Abstract
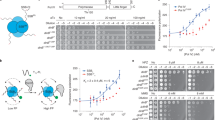
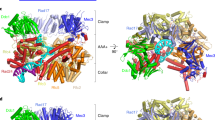
Sliding clamps are loaded onto DNA by ATP-dependent clamp loader complexes. A recent crystal structure of a clamp loader–clamp complex suggested an unexpected mechanism for DNA recognition, in which the ATPase subunits of the loader spiral around primed DNA. We report the results of fluorescence-based assays that probe the mechanism of the Escherichia coli clamp loader and show that conserved residues clustered within the inner surface of the modeled clamp loader spiral are critical for DNA recognition, DNA-dependent ATPase activity and clamp release. Duplex DNA with a 5′-overhang single-stranded region (corresponding to correctly primed DNA) stimulates clamp release, as does blunt-ended duplex DNA, whereas duplex DNA with a 3′ overhang and single-stranded DNA are ineffective. These results provide evidence for the recognition of DNA within an inner chamber formed by the spiral organization of the ATPase domains of the clamp loader.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
O'Donnell, M. & Studwell, P.S. Total reconstitution of DNA polymerase III holoenzyme reveals dual accessory protein clamps. J. Biol. Chem. 265, 1179–1187 (1990).
Stukenberg, P.T., Studwell-Vaughan, P.S. & O'Donnell, M. Mechanism of the sliding β-clamp of DNA polymerase III holoenzyme. J. Biol. Chem. 266, 11328–11334 (1991).
Kong, X.P., Onrust, R., O'Donnell, M. & Kuriyan, J. Three-dimensional structure of the beta subunit of E. coli DNA polymerase III holoenzyme: a sliding DNA clamp. Cell 69, 425–437 (1992).
Waga, S. & Stillman, B. Anatomy of a DNA replication fork revealed by reconstitution of SV40 DNA replication in vitro. Nature 369, 207–212 (1994).
Hacker, K.J. & Alberts, B.M. The slow dissociation of the T4 DNA polymerase holoenzyme when stalled by nucleotide omission. An indication of a highly processive enzyme. J. Biol. Chem. 269, 24209–24220 (1994).
Krishna, T.S.R., Kong, X.-P., Gary, S., Burgers, P. & Kuriyan, J. Crystal structure of the eukaryotic DNA polymerase processivity factor PCNA. Cell 79, 1233–1243 (1994).
Kelman, Z. & O'Donnell, M. DNA polymerase III holoenzyme: Structure and function of a chromosomal replicating machine. Annu. Rev. Biochem. 64, 171–200 (1995).
Waga, S. & Stillman, B. The DNA replication fork in eukaryotic cells. Annu. Rev. Biochem. 67, 721–751 (1998).
Cann, I.K. & Ishino, Y. Archaeal DNA replication: identifying the pieces to solve a puzzle. Genetics 152, 1249–1267 (1999).
Guenther, B.D., Onrust, R., Sali, A., O'Donnell, M. & Kuriyan, J. Crystal structure of the δ′ subunit of the clamp-loader complex of E. coli DNA polymerase III. Cell 91, 335–345 (1997).
Lupas, A.N. & Martin, J. AAA proteins. Curr. Opin. Struct. Biol. 12, 746–753 (2002).
Neuwald, A.F., Aravind, L., Spouge, J.L. & Koonin, E.V. AAA+: a class of chaperone-like ATPases associated with the assembly, operation, and disassembly of protein complexes. Genome Res. 9, 27–43 (1999).
Stewart, J., Hingorani, M.M., Kelman, Z. & O'Donnell, M. Mechanism of β clamp opening by the δ subunit of Escherichia coli DNA polymerase III holoenzyme. J. Biol. Chem. 276, 19182–19189 (2001).
Onrust, R., Stukenberg, P.T. & O'Donnell, M. Analysis of the ATPase subassembly which initiates processive DNA synthesis by DNA polymerase. J. Biol. Chem. 266, 21681–21686 (1991).
