Abstract
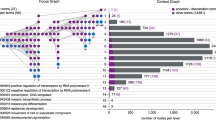
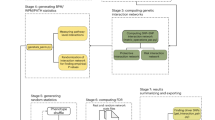
Epistatic or genetic interactions, representing the effects of mutations on the phenotypes caused by other mutations, can be very helpful for uncovering functional relationships between genes. Recently, the Epistasis Miniarray Profile (E-MAP) method has emerged as a powerful approach for identifying such interactions systematically. As part of this approach, hierarchical clustering is used to partition genes into groups on the basis of the similarity between their global interaction profiles. Here we present an original biclustering algorithm for identifying groups of functionally related genes from E-MAP data in a manner that allows individual genes to be assigned to more than one functional group. This enables investigation of the pleiotropic nature of gene function, a goal that cannot be achieved with hierarchical clustering. The performance of our algorithm is illustrated by applying it to two E-MAP datasets and an E-MAP-like in silico dataset for the yeast S. cerevisiae. In addition to identifying the majority of the functional modules reported in these studies, our algorithm uncovers many recently documented and novel multi-functional relationships between genes and gene groups.
Similar content being viewed by others
Article PDF
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Pu, S., Ronen, K., Vlasblom, J. et al. Analysis of Genetic Interaction Maps Reveals Functional Pleiotropy. Nat Prec (2008). https://doi.org/10.1038/npre.2008.2182.1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/npre.2008.2182.1