Abstract
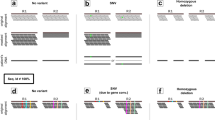
Next-generation sequencing produces high-throughput data, albeit with greater error and shorter reads than traditional Sanger sequencing methods. This complicates the detection of genomic variations, especially, small insertions and deletions. Here we describe ParMap, a statistical algorithm for the identification of complex genetic variants using partially mapped reads in nextgen sequencing data. We also report ParMap’s successful application to the mutation analysis of chromosome X exome-captured leukemia DNA samples.
Similar content being viewed by others
Article PDF
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Khiabanian, H., Van Vlierberghe, P., Palomero, T. et al. ParMap, an Algorithm for the Identification of Complex Genomic Variations in Nextgen Sequencing Data. Nat Prec (2010). https://doi.org/10.1038/npre.2010.4145.1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/npre.2010.4145.1