Tsurimoto, T. & Stillman, B. Replication factors required for SV40 DNA replication in vitro. I. DNA structure-specific recognition of a primer-template junction by eukaryotic DNA polymerases and their accessory proteins. J. Biol. Chem. 266, 1950–1960 (1991).
Yao, N., Leu, F.P., Anjelkovic, J., Turner, J. & O'Donnell, M. DNA structure requirements for the Escherichia coli γ complex clamp loader and DNA polymerase III holoenzyme. J. Biol. Chem. 275, 11440–11450 (2000).
Jeruzalmi, D., O'Donnell, M. & Kuriyan, J. Crystal structure of the processivity clamp loader γ complex of E. coli DNA polymerase III. Cell 106, 429–441 (2001).
Oyama, T., Ishino, Y., Cann, I.K., Ishino, S. & Morikawa, K. Atomic structure of the clamp loader small subunit from Pyrococcus furiosus. Mol. Cell 8, 455–463 (2001).
Bowman, G.D., O'Donnell, M. & Kuriyan, J. Structural analysis of a eukaryotic sliding DNA clamp-clamp loader complex. Nature 429, 724–730 (2004).
Kazmirski, S.P., Podobnik, M., Weitze, T.F., O'Donnell, M. & Kuriyan, J. Structural analysis of the inactive state of the Escherichia coli DNA polymerase clamp-loader complex. Proc. Natl. Acad. Sci. USA 101, 16750–16755 (2004).
Jeruzalmi, D., O'Donnell, M. & Kuriyan, J. Clamp loaders and sliding clamps. Curr. Opin. Struct. Biol. 12, 217–224 (2002).
Naktinis, V., Onrust, R., Fang, L. & O'Donnell, M. Assembly of a chromosomal replication machine: two DNA polymerases, a clamp loader, and sliding clamps in one holoenzyme particle. II. Intermediate complex between the clamp loader and its clamp. J. Biol. Chem. 270, 13358–13365 (1995).
Leu, F.P. & O'Donnell, M. Interplay of clamp loader subunits in opening the beta sliding clamp of Escherichia coli DNA polymerase III holoenzyme. J. Biol. Chem. 276, 47185–47194 (2001).
Turner, J., Hingorani, M.M., Kelman, Z. & O'Donnell, M. The internal workings of a DNA polymerase clamp-loading machine. EMBO J. 18, 771–783 (1999).
Goedken, E.R. et al. Fluorescence measurements on the E. coli DNA polymerase clamp loader: implications for conformational changes during ATP and clamp binding. J. Mol. Biol. 336, 1047–1059 (2004).
Jeruzalmi, D. et al. Mechanism of processivity clamp opening by the δ subunit wrench of the clamp loader complex of E. coli DNA polymerase III. Cell 106, 417–428 (2001).
Haugland, R.P. Handbook of Fluorescent Probes and Research Products 9th edn. (Molecular Probes, Eugene, Oregon, USA, 2002).
Bertram, J.G. et al. Pre-steady state analysis of the assembly of wild type and mutant circular clamps of Escherichia coli DNA polymerase III onto DNA. J. Biol. Chem. 273, 24564–24574 (1998).
Ason, B. et al. Mechanism of loading the Escherichia coli DNA polymerase III beta sliding clamp on DNA: Bona fide primer/templates preferentially trigger the γ complex to hydrolyze ATP and load the clamp. J. Biol. Chem. 278, 10033–10040 (2003).
Combeau, C. & Carlier, M.F. Probing the mechanism of ATP hydrolysis on F-actin using vanadate and the structural analogs of phosphate BeF-3 and A1F-4. J. Biol. Chem. 263, 17429–17436 (1988).
Chabre, M. Aluminofluoride and beryllofluoride complexes: a new phosphate analogs in enzymology. Trends Biochem. Sci. 15, 6–10 (1990).
