Abstract
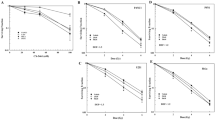
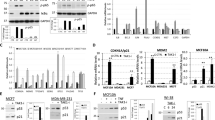
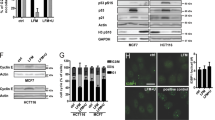
Many drugs currently used in chemotherapy work by hindering the process of ribosome biogenesis. In tumors with functional p53, the inhibition of ribosome biogenesis may contribute to the efficacy of this treatment by inducing p53 stabilization. As the level of stabilized p53 is critical for the induction of cytotoxic effects, it seems useful to highlight those cancer cell characteristics that can predict the degree of p53 stabilization following the treatment with inhibitors of ribosome biogenesis. In the present study we exposed a series of p53 wild-type human cancer cell lines to drugs such as actinomycin D (ActD), doxorubicin, 5-fluorouracil and CX-5461, which hinder ribosomal RNA (rRNA) synthesis. We found that the amount of stabilized p53 was directly related to the level of ribosome biogenesis in cells before the drug treatment. This was due to different levels of inactivation of the ribosomal proteins–MDM2 pathway of p53 digestion. Inhibition of rRNA synthesis always caused cell cycle arrest, independent of the ribosome biogenesis rate of the cells, whereas apoptosis occurred only in cells with a high rDNA transcription rate. The level of p53 stabilization induced by drugs acting in different ways from the inhibition of ribosome biogenesis, such as hydroxyurea (HU) and nutlin-3, was independent of the level of ribosome biogenesis in cells and always lower than that occurring after the inhibition of rRNA synthesis. Interestingly, in cells with a low ribosome biogenesis rate, the combined treatment with ActD and HU exerted an additive effect on p53 stabilization. These results indicated that (i) drugs inhibiting ribosome biogenesis may be highly effective in p53 wild-type cancers with a high ribosome biogenesis rate, as they induce apoptotic cell death, and (ii) the combination of drugs capable of stabilizing p53 through different mechanisms may be useful for treating cancers with a low ribosome biogenesis rate.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
References
Vogelstein B, Lane D, Levine AJ . Surfing the p53 network. Nature 2000; 408: 307–310.
Johnstone RW, Ruefli AA, Lowe SW . Apoptosis. Cell 2002; 108: 153–164.
Vousden KH, Prives C . Blinded by the light: the growing complexity of p53. Cell 2009; 137: 413–431.
Harris SL, Levine AJ . The p53 pathway: positive and negative feedback loops. Oncogene 2005; 24: 2899–2908.
Vousden KH, Lu X . Live or let die: the cell’s response to p53. Nat Rev Cancer 2002; 2: 594–604.
Mayer C, Grummt I . Cellular stress and nucleolar function. Cell Cycle 2005; 4: 1036–1038.
Zhang Y, Lu H . Signaling to p53: ribosomal proteins find their way. Cancer Cell 2009; 16: 369–377.
Deisenroth C, Zhang Y . Ribosome biogenesis surveillance: probing the ribosomal protein-Mdm2-p53 pathway. Oncogene 2010; 29: 4253–4260.
Hu W, Feng Z, Levine AJ . The regulation of multiple p53 stress responses is mediated through MDM2. Genes Cancer 2012; 3: 199–208.
Momand J, Zambetti GP, Olson DC, George D, Levine AJ . The mdm-2 oncogene product forms a complex with the p53 protein and inhibits p53-mediated transactivation. Cell 1992; 69: 1237–1245.
Haupt Y, Maya R, Kazaz A, Oren M . Mdm2 promotes the rapid degradation of p53. Nature 1997; 387: 296–299.
Kubbutat MH, Jones SN, Vousden KH . Regulation of p53 stability by Mdm2. Nature 1997; 387: 299–303.
Gajjar M, Candeias MM, Malbert-Colas L, Mazars A, Fujita J, Olivares-Illana V et al. The p53 mRNA-Mdm2 interaction controls Mdm2 nuclear trafficking and is required for p53 activation following DNA damage. Cancer Cell 2012; 21: 25–35.
Lowe SW . Activation of p53 by oncogenes. Endocr Relat Cancer 1999; 6: 45–48.
Zhang Y, Wolf GW, Bhat K, Jin A, Allio T, Burkhart WA et al. Ribosomal protein L11 negatively regulates oncoprotein MDM2 and mediates a p53-dependent ribosomal-stress checkpoint pathway. Mol Cell Biol 2003; 23: 8902–8912.
