Abstract
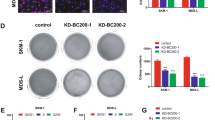
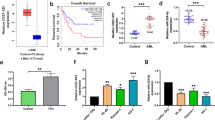
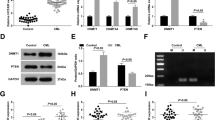
Aberrant expression of long noncoding RNAs (lncRNAs) is associated with various human cancers. However, the role of lncRNAs in Bcr-Abl-mediated chronic myeloid leukemia (CML) is unknown. In this study, we performed a comprehensive analysis of lncRNAs in human CML cells using an lncRNA cDNA microarray and identified an lncRNA termed lncRNA-BGL3 that acted as a key regulator of Bcr-Abl-mediated cellular transformation. Notably, we observed that lncRNA-BGL3 was highly induced in response to disruption of Bcr-Abl expression or by inhibiting Bcr-Abl kinase activity in K562 cells and leukemic cells derived from CML patients. Ectopic expression of lncRNA-BGL3 sensitized leukemic cells to undergo apoptosis and inhibited Bcr-Abl-induced tumorigenesis. Furthermore, transgenic (TG) mice expressing lncRNA-BGL3 were generated. We found that TG expression of lncRNA-BGL3 alone in mice was sufficient to impair primary bone marrow transformation by Bcr-Abl. Interestingly, we identified that lncRNA-BGL3 was a target of miR-17, miR-93, miR-20a, miR-20b, miR-106a and miR-106b, microRNAs that repress mRNA of phosphatase and tensin homolog (PTEN). Further experiments demonstrated that lncRNA-BGL3 functioned as a competitive endogenous RNA for binding these microRNAs to cross-regulate PTEN expression. Additionally, our experiments have begun to address the mechanism of how lncRNA-BGL3 is regulated in the leukemic cells and showed that Bcr-Abl repressed lncRNA-BGL3 expression through c-Myc-dependent DNA methylation. Taken together, these results reveal that Bcr-Abl-mediated cellular transformation critically requires silence of tumor-suppressor lncRNA-BGL3 and suggest a potential strategy for the treatment of Bcr-Abl-positive leukemia.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Chen Y, Peng C, Sullivan C, Li D, Li S . Critical molecular pathways in cancer stem cells of chronic myeloid leukemia. Leukemia 2010; 24: 1545–1554.
Cilloni D, Saglio G . Molecular pathways: BCR-ABL. Clin Cancer Res 2012; 18: 930–937.
Guo G, Qiu X, Wang S, Chen Y, Rothman PB, Wang Z et al. Oncogenic E17K mutation in the pleckstrin homology domain of AKT1 promotes v-Abl-mediated pre-B-cell transformation and survival of Pim-deficient cells. Oncogene 2010; 29: 3845–3853.
Yang J, Wang J, Chen K, Guo G, Xi R, Rothman PB et al. eIF4B phosphorylation by pim kinases plays a critical role in cellular transformation by Abl oncogenes. Cancer Res 2013; 73: 4898–4908.
Kapranov P, Cheng J, Dike S, Nix DA, Duttagupta R, Willingham AT et al. RNA maps reveal new RNA classes and a possible function for pervasive transcription. Science 2007; 316: 1484–1488.
Guttman M, Garber M, Levin JZ, Donaghey J, Robinson J, Adiconis X et al. Ab initio reconstruction of cell type-specific transcriptomes in mouse reveals the conserved multi-exonic structure of lincRNAs. Nat Biotechnol 2010; 28: 756–756.
Guttman M, Amit I, Garber M, French C, Lin MF, Feldser D et al. Chromatin signature reveals over a thousand highly conserved large non-coding RNAs in mammals. Nature 2009; 458: 223–227.
Ponting CP, Oliver PL, Reik W . Evolution and functions of long noncoding RNAs. Cell 2009; 136: 629–641.
Klattenhoff CA, Scheuermann JC, Surface LE, Bradley RK, Fields PA, Steinhauser ML et al. Braveheart, a long noncoding RNA required for cardiovascular lineage commitment. Cell 2013; 152: 570–583.
