269: 16075–16081
Goldman AS 2000 Modulation of the gastrointestinal tract of infants by human milk. Interfaces and interactions. An evolutionary perspective. J Nutr 130: 426S–431S
Koldovsky O, Goldman AS 1999 Koldovsky O, Goldman AS 1999 Growth factors and cytokines in milk. In: Ogra PL, Mestecky J, Lamm ME, Strober W, Bienenstock J, McGhee JR (eds) Mucosal Immunology. Academic Press, San Diego pp 1523–1530
Gale SM, Read LC, George-Nascimento C, Wallace JC, Ballard FJ 1989 Is dietary epidermal growth factor absorbed by premature human infants?. Biol Neonate 55: 104–110
Juul SE, Christensen RD 2003 Absorption of enteral recombinant human erythropoietin by neonates. Ann Pharmacother 37: 782–786
Calhoun DA, Maheshwari A, Christensen RD 2003 Recombinant granulocyte colony-stimulating factor administered enterally to neonates is not absorbed. Pediatrics 112: 421–423
Acknowledgements
We thank Drs. D.J. Jacobs, Leslie A. Kuhn, and M.F. Thorpe (Michigan State University, East Lansing, MI) for permission to use the algorithm FIRST, and Dr. Gert Vriend (University of Groningen, The Netherlands) for WHAT IF. We are grateful to Nicolas Guex, Alexandre Diemand, Manuel C. Peitsch, Torsten Schwede, and GlaxoSmithKline R&D for allowing access to the Swiss deep view pdb viewer (http://www.expasy.org/spdbv) and to the Nussinov-Wolfson Structural Bioinformatics Group (Tel-Aviv University, Israel) for allowing access to the FlexProt protein alignment algorithm. We thank Drs. Deepak Agrawal (Moffitt Cancer Center, Tampa, FL) and Jane Carver for thoughtful discussions on the manuscript; Dawn Bruton, Sue Stripling, and Debbie Patrick, RN, for help with sample and data collection; and Ingrid Williams for managerial support.
Author information
Authors and Affiliations
Corresponding author
Additional information
This work was supported by grants HD-01180, HD-42326, and HL-61798 from the National Institutes of Health.
Rights and permissions
About this article
Cite this article
Maheshwari, A., Lu, W., Guida, W. et al. IL-8/CXC Ligand 8 Survives Neonatal Gastric Digestion as a Result of Intrinsic Aspartyl Proteinase Resistance. Pediatr Res 57, 438–444 (2005). https://doi.org/10.1203/01.PDR.0000151317.08180.7E
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1203/01.PDR.0000151317.08180.7E