Sun X-X, Dai M-S, Lu H . 5-fluorouracil activation of p53 involves an MDM2-ribosomal protein interaction. J Biol Chem 2007; 282: 8052–8059.
Sun X-X, Dai M-S, Lu H . Mycophenolic acid activation of p53 requires ribosomal proteins L5 and L11. J Biol Chem 2008; 283: 12387–12392.
Yuan X, Zhou Y, Casanova E, Chai M, Kiss E, Gröne H-J et al. Genetic inactivation of the transcription factor TIF-IA leads to nucleolar disruption, cell cycle arrest, and p53-mediated apoptosis. Mol Cell 2005; 19: 77–87.
Donati G, Bertoni S, Brighenti E, Vici M, Treré D, Volarevic S et al. The balance between rRNA and ribosomal protein synthesis up- and downregulates the tumour suppressor p53 in mammalian cells. Oncogene 2011; 30: 3274–3288.
Lohrum MAE, Ludwig RL, Kubbutat MHG, Hanlon M, Vousden KH . Regulation of HDM2 activity by the ribosomal protein L11. Cancer Cell 2003; 3: 577–587.
Bhat KP, Itahana K, Jin A, Zhang Y . Essential role of ribosomal protein L11 in mediating growth inhibition-induced p53 activation. EMBO J 2004; 23: 2402–2412.
Dai M-S, Lu H . Inhibition of MDM2-mediated p53 ubiquitination and degradation by ribosomal protein L5. J Biol Chem 2004; 279: 44475–44482.
Dai M-S, Zeng SX, Jin Y, Sun X-X, David L, Lu H . Ribosomal protein L23 activates p53 by inhibiting MDM2 function in response to ribosomal perturbation but not to translation inhibition. Mol Cell Biol 2004; 24: 7654–7668.
Jin A, Itahana K, O’Keefe K, Zhang Y . Inhibition of HDM2 and activation of p53 by ribosomal protein L23. Mol Cell Biol 2004; 24: 7669–7680.
Burger K, Mühl B, Harasim T, Rohrmoser M, Malamoussi A, Orban M et al. Chemotherapeutic drugs inhibit ribosome biogenesis at various levels. J Biol Chem 2010; 285: 12416–12425.
Drygin D, Lin A, Bliesath J, Ho CB, O’Brien SE, Proffitt C et al. Targeting RNA polymerase I with an oral small molecule CX-5461 inhibits ribosomal RNA synthesis and solid tumor growth. Cancer Res 2011; 71: 1418–1430.
Bywater MJ, Poortinga G, Sanij E, Hein N, Peck A, Cullinane C et al. Inhibition of RNA polymerase I as a therapeutic strategy to promote cancer-specific activation of p53. Cancer Cell 2012; 22: 51–65.
Drygin D, Siddiqui-Jain A, O’Brien S, Schwaebe M, Lin A, Bliesath J et al. Anticancer activity of CX-3543: a direct inhibitor of rRNA biogenesis. Cancer Res 2009; 69: 7653–7661.
Drygin D, Rice WG, Grummt I . The RNA polymerase I transcription machinery: an emerging target for the treatment of cancer. Annu Rev Pharmacol Toxicol 2010; 50: 131–156.
Chen X, Ko LJ, Jayaraman L, Prives C . p53 levels, functional domains, and DNA damage determine the extent of the apoptotic response of tumor cells. Genes Dev 1996; 10: 2438–2451.
Perry RP, Kelley DE . Inhibition of RNA synthesis by actinomycin D: characteristic dose-response of different RNA species. J Cell Physiol 1970; 76: 127–139.
Derenzini M, Montanaro L, Chillà A, Tosti E, Vici M, Barbieri S et al. Key role of the achievement of an appropriate ribosomal RNA complement for G1-S phase transition in H4-II-E-C3 rat hepatoma cells. J Cell Physiol 2005; 202: 483–491.
Poortinga G, Quinn LM, Hannan RD . Targeting RNA polymerase I to treat MYC-driven cancer. Oncogene 2014 2015; 34: 403–412.
Burns TF, El-Deiry WS . The p53 pathway and apoptosis. J Cell Physiol 1999; 181: 231–239.
Sax JK, El-Deiry WS . p53-induced gene expression analysis. Methods Mol Biol 2003; 234: 65–71.
Xiong Y, Hannon GJ, Zhang H, Casso D, Kobayashi R, Beach D . p21 is a universal inhibitor of cyclin kinases. Nature 1993; 366: 701–704.