Sigova AA, Mullen AC, Molinie B, Gupta S, Orlando DA, Guenther MG et al. Divergent transcription of long noncoding RNA/mRNA gene pairs in embryonic stem cells. Proc Natl Acad Sci USA 2013; 110: 2876–2881.
Wang KC, Chang HY . Molecular mechanisms of long noncoding RNAs. Mol Cell 2011; 43: 904–914.
Yoon JH, Abdelmohsen K, Srikantan S, Yang X, Martindale JL, De S et al. LincRNA-p21 suppresses target mRNA translation. Mol Cell 2012; 47: 648–655.
Prensner JR, Iyer MK, Balbin OA, Dhanasekaran SM, Cao Q, Brenner JC et al. Transcriptome sequencing across a prostate cancer cohort identifies PCAT-1, an unannotated lincRNA implicated in disease progression. Nat Biotechnol 2011; 29: 742–749.
Yang F, Huo XS, Yuan SX, Zhang L, Zhou WP, Wang F et al. Repression of the long noncoding RNA-LET by histone deacetylase 3 contributes to hypoxia-mediated metastasis. Mol Cell 2013; 49: 1083–1096.
Tsai MC, Spitale RC, Chang HY . Long intergenic noncoding RNAs: new links in cancer progression. Cancer Res 2011; 71: 3–7.
Gupta RA, Shah N, Wang KC, Kim J, Horlings HM, Wong DJ et al. Long non-coding RNA HOTAIR reprograms chromatin state to promote cancer metastasis. Nature 2010; 464: 1071–1076.
Tsai MC, Manor O, Wan Y, Mosammaparast N, Wang JK, Lan F et al. Long noncoding RNA as modular scaffold of histone modification complexes. Science 2010; 329: 689–693.
Ji P, Diederichs S, Wang W, Boing S, Metzger R, Schneider PM et al. MALAT-1, a novel noncoding RNA, and thymosin beta4 predict metastasis and survival in early-stage non-small cell lung cancer. Oncogene 2003; 22: 8031–8041.
Tripathi V, Ellis JD, Shen Z, Song DY, Pan Q, Watt AT et al. The nuclear-retained noncoding RNA MALAT1 regulates alternative splicing by modulating SR splicing factor phosphorylation. Mol Cell 2010; 39: 925–938.
Guffanti A, Iacono M, Pelucchi P, Kim N, Solda G, Croft LJ et al. A transcriptional sketch of a primary human breast cancer by 454 deep sequencing. BMC Genomics 2009; 10: 163.
Luo JH, Ren B, Keryanov S, Tseng GC, Rao UN, Monga SP et al. Transcriptomic and genomic analysis of human hepatocellular carcinomas and hepatoblastomas. Hepatology 2006; 44: 1012–1024.
Heuston EF, Lemon KT, Arceci RJ . The beginning of the road for non-coding RNAs in normal hematopoiesis and hematologic malignancies. Front Genet 2011; 2: 94.
Wang KC, Yang YW, Liu B, Sanyal A, Corces-Zimmerman R, Chen Y et al. Long noncoding RNA programs active chromatin domain to coordinate homeotic gene expression. J Invest Dermatol 2011; 131: S63–S63.
Salmena L, Poliseno L, Tay Y, Kats L, Pandolfi PP . A ceRNA hypothesis: the Rosetta stone of a hidden RNA language? Cell 2011; 146: 353–358.
Tay Y, Kats L, Salmena L, Weiss D, Tan SM, Ala U et al. Coding-independent regulation of the tumor suppressor PTEN by competing endogenous mRNAs. Cell 2011; 147: 344–357.
Karreth FA, Tay Y, Perna D, Ala U, Tan SM, Rust AG et al. In vivo identification of tumor- suppressive PTEN ceRNAs in an oncogenic BRAF-induced mouse model of melanoma. Cell 2011; 147: 382–395.
Cesana M, Cacchiarelli D, Legnini I, Santini T, Sthandier O, Chinappi M et al. A long noncoding RNA controls muscle differentiation by functioning as a competing endogenous RNA. Cell 2011; 147: 358–369.
