Abstract
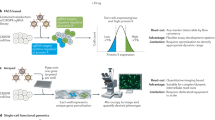
The accelerating pace of the discovery of genes has far surpassed our capabilities to understand their biological function — in other words, the phenotypes they engender. We need efficient and comprehensive large-scale phenotyping technologies. This presents a difficult challenge because phenotypes are numerous and diverse, and they can be observed and annotated at the molecular, cellular and organismal level. New technologies and approaches will therefore be required. Here, I describe recent efforts to develop new and efficient technologies for assessing cellular phenotypes.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Miller, J. H. A Short Course in Bacterial Genetics: A Laboratory Manual and Handbook for Escherichia coli and Related Bacteria (Cold Spring Harbor Lab. Press, New York, 1992).
Blattner, F. R. et al. The complete genome sequence of Escherichia coli K-12. Science 277, 1453–1474 (1997).
Goffeau, A. Life with 6,000 genes. Science 274, 563–567 (1996).
Kaul, S. et al. Analysis of the genome sequence of the flowering plant Arabidopsis thaliana. Nature 408, 796–815 (2000).
Goff, S. A. et al. A draft sequence of the rice genome (Oryza sativa L. ssp. japonica). Science 296, 92–100 (2002).
The C. elegans Sequencing Consortium. Genome sequence of C. elegans: a platform for investigating biology. Science 282, 2012–2018 (1998).
Adams, M. D. et al. The genome sequence of Drosophila melanogaster. Science 287, 2185–2195 (2000).
Waterston, R. H. et al. Initial sequencing and comparative analysis of the mouse genome. Nature 420, 520–562 (2002).
International Human Genome Sequencing Consortium. Initial sequencing and analysis of the human genome. Nature 409, 860–921 (2001).
Venter, J. C. et al. The sequence of the human genome. Science 291, 1304–1351 (2001).
Nadeau, J. H. et al. Functional annotation of mouse genome sequences. Science 291, 1251–1255 (2001).
Bassett, D. E. et al. Genome cross-referencing and XREFdb: implications for the identification and analysis of genes mutated in human diseases. Nature Genet. 15, 339–344 (1997).
Rubin, G. M. et al. Comparative genomics of eukaryotes. Science 287, 2204–2215 (2000).
LaRossa, R. A. in Escherichia coli and Salmonella Cellular and Molecular Biology 2nd edn Vol. 2 Ch. 39 (ed. Neidhart, F. C.) 2527–2587 (ASM, Washington DC, 1996).
Cheung, V. G. & Spielman, R. S. The genetics of variation in gene expression. Nature Genet. 32 (Suppl.), 522–525 (2002).
Jansen, R. C. Studying complex biological systems using multifactorial perturbation. Nature Rev. Genet. 4, 1–7 (2003).
Rosenthal, N. & Ashburner, M. Taking stock of our models: the function and future of stock centres. Nature Rev. Genet. 3, 711–717 (2002).
Eppig, J. T. Algorithms for mutant sorting: the need for phenotype vocabularies. Mamm. Genome 11, 584–589 (2000).
Pargent, W. et al. MouseNet database: digital management of a large-scale mutagenesis project. Mamm. Genome 11, 590–593 (2000).
Bochner, B. R. Sleuthing out bacterial identities. Nature 339, 157–158 (1989).
Brown, S. D. & Peters, J. Combining mutagenesis and genomics in the mouse — closing the phenotype gap. Trends Genet. 12, 433–435 (1996).
Paigen, K. & Epping, J. T. A mouse phenome project. Mamm. Genome 11, 715–717 (2000).
Moldin, S. O. et al. Trans-NIH neuroscience initiatives on mouse phenotyping and mutagenesis. Mamm. Genome 12, 575–581 (2001).
Beckers, J. & Hrabe de Angelis, M. Large-scale mutational analysis for the annotation of the mouse genome. Curr. Opin Chem. Biol. 6, 17–23 (2001).
Somerville, C. & Dangl, L. Genomics: plant biology in 2010. Science 290, 2077–2078 (2000).
Oliver, S. G. A network approach to the systematic analysis of yeast gene function. Trends Genet. 12, 241–242 (1996).
Hampsey, M. H. A review of phenotypes of Saccharomyces cerevisiae. Yeast 13, 1099–1133 (1997).
Niedenthal, R. et al. Systematic analysis of S. cerevisiae chromosome VII genes. Yeast 15, 1775–1796 (1999).
Bochner, B. R. 'Breathprints' at the microbial level. ASM News 55, 536–539 (1989).
Bochner, B. R., Gadzinski, P. & Panomitros, E. Phenotype MicroArrays for high-throughput phenotypic testing and assay of gene function. Genome Res. 11, 1246–1255 (2001).
Vazquez-Boland, J. A. et al. Listeria pathogenesis and molecular virulence determinants. Clin. Micro. Rev. 14, 584–640 (2001).