Sherr CJ, McCormick F . The RB and p53 pathways in cancer. Cancer Cell 2002; 2: 103–112.
Nakano K, Vousden KH . PUMA, a novel proapoptotic gene, is induced by p53. Mol Cell 2001; 7: 683–694.
Yu J, Zhang L . PUMA, a potent killer with or without p53. Oncogene 2008; 27: S71–S83.
Oltvai ZN, Milliman CL, Korsmeyer SJ . Bcl-2 heterodimerizes in vivo with a conserved homolog, Bax, that accelerates programmed cell death. Cell 1993; 74: 609–619.
Yin C, Knudson CM, Korsmeyer SJ, Van Dyke T . Bax suppresses tumorigenesis and stimulates apoptosis in vivo. Nature 1997; 385: 637–640.
Koopman G, Reutelingsperger CP, Kuijten GA, Keehnen RM, Pals ST, van Oers MH . Annexin V for flow cytometric detection of phosphatidylserine expression on B cells undergoing apoptosis. Blood 1994; 84: 1415–1420.
Kaufmann SH, Desnoyers S, Ottaviano Y, Davidson NE, Poirier GG . Specific proteolytic cleavage of poly(ADP-ribose) polymerase: an early marker of chemotherapy-induced apoptosis. Cancer Res 1993; 53: 3976–3985.
Porter AG, Jänicke RU . Emerging roles of caspase-3 in apoptosis. Cell Death Differ 1999; 6: 99–104.
Elledge SJ, Zhou Z, Allen JB . Ribonucleotide reductase: regulation, regulation, regulation. Trends Biochem Sci 1992; 17: 119–123.
Gottifredi V, Shieh S, Taya Y, Prives C . p53 accumulates but is functionally impaired when DNA synthesis is blocked. Proc Natl Acad Sci USA 2001; 98: 1036–1041.
Kapoor M, Hamm R, Yan W, Taya Y, Lozano G . Cooperative phosphorylation at multiple sites is required to activate p53 in response to UV radiation. Oncogene 2000; 19: 358–364.
Vassilev LT, Vu BT, Graves B, Carvajal D, Podlaski F, Filipovic Z et al. In vivo activation of the p53 pathway by small-molecule antagonists of MDM2. Science 2004; 303: 844–848.
Longley DB, Harkin DP, Johnston PG . 5-fluorouracil: mechanisms of action and clinical strategies. Nat Rev Cancer 2003; 3: 330–338.
Oskarsson T, Trumpp A . The Myc trilogy: lord of RNA polymerases. Nat Cell Biol 2005; 7: 215–217.
Maggi LB, Weber JD . Nucleolar adaptation in human cancer. Cancer Invest 2005; 23: 599–608.
Montanaro L, Treré D, Derenzini M . Nucleolus, ribosomes, and cancer. Am J Pathol 2008; 173: 301–310.
Derenzini M, Montanaro L, Treré D . What the nucleolus says to a tumour pathologist. Histopathology 2009; 54: 753–762.
Derenzini M . The AgNORs. Micron 2000; 31: 117–120.
Bland JM . The tyranny of power: is there a better way to calculate sample size? BMJ 2009; 339: b3985.
Acknowledgements
This work was supported by the Roberto and Cornelia Pallotti’s Legacy for Cancer Research and Associazione Italiana per la Ricerca sul Cancro (AIRC, grant number IG13480).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on the Oncogene website
Supplementary information
Rights and permissions
About this article
Cite this article
Scala, F., Brighenti, E., Govoni, M. et al. Direct relationship between the level of p53 stabilization induced by rRNA synthesis-inhibiting drugs and the cell ribosome biogenesis rate. Oncogene 35, 977–989 (2016). https://doi.org/10.1038/onc.2015.147
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/onc.2015.147
This article is cited by
-
Single-cell analysis of p53 transitional dynamics unravels stimulus- and cell type-dependent signaling output motifs
BMC Biology (2022)
-
BOP1 Silencing Suppresses Gastric Cancer Proliferation through p53 Modulation
Current Medical Science (2021)
-
Cell trapping microfluidic chip made of Cyclo olefin polymer enabling two concurrent cell biology experiments with long term durability
Biomedical Microdevices (2020)
-
The nucleolus: a central response hub for the stressors that drive cancer progression
Cellular and Molecular Life Sciences (2019)
-
The nucleolus, an ally, and an enemy of cancer cells
Histochemistry and Cell Biology (2018)