Poliseno L, Salmena L, Zhang J, Carver B, Haveman WJ, Pandolfi PP . A coding-independent function of gene and pseudogene mRNAs regulates tumour biology. Nature 2010; 465: 1033–1038.
de Giorgio A, Krell J, Harding V, Stebbing J, Castellano L . Emerging roles of ceRNAs in cancer: insights from the regulation of PTEN. Mol Cell Biol 2013; 33: 3976–3982.
Kiefer CM, Lee J, Hou C, Dale RK, Lee YT, Meier ER et al. Distinct Ldb1/NLI complexes orchestrate gamma-globin repression and reactivation through ETO2 in human adult erythroid cells. Blood 2011; 118: 6200–6208.
Chan CM, Fulton J, Montiel-Duarte C, Collins HM, Bharti N, Wadelin FR et al. A signature motif mediating selective interactions of BCL11A with the NR2E/F subfamily of orphan nuclear receptors. Nucleic Acids Res 2013; 41: 9663–9679.
Bu D, Yu K, Sun S, Xie C, Skogerbo G, Miao R et al. NONCODE v3.0: integrative annotation of long noncoding RNAs. Nucleic Acids Res 2012; 40: D210–215.
Nishikawa T, Ota T, Isogai T . Prediction whether a human cDNA sequence contains initiation codon by combining statistical information and similarity with protein sequences. Bioinformatics 2000; 16: 960–967.
Kong L, Zhang Y, Ye ZQ, Liu XQ, Zhao SQ, Wei L et al. CPC: assess the protein-coding potential of transcripts using sequence features and support vector machine. Nucleic Acids Res 2007; 35: W345–W349.
Lin MF, Jungreis I, Kellis M . PhyloCSF: a comparative genomics method to distinguish protein coding and non-coding regions. Bioinformatics 2011; 27: I275–I282.
Gesbert F, Griffin JD . Bcr/Abl activates transcription of the Bcl-X gene through STAT5. Blood 2000; 96: 2269–2276.
Lu D, Ma Y, Zhang W, Bao D, Dong W, Lian H et al. Knockdown of cytochrome P450 2E1 inhibits oxidative stress and apoptosis in the cTnT(R141W) dilated cardiomyopathy transgenic mice. Hypertension 2012; 60: 81–89.
Qiu X, Guo G, Chen K, Kashiwada M, Druker BJ, Rothman PB et al. A requirement for SOCS-1 and SOCS-3 phosphorylation in Bcr-Abl-induced tumorigenesis. Neoplasia 2012; 14: 547–558.
Chen Y, Chen J, Wang H, Shi J, Wu K, Liu S et al. HCV-induced miR-21 contributes to evasion of host immune system by targeting MyD88 and IRAK1. PLoS Pathog 2013; 9: e1003248.
Venturini L, Battmer K, Castoldi M, Schultheis B, Hochhaus A, Muckenthaler MU et al. Expression of the miR-17-92 polycistron in chronic myeloid leukemia (CML) CD34(+) cells. Blood 2007; 109: 4399–4405.
Prasanth KV, Prasanth SG, Xuan Z, Hearn S, Freier SM, Bennett CF et al. Regulating gene expression through RNA nuclear retention. Cell 2005; 123: 249–263.
Ma F, Xu S, Liu X, Zhang Q, Xu X, Liu M et al. The microRNA miR-29 controls innate and adaptive immune responses to intracellular bacterial infection by targeting interferon-gamma. Nat Immunol 2011; 12: 861–869.
Mizuno S, Chijiwa T, Okamura T, Akashi K, Fukumaki Y, Niho Y et al. Expression of DNA methyltransferases DNMT1, 3A, and 3B in normal hematopoiesis and in acute and chronic myelogenous leukemia. Blood 2001; 97: 1172–1179.
Sawyers CL, Callahan W, Witte ON . Dominant negative Myc blocks transformation by Abl oncogenes. Cell 1992; 70: 901–910.
Xie S, Lin H, Sun T, Arlinghaus RB . Jak2 is involved in c-Myc induction by Bcr-Abl. Oncogene 2002; 21: 7137.
