Abstract
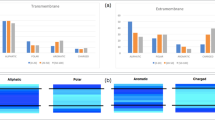
Polar residues in transmembrane α-helices may strongly influence the folding or association of integral membrane proteins. To test whether a motif that promotes helix association in a soluble protein could do the same within a membrane, we designed a model transmembrane helix based on the GCN4 leucine zipper. We found in both detergent miscelles and biological membranes that helix association is driven strongly by asparagine, independent of the rest of the hydrophobic leucine and/or valine sequence. Hydrogen bonding between membrane helices gives stronger associations than the packing of surfaces in glycophorin A helices, creating an opportunity to stabilize structures, but also implying a danger that non-specific interactions might occur. Thus, membrane proteins may fold to avoid exposure of strongly hydrogen bonding groups at their lipid exposed surfaces.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
References
Popot, J.-L. & Engelman, D.M. Membrane protein folding and oligomerization: the two-stage model. Biochemistry 29, 4031–4037 (1990).
Engelman, D.M. & Steitz, T.A. On the folding and insertion of globular membrane proteins. in The protein folding problem (ed. Wetlaufer, D.B.) 87–113 (Westview, Boulder, Colorado, 1984).
Lemmon, M.A. et al. Glycophorin A dimerization is driven by specific interactions between transmembrane α-helices. J. Biol. Chem. 267, 7683–7689 (1992).
Lemmon, M.A., Flanagan, J.M., Treutlein, H.R., Zhang, J. & Engelman, D.M. Sequence specificity in the dimerization of transmembrane α-helices. Biochemistry 31, 12719–12725 (1992).
Lemmon, M.A., Treutlein, H.R., Adams, P.D., Brunger, A.T. & Engelman, D.M. A dimerization motif for transmembrane α-helices . Nature Struct. Biol. 1, 157– 163 (1994).
Treutlein, H.R., Lemmon, M.A., Engelman, D.M. & Brunger, A.T. The glycophorin A transmembrane domain dimer: sequence-specific propensity for a right-handed supercoil of helices. Biochemistry 31, 12726–12733 (1992).
Langosch, D., Brosig, B., Kolmar, H. & Fritz, H.-J. Dimerisation of the glycophorin A transmembrane segment in membranes probed with the ToxR transcription activator. J. Mol. Biol., 525– 530 (1996).
MacKenzie, K.R., Prestegard, J.H. & Engelman, D.M. A transmembrane helix dimer: structure and implications . Science 276, 131–133 (1997).
MacKenzie, K.R. & Engelman, D.M. Structure-based prediction of the stability of transmembrane helix-helix interactions: the sequence dependence of glycophorin A dimerization. Proc. Natl. Acad. Sci. USA 95, 3583–3590 (1998).
Russ, W.P. & Engelman, D.M. TOXCAT: a measure of transmembrane helix association in a biological membrane. Proc. Natl. Acad. Sci. USA 96, 863–868 ( 1999).
Fleming, K.G., Ackerman, A.L. & Engelman, D.M. The effect of point mutations on the free energy of transmembrane α-helix dimerization. J. Mol. Biol. 272, 266–275 (1997).
Deisenhofer, J., Epp, O., Miki, K., Huber, R. & Michel, H. Structure of the protein subunits in the photosynthetic reaction centre of Rhodopseudomonas viridis at 3A resolution. Nature 318, 618–624 ( 1985).
Henderson, R. et al. Model for the structure of bacteriorhodopsin based on high-resolution electron cryo-microscopy. J. Mol. Biol. 213, 899–929 (1990).
O' Shea, E.K., Klemm, J.D., Kim, P.S. & Alber, T. X-ray structure of the GCN4 leucine zipper, a two-stranded, parallel coiled coil. Science 254, 539–544 ( 1991).
Jones, D.T., Taylor, W.R. & Thornton, J.M. A mutation data matrix for transmembrane proteins . FEBS Lett. 339, 269–275 (1994).
Samatey, F.A., Xu, C. & Popot, J.-L. On the distribution of amino acid residues in transmembrane α-helix bundles. Proc. Natl. Acad. Sci. USA 92, 4577–4581 (1995).
Arkin, I.T. & Brunger, A.T. Statistical analysis of predicted transmembrane α-helices. Biochim. Biophys. Acta 1429, 113–128 (1998).
Arkin, I. et al. Structural organization of the pentameric transmembrane α-helices of phospholamban, a cardiac ion channel. EMBO J. 13 , 4757–4764 (1994).
Kolmar, H. et al. Membrane insertion of the bacterial signal transduction protein ToxR and requirements of transcription activation studied by modular replacement of different protein substitution. EMBO J. 14, 3895–3904 (1995).
Harbury, P.B., Zhang, T., Kim, P.S. & Alber, T. A switch between two-, three- and four-stranded coiled coils in GCN4 leucine zipper mutants . Science 262, 1401–1407 (1993).
Lovejoy, B. et al. Crystal structure of a synthetic triple-stranded α-helical bundle. Science 259, 1288– 1293 (1993).
Zhu, B.-Y., Zhou, N.E., Kay, C.M. & Hodges, R.S. Packing and hydrophobicity effects on protein folding and stability: Effects of β-branched amino acids, valine and isoleucine, on the formation of stability of two-stranded α-helical coiled coils/leucine zippers. Protein Sci. 2, 383–394 (1993).
Zhou, N.E., Kay, C.M. & Hodges, R.S. The role of interhelical ionic interactions in controlling protein folding and stability: De novo designed synthetic two-stranded α-helical coiled coils. J. Mol. Biol. 237, 500– 512 (1994).