Guzman, L. M., Belin, D., Carson, M. J. & Beckwith, J. Tight regulation, modulation, and high-level expression by vectors containing the arabinose PBAD promoter. J. Bacteriol. 177, 4121–4130 (1995).
Ji, Y. et al. Identification of critical staphylococcal genes using conditional phenotypes generated by antisense RNA. Science 293, 2266–2269 (2001).
Smith, V. et al. Functional analysis of the genes of yeast chromosome V by genetic footprinting. Science 274, 2069–2074 (1996).
Shoemaker, D. D. et al. Quantitative phenotypic analysis of yeast deletion mutants using a highly parallel molecular bar-coding strategy. Nature Genet. 14, 450–456 (1996).
Winzeler, E. A. et al. Functional characterization of the S. cerevisiae genome by gene deletion and parallel analysis. Science 285, 901–906 (1999).
Giaever, G. et al. Functional profiling of the Saccharomyces cerevisiae genome. Nature 418, 387–391 (2002).
Rieger, K. J. et al. Large-scale phenotypic analysis — the pilot project on yeast chromosome III. Yeast 13, 1547–1562 (1997).
Rieger, K. J. et al. in Methods in Microbiology Vol. 28 (ed. Craig, A.G.) 205–227 (Academic Press, London, 1999).
Rieger, K. J. et al. Chemotyping of yeast mutants using robotics. Yeast 15, 973–986 (1999).
Entian, K. D. et al. Functional analysis of 150 deletion mutants in Saccharomyces cerevisiae by a systematic approach. Mol. Gen. Genet. 262, 683–702 (1999).
Ross-MacDonald, P. et al. Large-scale analysis of the yeast genome by transposon tagging and gene disruption. Nature 402, 413–418 (1999).
True, H. L. & Lindquist, S. L. A yeast prion provides a mechanism for genetic variation and phenotypic diversity. Nature 407, 477–483 (2000).
Sakumoto, N. et al. A series of double disruptants for protein phosphatase genes in Saccharomyces cerevisiae and their phenotypic analysis. Yeast 19, 587–599 (2002).
Warringer, J. & Blomberg, A. Automated screening in environmental arrays allows analysis of quantitative phenotypic profiles in Saccharomyces cerevisiae. Yeast 20, 53–67 (2003).
Acknowledgements
The author gratefully acknowledges and thanks his colleagues that have participated in the development of Phenotype MicroArray technology: X. H. Lei, A. Franco-Buff, R. Kostriken, J. Argyle, L. Wiater, J. Carlson, A. Morgan, C. Gorman, P. Gadzinski, E. Olender, E. Panomitros and L. He. We are also grateful for partial financial support of this project by Small Business Innovation Research Grants from the National Institutes of Health/National Institute of General Medical Sciences and from the National Aeronautics and Space Administration
Author information
Authors and Affiliations
Related links
Related links
DATABASES
Entrez
FURTHER INFORMATION
Glossary
- HETEROLOGOUS GENE
-
A gene that is transferred into a cell but originated in a cell from a different species.
- ISOGENIC
-
Cells or organisms that are derived from the same parent and have almost identical genomes.
- MASS SPECTROMETRY
-
An analytical tool for determining the molecular weight of a chemical.
- MULTI-STATE AUTOMATON
-
A self-acting and self-responding machine that has the ability to change itself into multiple states.
- PASSAGED STRAINS
-
Cells that have been repeatedly subcultured, typically under artificial in vitro laboratory-culture conditions and not in more natural in vivo conditions.
- PATHOGENICITY ISLAND
-
A contiguous block of genes, found in pathogenic microorganisms, in which at least a subset of the genes code for virulence factors.
- TETRAZOLIUM REDOX CHEMISTRY
-
A dye chemistry that absorbs the electrons produced by cellular respiration, causing a colour change as the tetrazolium dye is reduced.
- TN10 CASSETTE
-
A contiguous block of genes that is derived from the bacterial transposon Tn10, which confers resistance to tetracycline antibiotics.
Rights and permissions
About this article
Cite this article
Bochner, B. New technologies to assess genotype–phenotype relationships. Nat Rev Genet 4, 309–314 (2003). https://doi.org/10.1038/nrg1046
Issue Date:
DOI: https://doi.org/10.1038/nrg1046
This article is cited by
-
The Insertion in the 3′ UTR of Pmel17 Is the Causal Variant for Golden Skin Color in Tilapia
Marine Biotechnology (2022)
-
Population structure and uropathogenic potential of extended-spectrum cephalosporin-resistant Escherichia coli from retail chicken meat
BMC Microbiology (2021)
-
Diversity of soil microscopic filamentous fungi in Dystric Cambisol at the Banská Štiavnica – Šobov (Slovakia) locality after application of remediation measures
Biologia (2021)
-
A consensus S. cerevisiae metabolic model Yeast8 and its ecosystem for comprehensively probing cellular metabolism
Nature Communications (2019)
-
Developing genome-reduced Pseudomonas chlororaphis strains for the production of secondary metabolites
BMC Genomics (2017)