Mott JL, Kurita S, Cazanave SC, Bronk SF, Werneburg NW, Fernandez-Zapico ME . Transcriptional suppression of mir-29b-1/mir-29a promoter by c-Myc, hedgehog, and NF-kappaB. J Cell Biochem 2010; 110: 1155–1164.
Mishra A, Liu S, Sams GH, Curphey DP, Santhanam R, Rush LJ et al. Aberrant overexpression of IL-15 initiates large granular lymphocyte leukemia through chromosomal instability and DNA hypermethylation. Cancer Cell 2012; 22: 645–655.
Barcelona SL, Thompson AA, Cote CJ . Intraoperative pediatric blood transfusion therapy: a review of common issues. Part I: Hematologic and physiologic differences from adults; metabolic and infectious risks. Pediatr Anesth 2005; 15: 716–726.
O'Hare T, Deininger MW, Eide CA, Clackson T, Druker BJ . Targeting the BCR-ABL signaling pathway in therapy-resistant Philadelphia chromosome-positive leukemia. Clin Cancer Res 2011; 17: 212–221.
Peng C, Chen YY, Yang ZF, Zhang HJ, Osterby L, Rosmarin AG et al. PTEN is a tumor suppressor in CML stem cells and BCR-ABL-induced leukemias in mice. Blood 2010; 115: 626–635.
Fontana L, Fiori ME, Albini S, Cifaldi L, Giovinazzi S, Forloni M et al. Antagomir-17-5p abolishes the growth of therapy-resistant neuroblastoma through p21 and BIM. PLoS One 2008; 3: e2236.
Chen CC, Stairs DB, Boxer RB, Belka GK, Horseman ND, Alvarez JV et al. Autocrine prolactin induced by the Pten-Akt pathway is required for lactation initiation and provides a direct link between the Akt and Stat5 pathways. Genes Dev 2012; 26: 2154–2168.
Ørom UA, Derrien T, Beringer M, Gumireddy K, Gardini A, Bussotti G et al. Long noncoding RNAs with enhancer-like function in human cells. Cell 2010; 143: 46–58.
Wang S, Li H, Chen YH, Wei HT, Gao GF, Liu HQ et al. Transport of influenza virus neuraminidase (NA) to host cell surface is regulated by ARHGAP21 and Cdc42 proteins. J Biol Chem 2012; 287: 9804–9816.
Chen JL, Limnander A, Rothman PB . Pim-1 and Pim-2 kinases are required for efficient pre-B-cell transformation by v-Abl oncogene. Blood 2008; 111: 1677–1685.
Acknowledgements
We thank Dr Paul B Rothman from the School of Medicine, The Johns Hopkins University for valuable discussions. This work was supported by the National Key Technologies Research and Development Program of China (2013ZX10004-611), National Basic Research Program (973) of China (2014CB541804), Intramural grant of the Chinese Academy of Sciences (KJZD-EW-L01-3), Natural Science Foundation of China (81171943, U1305212) and Hundreds of Talents Program of Chinese Academy of Sciences 2009–2014 to J-LC.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on the Oncogene website
Supplementary information
Rights and permissions
About this article
Cite this article
Guo, G., Kang, Q., Zhu, X. et al. A long noncoding RNA critically regulates Bcr-Abl-mediated cellular transformation by acting as a competitive endogenous RNA. Oncogene 34, 1768–1779 (2015). https://doi.org/10.1038/onc.2014.131
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/onc.2014.131
This article is cited by
-
Hypoxia-induced signaling in the cardiovascular system: pathogenesis and therapeutic targets
Signal Transduction and Targeted Therapy (2023)
-
Epigenetic regulation in hematopoiesis and its implications in the targeted therapy of hematologic malignancies
Signal Transduction and Targeted Therapy (2023)
-
A novel imatinib-upregulated long noncoding RNA plays a critical role in inhibition of tumor growth induced by Abl oncogenes
Molecular Cancer (2022)
-
BGL3 inhibits papillary thyroid carcinoma progression via regulating PTEN stability
Journal of Endocrinological Investigation (2021)
-
Role of non-coding RNA networks in leukemia progression, metastasis and drug resistance
Molecular Cancer (2020)