Zhou, N.E., Kay, C.M. & Hodges, R.S. The net energetic contribution of interhelical electrostatic attractions to coiled-coil stability. Protein Eng. 7, 1365–1372 (1994).
Lumb, K.J. & Kim, P.S. A buried polar interaction imparts structural uniqueness in a designed heterodimeric coiled coil. Biochemistry 34, 8642–8648 (1995).
Gurezka, R., Laage, R., Brosig, B. & Langosch, D. A heptad motif of leucine residues found in membrane proteins can drive self-assembly of artificial transmembrane segments. J. Biol. Chem. 274 , 9265–9270 (1999).
Potekhin, S.A., Medvedkin, V.N., Kashparov, I.A. & Venyaminov, S.Y. Synthesis and properties of the peptide corresponding to the mutant form of the leucine zipper of the transcriptional activator GCN4 form yeast. Protein Eng. 7, 1097–1101 (1994).
Vieth, M., Kolinski, A. & Skolnick, J. Method for predicting the state of association of discretized protein models: Applications to leucine zippers. Biochemistry 35, 955–967 (1996).
Zeng, X., Herndon, A.M. & Hu, J.C. Buried asparagines determine the dimerization specificities of leucine zipper mutants. Proc. Natl. Acad. Sci. USA 94, 3673–3678 (1997).
Choma, C., Gratkowski, H., Lear, J.D. & DeGrado, W.F. A membrane-soluble analogue of the two-stranded coiled coil from GCN4. Nature Struct. Biol. 7, 161–166 (2000).
Gonzalez, L., Woolfson, D.N. & Alber, T. Buried polar residues and structural specificity in the GCN4 leucine zipper. Nature Struct. Biol. 3, 1011–1018 (1996).
Gonzalez, L., Brown, R.A., Richardson, D. & Alber, T. Crystal structures of a single coiled-coil peptide in two oligomeric states reveal the basis for structural polymorphism. Nature Struct. Biol. 3, 1002–1010 ( 1996).
Gonzalez, L., Plecs, J.J. & Alber, T. An engineered allosteric switch in leucine-zipper oligomerization . Nature Struct. Biol. 3, 510– 515 (1996).
Drees, B.L., Grotkopp, E.K. & Nelson, H.C.M. The GCN4 leucine zipper can functionally substitute for the heat shock transcription factor's trimerization domain. J. Mol. Biol. 273, 61–74 (1997).
Junius, F.K. et al. Nuclear magnetic resonance characterization of the Jun leucine zipper domain: unusual properties of coiled-coil interfacial polar residues . Biochemistry 34, 6164– 6174 (1995).
Engelman, D.M., Steitz, T.A. & Goldman, A. Identifying nonpolar transbilayer helices in amino acid sequences of membrane proteins. Annu. Rev. Biophys. Biophys. Chem. 15, 321–353 ( 1986).
Bargmann, C.I., Hung, M.-C. & Weinberg, R.A. Multiple Independent activations of the neu oncogene by a point mutation altering the transmembrane domain of p185. Cell 45, 649–657 ( 1986).
Weiner, D.B., Liu, J., Cohen, J.A., Williams, W.V. & Greene, M.I. A point mutation in the neu oncogene mimics ligand induction of receptor aggregation. Nature 339, 230–231 (1989).
Hynes, N.E. & Stern, D.F. The biology of erbB-2/neu/HER-2 and its role in cancer. Biochim. Biophys. Acta 1198 , 165–184 (1994).
Smith, S.O., Smith, C.S. & Bormann, B.J. Strong hydrogen bonding interactions involving a buried glutamic acid in the transmembrane sequence of the neu/erbB-2 receptor. Nature Struct. Biol. 3, 252–258 (1996).
Fields, G.B. & Noble, R.L. Solid phase peptide synthesis utilizing 9-fluorenylmethoxycarbonyl amino acids. Int. J. Peptide Protein Res. 35, 161–214 ( 1990).
Piotto, M., Saudek, V. & Sklenar, V. Gradient-tailored excitation for single-quantum NMR spectroscopy of aqueous solutions. J. Biomol. NMR 2 , 661–665 (1992).
Altieri, A. & Byrd, R.A. Randomization approach to water suppression in multidimensional NMR using pulsed field gradients. J. Magn. Reson. 107B, 260–266 ( 1995).
Delaglio, F. et al. NMRPipe: a multidimensional spectral processing system based on UNIX pipes. J. Biomol. NMR 6, 277– 293 (1995).
Acknowledgements
We thank P.D. Adams, K. R. MacKenzie, A. Senes and I. Ubarretxena for helpful discussions. We are also indebted to L. Fisher for advice and assistance in peptide synthesis. We kindly acknowledge G. King for permission to adapt a figure. This research is funded with a program project grant on helix interactions in membrane proteins (National Institute of Health) and the National Foundation for Cancer Research.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Xiao Zhou, F., Cocco, M., Russ, W. et al. Interhelical hydrogen bonding drives strong interactions in membrane proteins. Nat Struct Mol Biol 7, 154–160 (2000). https://doi.org/10.1038/72430
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/72430
This article is cited by
-
Crystal structures of the human neurokinin 1 receptor in complex with clinically used antagonists
Nature Communications (2019)
-
Identifying ionic interactions within a membrane using BLaTM, a genetic tool to measure homo- and heterotypic transmembrane helix-helix interactions
Scientific Reports (2017)
-
Ser/Thr Motifs in Transmembrane Proteins: Conservation Patterns and Effects on Local Protein Structure and Dynamics
The Journal of Membrane Biology (2012)
-
Protein folding in membranes
Cellular and Molecular Life Sciences (2010)
-
Model of the whole rat AT1 receptor and the ligand-binding site
Journal of Molecular Modeling (2006)