Abstract
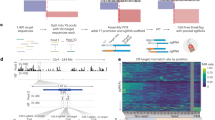
Methylation-sensitive single-nucleotide primer extension (Ms-SNuPE) is a technique that can be used for rapid quantitation of methylation at individual CpG sites. Treatment of genomic DNA with sodium bisulfite is used to convert unmethylated Cytosine to Uracil while leaving 5-methylcytosine unaltered. Strand-specific PCR is performed to generate a DNA template for quantitative methylation analysis using Ms-SNuPE. SNuPE is then performed with oligonucleotide(s) designed to hybridize immediately upstream of the CpG site(s) being interrogated. Reaction products are electrophoresed on polyacrylamide gels for visualization and quantitation by phosphorimage analysis. The Ms-SNuPE technique is similar to other quantitative assays that use bisulfite treatment of genomic DNA to discriminate unmethylated from methylated Cytosines (i.e., COBRA, pyrosequencing). Ms-SNuPE can be used for high-throughput methylation analysis and rapid quantitation of Cytosine methylation suitable for a wide range of biological investigations, such as checking aberrant methylation changes during tumorigenesis, monitoring methylation changes induced by DNA methylation inhibitors or for measuring hemimethylation. Approximately two to four CpG sites can be interrogated in up to 40 samples by Ms-SNuPE in less than 5 h, after PCR amplification of the desired target sequence and preparation of PCR amplicons.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Laird, P.W. & Jaenisch, R. The role of DNA methylation in cancer genetic and epigenetics. Annu. Rev. Genet. 30, 441–464 (1996).
Gaudet, F. et al. Induction of tumors in mice by genomic hypomethylation. Science 300, 489–492 (2003).
Venter, J.C. et al. The sequence of the human genome. Science 291, 1304–1351 (2001).
Jones, P.A. & Baylin, S.B. The fundamental role of epigenetic events in cancer. Nat. Rev. Genet. 3, 415–428 (2002).
Jones, P.A. & Baylin, S.B. The epigenomics of cancer. Cell 128, 683–692 (2007).
Feinberg, A.P. & Vogelstein, B. Hypomethylation distinguishes genes of some human cancers from their normal counterparts. Nature 301, 89–92 (1983).
Singer-Sam, J., LeBon, J.M., Tanguay, R.L. & Riggs, A.D. A quantitative HpaII-PCR assay to measure methylation of DNA from a small number of cells. Nucleic Acids Res. 18, 687(1990).
Herman, J.G., Graff, J.R., Myohanen, S., Nelkin, B.D. & Baylin, S.B. Methylation-specific PCR: a novel PCR assay for methylation status of CpG islands. Proc. Natl. Acad. Sci. USA 93, 9821–9826 (1996).
Kuppuswamy, M.N. et al. Single nucleotide primer extension to detect genetic diseases: experimental application to hemophilia B (factor IX) and cystic fibrosis genes. Proc. Natl. Acad. Sci. USA 88, 1143–1147 (1991).
Singer-Sam, J., LeBon, J.M., Dai, A. & Riggs, A.D. A sensitive, quantitative assay for measurement of allele-specific transcripts differing by a single nucleotide. PCR Methods Appl. 1, 160–163 (1992).
Szabo, P.E. & Mann, J.R. Allele-specific expression and total expression levels of imprinted genes during early mouse development: implications for imprinting mechanisms. Genes Dev. 9, 3097–3108 (1995).
Greenwood, A.D. & Burke, D.T. Single nucleotide primer extension: quantitative range, variability, and multiplex analysis. Genome Res. 6, 336–348 (1996).
Gonzalgo, M.L. & Jones, P.A. Rapid quantitation of methylation differences at specific sites using methylation-sensitive single nucleotide primer extension (Ms-SNuPE). Nucleic Acids Res. 25, 2529–2531 (1997).
Saito, Y. et al. Specific activation of microRNA-127 with downregulation of the proto-oncogene BCL6 by chromatin-modifying drugs in human cancer cells. Cancer Cell 9, 435–443 (2006).
Liang, G., Robertson, K.D., Talmadge, C., Sumegi, J. & Jones, P.A. The gene for a novel transmembrane protein containing epidermal growth factor and follistatin domains is frequently hypermethylated in human tumor cells. Cancer Res. 60, 4907–4912 (2000).
Cheng, J.C. et al. Inhibition of DNA methylation and reactivation of silenced genes by zebularine. J. Natl. Cancer Inst. 95, 399–409 (2003).
Cheng, J.C. et al. Continuous zebularine treatment effectively sustains demethylation in human bladder cancer cells. Mol. Cell. Biol. 24, 1270–1278 (2004).
Egger, G. et al. Identification of DNMT1 (DNA methyltransferase 1) hypomorphs in somatic knockouts suggests an essential role for DNMT1 in cell survival. Proc. Natl. Acad. Sci. USA 103, 14080–14085 (2006).
Liang, G. et al. Cooperativity between DNA methyltransferases in the maintenance methylation of repetitive elements. Mol. Cell. Biol. 22, 480–491 (2002).
Kaminsky, Z.A., Assadzadeh, A., Flanagan, J. & Petronis, A. Single nucleotide extension technology for quantitative site-specific evaluation of metC/C in GC-rich regions. Nucleic Acids Res. 33, e95 (2005).
Frommer, M. et al. A genomic sequencing protocol that yields a positive display of 5-methylcytosine residues in individual DNA strands. Proc. Natl. Acad. Sci. USA 89, 1827–1831 (1992).
Fatemi, M. et al. Footprinting of mammalian promoters: use of a CpG DNA methyltransferase revealing nucleosome positions at a single molecule level. Nucleic Acids Res. 33, e176 (2005).
Markl, I.D. & Jones, P.A. Presence and location of TP53 mutation determines pattern of CDKN2A/ARF pathway inactivation in bladder cancer. Cancer Res. 58, 5348–5353 (1998).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
M.L.G. and G.L. have a patent (United States patent 6,811,982) entitled “A cancer diagnostic method based upon DNA methylation differences.“
Rights and permissions
About this article
Cite this article
Gonzalgo, M., Liang, G. Methylation-sensitive single-nucleotide primer extension (Ms-SNuPE) for quantitative measurement of DNA methylation. Nat Protoc 2, 1931–1936 (2007). https://doi.org/10.1038/nprot.2007.271
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2007.271
This article is cited by
-
Examinations of maternal uniparental disomy and epimutations for chromosomes 6, 14, 16 and 20 in Silver-Russell syndrome-like phenotypes
BMC Medical Genetics (2016)
-
Renal kallikrein excretion and epigenetics in human acute kidney injury: Expression, mechanisms and consequences
BMC Nephrology (2011)
-
An Overview of Epigenetic Assays
Molecular Biotechnology (2008)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.