Abstract
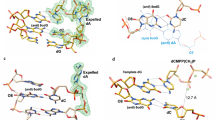
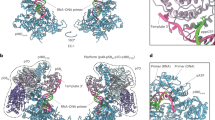
The structure of a large nucleic acid complex formed by the 10–23 DNA enzyme bound to an RNA substrate was determined by X–ray diffraction at 3.0 Å resolution. The 82–nucleotide complex contains two strands of DNA and two strands of RNA that form five double–helical domains. The spatial arrangement of these helices is maintained by two four–way junctions that exhibit extensive base–stacking interactions and sharp turns of the phosphodiester backbone stabilized by metal ions coordinated to nucleotides at these junctions. Although it is unlikely that the structure corresponds to the catalytically active conformation of the enzyme, it represents a novel nucleic acid fold with implications for the Holliday junction structure.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
References
Breaker, R.R. & Joyce, G.F. Chem. Biol. 1, 223–229 (1994).
Cuenoud, B. & Szostak, J.W. Nature 375, 611–614 (1995).
Breaker, R.R. Curr. Opin. Chem. Biol. 1, 26–31 (1997).
Santoro, S.W. & Joyce, G.F. Proc. Natl. Acad. Sci. USA 94, 4262–4266 (1997).
Santoro, S.W. & Joyce, G.F. Biochemistry 37,13330–13342 (1998).
Cate, J.H., Hanna, R.L. & Doudna, J.A. Nature Struct. Biol. 4, 553 –558 (1997).
Cowan, J.A. J. Inorg. Biochem. 49, 171–175 (1993).
Mollegaard, N.E., Murchie, A.I.H., Lilley, D.M.J. & Nielsen, P.E. EMBO J. 13, 1508–1513 ( 1994).
Heus, H.A. & Pardi, A. Science 253, 191–194 (1991).
Privé, G.G., Heinemann, U., Chandrasegaran, L.K., Kopka, M.L. & Dickerson, R.E. Science 238, 498–504 (1987).
Leonard, G.A. et al. Structure 2, 483–494 (1994).
Cheong, C.J., Varani, G. & Tinoco, I. Jr. Nature 346, 680–682 (1990).
Holbrook, S.R., Cheong, C., Tinoco, I. Jr. & Kim, S.H. Nature 353, 579–581 ( 1991).
Holliday, R. Genet. Res. 5, 282–304 ( 1964).
Sigal, N. & Alberts, B. J. Mol. Biol. 71, 789–793 (1972).
Lilley, D.M.J. & Clegg, R.M. Annu. Rev. Biophys. Biomol. Struct. 22, 299–328 (1993).
Seeman, N.C. & Kallenbach, N.R. Annu. Rev. Biophys. Biomol. Struct. 23, 53–86 ( 1994).
Hargreaves, D. et al. Nature Struct. Biol. 5, 441– 446 (1998).
Gopaul, D.N., Guo, F. & Van Duyne G.D. EMBO J. 17, 4175– 4187 (1998).
Murchie, A.I.H. et al. Nature 341, 763–766 (1989).
Cooper, J.P. & Hagerman, P.J. J. Mol. Biol. 198 , 711–719 (1987).
Duckett, D.R. et al. Cell 55, 79–89 (1988).
Churchill, M.E.A., Tullius, T.D., Kallenbach, N.R. & Seeman, N.C. Proc. Natl. Acad. Sci. USA 85, 4653– 4656 (1988).
Chen, S., Heffron, F. & Chazin, W.J. Biochemistry 32, 319– 326 (1993).
Duckett, D.R., Murchie, A.I.H. & Lilley, D.M.J. EMBO J. 9, 583– 590 (1990).
Miick, S.M., Fee, R.S., Millar, D.P. & Chazin, W.J. Proc. Natl. Acad. Sci. USA 94, 9080–9084 ( 1997).
Wincott, F. et al. Nucleic Acids Res. 23, 2677– 2684 (1995).
CCP4. Acta Crystallogr. D50, 760–763 (1994).
McRee, D.E. J. Mol. Graphics 10, 44–47 (1992).
Brünger, A.T. X–PLOR 3.1: A system for Crystallography and NMR (Yale University Press, New Haven, Connecticut; 1993).
Acknowledgements
The authors would like to thank E. Stura for crystallization advice, W. Chazin, D.E. McRee, S. Santoro and J. Williamson for helpful discussions, N. Kresge for help with data collection, and the staff at Stanford Synchrotron Radiation Laboratory for their generous assistance. NMR experiments were conducted at the TSRI Biomolecular NMR Laboratory with the assistance of W. Chazin. This work was funded by The Skaggs Institute for Chemical Biology (G.F.J.) and an NIH grant to C.D.S.
Author information
Authors and Affiliations
Corresponding authors
Rights and permissions
About this article
Cite this article
Nowakowski, J., Shim, P., Prasad, G. et al. Crystal structure of an 82-nucleotide RNA–DNA complex formed by the 10-23 DNA enzyme. Nat Struct Mol Biol 6, 151–156 (1999). https://doi.org/10.1038/5839
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/5839
This article is cited by
-
Structure of a 10-23 deoxyribozyme exhibiting a homodimer conformation
Communications Chemistry (2023)
-
Time-resolved structural analysis of an RNA-cleaving DNA catalyst
Nature (2022)
-
Crystal structure of an RNA-cleaving DNAzyme
Nature Communications (2017)
-
Aptasensor for environmental monitoring
Toxicology and Environmental Health Sciences (2017)
-
Crystal structure of a DNA catalyst
Nature (2016)