Abstract
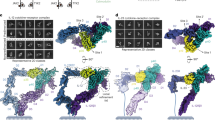
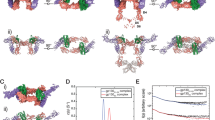
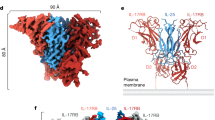
Interleukin-18 (IL-18), a cytokine formerly known as interferon-γ- (IFN-γ-) inducing factor, has pleiotropic immunoregulatory functions, including augmentation of IFN-γ production, Fas-mediated cytotoxicity and developmental regulation of T-lymphocyte helper type I. We determined the solution structure of IL-18 as a first step toward understanding its receptor activation mechanism. It folds into a β-trefoil structure that resembles that of IL-1. Extensive mutagenesis revealed the presence of three sites that are important for receptor activation: two serve as binding sites for IL-18 receptor α (IL-18Rα), located at positions similar to those of IL-1 for IL-1 receptor type I (IL-1RI), whereas the third site may be involved in IL-18 receptor β (IL-18Rβ) binding. The structure and mutagenesis data provide a basis for understanding the IL-18-induced heterodimerization of receptor subunits, which is necessary for receptor activation.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Okamura, H. et al. Cloning of a new cytokine that induces IFN-γ production by T cells. Nature 378, 88–91 (1995).
Dinarello, C.A. Interleukin-18, a proinflammatory cytokine. Eur. Cytokine Netw. 11, 483–486 (2000).
Nakanishi, K., Yoshimoto, T., Tsutsui H. & Okamura, H. Interleukin-18 is a unique cytokine that stimulates both Th1 and Th2 responses depending on its cytokine milieu. Cytokine Growth Factor Rev. 12, 53–72 (2001).
Konishi, H. et al. IL-18 contributes to the spontaneous development of atopic dermatitis-like inflammatory skin lesion independently of IgE/stat6 under specific pathogen-free conditions. Proc. Natl. Acad. Sci. USA 99, 11340–11345 (2002).
Wigginton, J.M., et al. Synergistic engagement of an ineffective endogenous anti-tumor immune response and induction of IFN-γ and Fas ligand-dependent tumor eradication by combined administration of IL-18 and IL-2. J. Immunol. 169, 4467–4474 (2002).
Vigers, G.P., Anderson, L.J., Caffes, P. & Brandhuber, B.J. Crystal structure of the type-I interleukin-1 receptor complexed with interleukin-1β. Nature 386, 190–194 (1997).
Schreuder, H. et al. A new cytokine-receptor binding mode revealed by the crystal structure of the IL-1 receptor with an antagonist. Nature 386, 194–200 (1997).
Bazan, J.F., Timans, J.C. & Kastelein, R.A. A newly defined interleukin-1? Nature 379, 591 (1996).
Kim, S.H. et al. Identification of amino acid residues critical for biological activity in human interleukin-18. J. Biol. Chem. 277, 10998–11003 (2002).
Born, T.L., Thomassen, E., Bird, T.A. & Sims, J.E. Cloning of a novel receptor subunit, AcPL, required for interleukin-18 signaling. J. Biol. Chem. 273, 29445–29450 (1998).
DeVos, A.M., Ultsch, M. & Kossiakoff, A.A. Human growth hormone and extracellular domain of its receptor: crystal structure of the complex. Science 255, 306–312 (1992).
Wiesmann, C. & de Vos, A.M. Variations on ligand-receptor complexes. Nat. Struct. Biol. 7, 440–442 (2000).
Wells, J.A., & de Vos, A.M. Hematopoietic receptor complexes. Annu. Rev. Biochem. 65, 609–634 (1996).
Hage, T., Sebald, W., & Reinemer, P. Crystal structure of the interleukin-4/receptor α chain complex reveals a mosaic binding interface. Cell 97, 271–281 (1999).
Schlessinger, J., et al. Crystal structure of a ternary FGF-FGFR-heparin complex reveals a dual role for heparin in FGFR binding and dimerization. Mol. Cell 6, 743–750 (2000).
Auron, P.E. The interleukin 1 receptor: ligand interactions and signal transduction. Cytokine Growth Factor Rev. 9, 221–237 (1998).
Kumar, S. et al. Identification and initial characterization of four novel members of the interleukin-1 family. J. Biol. Chem. 275, 10308–10314 (2000).
