Abstract
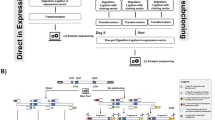
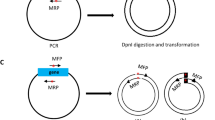
Extension of overlapping gene segments by PCR is a simple, versatile technique for site-directed mutagenesis and gene splicing. Initial PCRs generate overlapping gene segments that are then used as template DNA for another PCR to create a full-length product. Internal primers generate overlapping, complementary 3′ ends on the intermediate segments and introduce nucleotide substitutions, insertions or deletions for site-directed mutagenesis, or for gene splicing, encode the nucleotides found at the junction of adjoining gene segments. Overlapping strands of these intermediate products hybridize at this 3′ region in a subsequent PCR and are extended to generate the full-length product amplified by flanking primers that can include restriction enzyme sites for inserting the product into an expression vector for cloning purposes. The highly efficient generation of mutant or chimeric genes by this method can easily be accomplished with standard laboratory reagents in approximately 1 week.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Hunt, H.D., Pullen, J.K., Dick, R.F., Bluestone, J.A. & Pease, L.R. Structural basis of Kbm8 alloreactivity. Amino acid substitutions on the beta-pleated floor of the antigen recognition site. J. Immunol. 145, 1456–1462 (1990).
Pullen, J.K., Hunt, H.D. & Pease, L.R. Peptide interactions with the Kb antigen recognition site. J. Immunol. 146, 2145–2151 (1991).
Lohi, J., Lehti, K., Valtanen, H., Parks, W.C. & Keski-Oja, J. Structural analysis and promoter characterization of the human membrane-type matrix metalloproteinase-1 (MT1-MMP) gene. Gene 242, 75–86 (2000).
Seyboldt, C., Granzow, H. & Osterrieder, N. Equine herpesvirus 1 (EVH-1) glycoprotein M: effect of deletions of transmembrane domains. Virology 278, 477–489 (2000).
Lemberg, M.K. & Martoglio, B. Requirements for signal peptide peptidase-catalyzed intramembrane proteolysis. Mol Cell 10, 735–744 (2002).
Ho, S.N., Hunt, H.D., Horton, R.M., Pullen, J.K. & Pease, L.R. Site-directed mutagenesis by overlap extension using the polymerase chain reaction. Gene 77, 51–59 (1989).
Horton, R.M., Hunt, H.D., Ho, S.N., Pullen, J.K. & Pease, L.R. Engineering hybrid genes without the use of restriction enzymes: gene splicing by overlap extension. Gene 77, 61–68 (1989).
Zoller, M.J. & Smith, M. Oligonucleotide-directed mutagenesis using M13-derived vectors: an efficient and general procedure for the production of point mutations in any fragment of DNA. Nuc. Acids Res. 10, 6487–6500 (1982).
Morinaga, Y., Franceschini, T., Inouye, S. & Inouye, M. Improvement of oligonucleotide-directed site-specific mutagenesis using double-stranded plasmid DNA. Biotechnology 2, 636–639 (1984).
Lai, D., Zhu, X. & Pestka, S. A simple and efficient method for site-directed mutagenesis with double-stranded plasmid DNA. Nucleic Acids Res. 21, 3977–3980 (1993).
Brown, T. Hybridization analysis of DNA blots. in Current Protocols in Molecular Biology, Unit 2.10, January 1993 (ed. Harkins, B.) 2.10.1–2.10.16 (John Wiley & Sons Inc., Hoboken, NJ, 2003).
Duby, A., Jacobs, K.A. & Celeste, A. Using synthetic oligonucleotides as probes. in Current Protocols in Molecular Biology, Unit 6.4, January 1990 (ed. Harkins, B.) 6.4.1–6.4.10 (John Wiley & Sons Inc., Hoboken, NJ, 2003).
Engebrecht, J., Brent, R. & Kaderbhai, M.A. Minipreps of plasmid DNA. in Current Protocols in Molecular Biology, Unit 1.6, July 1991 (ed. Harkins, B.) 1.6.1–1.6.10 (John Wiley & Sons Inc., Hoboken, NJ, 2003).
Slatko, B.E., Eckert, R.L., Albright, L.M. & Ausubel, F.M. DNA sequencing strategies. in Current Protocols in Molecular Biology, Unit 7.1, April 1999 (ed. Harkins, B.) 7.1.1–7.1.7 (John Wiley & Sons Inc., Hoboken, NJ, 2003).
Kammann, M., Laufs, J., Schell, J. & Gronenborn, B. Rapid insertional mutagenesis of DNA by polymerase chain reaction (PCR). Nucleic Acids Res. 17, 5404 (1989).
Sarkar, G. & Sommer, S.S. The “megaprimer” method of site-directed mutagenesis. Biotechniques 8, 404–407 (1990).
Mikaelian, I. & Sergeant, A. Modification of the overlap extension method for extensive mutagenesis on the same template. Methods Mol. Biol. 57, 193–202 (1996).
Urban, A., Neukirchen, S. & Jaeger, K. A rapid and efficient method for site-directed mutagenesis using one-step overlap extension PCR. Nucleic Acids Res. 25, 2227–2228 (1997).
Kramer, M.F. & Coen, D.M. Enzymatic amplification of DNA by PCR: standard procedures and optimization. in Current Protocols in Molecular Biology, Unit 15.1, October 2001 (ed. Harkins, B.) 15.1.1–15.1.14 (John Wiley & Sons Inc., Hoboken, NJ, 2003).
Rohren, E.M., Pease, L.R., Ploegh, H.L. & Schumacher, T.N.M. Polymorphisms in pockets of major histocompatibility complex class I molecules influence peptide preference. J. Exp. Med. 177, 1713–1721 (1993).
Weiss, E. et al. The DNA sequence of the H-2Kb gene: evidence for gene conversion as a mechanism for the generation of polymorphism in histocompatibility antigens. EMBO J. 2, 453–462 (1983).
Acknowledgements
We thank Rudy Hanson for his technical contributions to this paper and Virginia Van Keulen and Michael Bell for helpful discussions.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
Mayo Clinic is assigned intellectual property rights related to the described technology that have been and continue to be licensed in a non-exclusive manner to a number of non-academic private interests. Mayo Clinic and I (author) have received royalty payments in excess of $10,000. K. Heckman has no competing financial interest.
Rights and permissions
About this article
Cite this article
Heckman, K., Pease, L. Gene splicing and mutagenesis by PCR-driven overlap extension. Nat Protoc 2, 924–932 (2007). https://doi.org/10.1038/nprot.2007.132
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2007.132
This article is cited by
-
A novel transcriptional repressor specifically regulates xylanase gene 1 in Trichoderma reesei
Biotechnology for Biofuels and Bioproducts (2023)
-
Genome-wide association analysis reveals a novel pathway mediated by a dual-TIR domain protein for pathogen resistance in cotton
Genome Biology (2023)
-
Computational remodeling of an enzyme conformational landscape for altered substrate selectivity
Nature Communications (2023)
-
Rational design of PD-1-CD28 immunostimulatory fusion proteins for CAR T cell therapy
British Journal of Cancer (2023)
-
Understanding the Necessity of Regulatory Protein Machinery in Heterologous Expression of Class-III Type of Ocins
The Protein Journal (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.