Fisher, A.J. et al. X-ray structures of the myosin motor domain of Dictyostelium discoideum complexed with MgADP.BeFx and MgADP.AlF4. Biochemistry 34, 8960–8972 (1995).
Park, S., Ajtai, K. & Burghardt, T.P. Mechanism for coupling free energy in ATPase to the myosin active site. Biochemistry 36, 3368–3372 (1997).
Kagawa, R., Montgomery, M.G., Braig, K., Leslie, A.G. & Walker, J.E. The structure of bovine F(1)-ATPase inhibited by ADP and beryllium fluoride. EMBO J. 23, 2734–2744 (2004).
Johnson, A. & O'Donnell, M. Ordered ATP hydrolysis in the γ complex clamp loader AAA+ machine. J. Biol. Chem. 278, 14406–14413 (2003).
Snyder, A.K., Williams, C.R., Johnson, A., O'Donnell, M. & Bloom, L.B. Mechanism of loading the Escherichia coli DNA polymerase III sliding clamp: II. Uncoupling the β and DNA binding activities of the γ complex. J. Biol. Chem. 279, 4386–4393 (2004).
Haran, T.E., Joachimiak, A. & Sigler, P.B. The DNA target of the trp repressor. EMBO J. 11, 3021–3030 (1992).
Ellison, V. & Stillman, B. Biochemical characterization of DNA damage checkpoint complexes: clamp loader and clamp complexes with specificity for 5′ recessed DNA. PLoS Biol. 1, 231–243 (2003).
Miyata, T. et al. The clamp-loading complex for processive DNA replication. Nat. Struct. Mol. Biol. 11, 632–636 (2004).
O'Donnell, M., Jeruzalmi, D. & Kuriyan, J. Clamp loader structure predicts the architecture of DNA polymerase III holoenzyme and RFC. Curr. Biol. 11, R935–R946. (2001).
Barker, S.C. et al. Characterization of pp60c-src tyrosine kinase activities using a continuous assay: autoactivation of the enzyme is an intermolecular autophosphorylation process. Biochemistry 34, 14843–14851 (1995).
Henikoff, S. & Henikoff, J.G. Amino acid substitution matrices from protein blocks. Proc. Natl. Acad. Sci. USA 89, 10915–10919 (1992).
Acknowledgements
We gratefully recognize the assistance of X. Cao for DNA mutagenesis, L. Leighton for the creation of illustrations, and M. Levitus, R. McNally, M. Lamers, J. Guenther, O. Rosenberg and H. Sondermann for helpful discussions. This work was supported by an American Cancer Society postdoctoral fellowship to E.R.G., and by grants from the US National Institutes of Health to J.K. (GM45547) and M.O.D. (GM38839).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Fig. 1
FRET controls. (PDF 511 kb)
Supplementary Fig. 2
δ raw FRET data. (PDF 547 kb)
Supplementary Fig. 3
δ ATPase data. (PDF 572 kb)
Supplementary Fig. 4
Alternate hairpin DNA competition. (PDF 162 kb)
Rights and permissions
About this article
Cite this article
Goedken, E., Kazmirski, S., Bowman, G. et al. Mapping the interaction of DNA with the Escherichia coli DNA polymerase clamp loader complex. Nat Struct Mol Biol 12, 183–190 (2005). https://doi.org/10.1038/nsmb889
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb889
This article is cited by
-
Autoinhibition of a clamp-loader ATPase revealed by deep mutagenesis and cryo-EM
Nature Structural & Molecular Biology (2024)
-
Clamp loader ATPases and the evolution of DNA replication machinery
BMC Biology (2012)
-
Analysis of the role of PCNA-DNA contacts during clamp loading
BMC Structural Biology (2010)
-
The replication clamp-loading machine at work in the three domains of life
Nature Reviews Molecular Cell Biology (2006)