Born, T.L. et al. Identification and characterization of two members of a novel class of the interleukin-1 receptor (IL-1R) family. J. Biol. Chem. 275, 29946–29954 (2000).
Kim, S.H. et al. Structural requirements of six naturally occurring isoforms of the IL-18 binding protein to inhibit IL-18. Proc. Natl. Acad. Sci. USA 97, 1190–1195 (2000).
Xiang, Y. & Moss, B. Determination of the functional epitopes of human interleukin-18-binding protein by site-directed mutagenesis. J. Biol. Chem. 276, 17380–17386 (2001).
Watanabe, M., et al. Predominant expression of 950delCAG of IL-18R α chain cDNA is associated with reduced IFN-γ production and high serum IgE levels in atopic Japanese children. J. Allergy Clin. Immunol. 109, 669–675 (2002).
Evans, R.J. et al. Mapping receptor binding sites in interleukin (IL)-receptor antagonist and IL-1β by site-directed mutagenesis. Identification of a single site in IL-1Rα and two sites in IL-1β. J. Biol. Chem. 270, 11477–11483 (1995).
Cavanagh, J., Fairbrother, W.J., Palmer, A.G. III. & Skelton, N.J. Protein NMR Spectroscopy (Academic Press, San Diego, 1996).
Hu, W. & Zuiderweg, E.R. Stereospecific assignments of Val and Leu methyl groups in a selectively 13C-labeled 18 kDa polypeptide using 3D CT-(H)CCH-COSY and 2D 1JC-C edited heteronuclear correlation experiments. J. Magn. Reson. B 113, 70–75 (1996).
Güntert, P., Mumenthaler, C. & Wüthrich, K. Torsion angle dynamics for NMR structure calculation with the new program DYANA. J. Mol. Biol. 273, 283–298 (1997).
Cornilescu, G., Delaglio, F. & Bax, A. Protein backbone angle restraints from searching a database for chemical shift and sequence homology. J. Biomol. NMR 13, 289–302 (1999).
Hu, J.-S., Grzesiek, S. & Bax, A. Two-dimensional NMR methods for determining χ1 angles of aromatic residues in proteins from three bond JC′Cγ and JNCγ couplings. J. Am. Chem. Soc. 119, 1803–1804 (1997).
Herrmann, T., Güntert, P. & Wüthrich, K. Protein NMR structure determination with automated NOE assignment using the new software CANDID and the torsion angle dynamics algorithm DYANA. J. Mol. Biol. 319, 209–227 (2002).
Brünger, A.T. et al. Crystallography & NMR system: a new software suite for macromolecular structure determination. Acta Crystallogr. D 54, 905–921 (1998).
Koradi, R., Billeter, M. & Wüthrich, K. MOLMOL: a program for the display and analysis of macromolecular structures. J. Mol. Graph. 14, 51–55 (1996).
Laskowski, R.A., Rullman, J.A., MacArthur, M.W., Kaptein, R. & Thornton, J.M. AQUA and PROCHECK-NMR: programs for checking the quality of protein structures solved by NMR. J. Biomol. NMR 8, 477–486 (1996).
Shikano, H. et al. IFN-γ production in response to IL-18 or IL-12 stimulation by peripheral blood mononuclear cells of atopic patients. Clin. Exp. Allergy 31, 1263–1270 (2001).
Acknowledgements
We thank Y. Matsuo, T. Ikegami, T. Furuya and K. Kasahara for their technical assistance and K. Sukegawa and T. Fukao for critical reading of the manuscript.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Kato, Z., Jee, J., Shikano, H. et al. The structure and binding mode of interleukin-18. Nat Struct Mol Biol 10, 966–971 (2003). https://doi.org/10.1038/nsb993
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsb993
This article is cited by
-
IRAK4 Deficiency Presenting with Anti-NMDAR Encephalitis and HHV6 Reactivation
Journal of Clinical Immunology (2021)
-
The IL-1 family of cytokines and receptors in rheumatic diseases
Nature Reviews Rheumatology (2019)
-
An innate interaction between IL-18 and the propeptide that inactivates its precursor form
Scientific Reports (2019)
-
Immunodeficiency in Two Female Patients with Incontinentia Pigmenti with Heterozygous NEMO Mutation Diagnosed by LPS Unresponsiveness
Journal of Clinical Immunology (2017)
-
The structural basis for receptor recognition of human interleukin-18
Nature Communications (2014)