Abstract
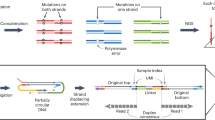
We describe a protocol that uses a bioinformatically optimized primer in an isothermal whole genome amplification (WGA) reaction. Overnight incubation at 37 °C efficiently generates several hundred- to several thousand-fold increases in input DNA. The amplified product retains reasonably faithful quantitative representation of unamplified whole genomic DNA (gDNA). We provide protocols for applying this isothermal primer extension WGA protocol in three different techniques of genomic analysis: comparative genomic hybridization (CGH), genotyping at simple tandem repeat (STR) loci and screening for single base mutations in a common monogenic disorder, β-thalassemia. gDNA extracted from formalin-fixed paraffin-embedded (FFPE) tissues can also be amplified with this protocol.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Dean, F.B. et al. Comprehensive human genome amplification using multiple displacement amplification. Proc. Natl. Acad. Sci. USA 99, 5261–5266 (2002).
Lage, J.M. et al. Whole genome analysis of genetic alterations in small DNA samples using hyperbranched strand displacement amplification and array-CGH. Genome Res. 13, 294–307 (2003).
Telenius, H. et al. Degenerate oligonucleotide-primed PCR: general amplification of target DNA by a single degenerate primer. Genomics 13, 718–725 (1992).
Zhang, L. et al. Whole genome amplification from a single cell: Implications for genetic analysis. Proc. Natl. Acad. Sci. USA 89, 5847–5851 (1992).
Klein, C.A. et al. Comparative genomic hybridization, loss of heterozygosity, and DNA sequence analysis of single cells. Proc. Natl. Acad. Sci. USA 96, 4494–4499 (1999).
Png, A.E.H., Choo, K.W., Lee, C.I.P., Leong, S.H. & Kon, O.L. Primer design for whole genome amplification using genetic algorithms. In Silico Biology 6, 47 (2006).
Lee, C.I.P. et al. An isothermal method for whole genome amplification of fresh and degraded DNA for comparative genomic hybridization, genotyping and mutation detection. DNA Res. 13, 77–88 (2006).
Feinberg, A.P. & Vogelstein, B. A technique for radiolabeling DNA restriction endonuclease fragments to high specific activity. Anal. Biochem. 132, 6–13 (1983).
Snijders, A.M. et al. Assembly of microarrays for genome-wide measurement of DNA copy number by CGH. Nat. Genet. 29 (3): 263–264 (2001).
Segraves, R., Albertson, D. & Pinkel, D. CGH using BAC genomic microarrays. In DNA Microarrays: A Molecular Cloning Manual (eds. Botwell, D. & Sambrook, J.) 6.380–6.385 (Cold Spring Harbor Laboratory Press, Cold Spring Harbor, New York, USA, 2003).
Acknowledgements
This work was supported by the National Medical Research Council, Singapore.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Lee, C., Leong, S., Png, A. et al. An isothermal primer extension method for whole genome amplification of fresh and degraded DNA: applications in comparative genomic hybridization, genotyping and mutation screening. Nat Protoc 1, 2185–2194 (2006). https://doi.org/10.1038/nprot.2006.398
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2006.398
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.