Abstract
Dynamic interplay between the plasma membrane and underlying cytoskeleton is essential for cellular shape change. Spatial organization of actin filaments, the growth of which generates membrane deformations during motility1, phagocytosis2, endocytosis3 and cytokinesis4, is mediated by specific protein–protein interactions that branch, crosslink and bundle filaments into networks that interact with the membrane. Although membrane curvature has been found to influence binding of proteins with curvature-sensitive domains5, the direct effect of membrane elasticity on cytoskeletal network organization is not clear. Here, we show through in vitro reconstitution and elastic modelling that a lipid bilayer can drive the emergence of bundled actin filament protrusions from branched actin filament networks, thus playing a role normally attributed to actin-binding proteins. Formation of these filopodium-like protrusions with only a minimal set of purified proteins points to an active participation of the membrane in organizing actin filaments at the plasma membrane. In this way, elastic interactions between the membrane and cytoskeleton can cooperate with accessory proteins to drive cellular shape change.
Similar content being viewed by others
Main
Changes in cell shape require reorganization of both the cytoskeleton and the plasma membrane. The cytoskeleton provides cells with internal organization and mechanical rigidity and is composed of filamentous protein polymers assembled by accessory proteins into networks. In contrast, the membrane is a comparatively soft barrier that serves as a substrate for enzymatic reactions and compartmentalizes cells by restricting diffusion. In many modes of cellular shape change, the actin cytoskeleton adjacent to the membrane dynamically reorganizes from one specific arrangement of filaments to another. The growth of dendritic actin networks against the membrane at the leading edge of crawling cells is a well-studied example. A model that details the interactions of actin and actin-binding proteins in protruding lamellipodia, known as the dendritic nucleation model, has been supported by reconstitution of dendritic actin networks that propel solid objects6,7,8. However, the influence of the cell membrane, which is both deformable and fluid, on actin network assembly is unclear. Cell-free reconstitution of vesicle motility in cytoplasmic extracts demonstrated that growth of a dendritic actin network can deform spherical vesicles to teardrop shapes in cell extracts9,10, but it is difficult to attribute the shape change to specific molecular interactions owing to the complexity of the extracts.
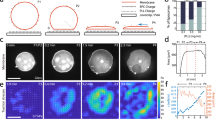
Here, we used giant unilamellar vesicles (GUVs) that nucleated assembly of a dendritic actin network to study the interactions between actin network growth and deformable membranes. Reconstituted actin networks containing actin, neural-Wiscott Aldrich Syndrome Protein (N-WASP), and the seven-subunit Actin-related protein 2/3 (Arp2/3) complex were assembled onto synthetic lipid bilayers as previously described11. When the vesicles were incubated with the purified proteins, local activation of the nucleation promotion factor N-WASP led to Arp2/3-branched dendritic actin network formation on the external leaflet of the GUVs. Surprisingly, we observed thin protrusions emanating from the Arp2/3-branched network (Fig. 1) in nearly 95% of the vesicles over 10 μm in diameter. We observed similar phenotypes for N-WASP activation by the synergistic activity of phosphoinositol 4,5 bisphosphate (PIP2) and small Rho GTPase Cdc42 (ref. 12), by dipyrromethene boron difluoride (BODIPY) tetramethylrhodamine-PIP2 (TMR-PIP2) alone11 and by His-tagged protein-binding lipid DOGS-Ni-NTA, which binds and activates His-tagged N-WASP, in the absence of both PIP2 and Cdc42.
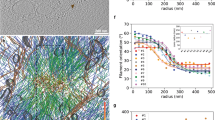
a, Phase-contrast and spinning–disc confocal images of membrane (green) and actin (red) show multiple protrusions in the lumen of a GUV. Overlay of the fluorescence images confirms that the membrane protrusions are supported by actin filaments. Scale bar, 5 μm. b, Localization of AF555 actin and AF488 capping protein (CP) fluorescence in thin actin filament protrusions shows that filament barbed ends are concentrated at the tip of the protrusions. Scale bar, 10 μm. c, Localization of actin, Arp2/3 complex and capping protein along thin actin filament protrusions. The normalized Arp2/3 complex (n=4) and capping protein (n=3) traces were divided by the normalized actin (n=7) line scans. d, Elongation of a thin protrusion visualized by phase-contrast microscopy. A second, independent protrusion enters the field at 10 min and crosses the path of the protrusion that is tracked (red arrows). Scale bar, 3 μm. The length of the protrusion was tracked through time showing that growth initially occurs quickly but slows down over time. Inset: Kymograph of fluorescently labelled membrane.
The membrane deformations generated by actin network growth and protruding into the vesicles can be easily identified with phase-contrast microscopy. Using confocal microscopy, we confirmed that the thin protrusions consisted of actin filaments encapsulated by a membrane tube (Fig. 1a). To determine the arrangement of actin filaments within the membrane protrusions, we used fluorescently labelled capping protein as a marker for F-actin barbed ends after elongation halted (Fig. 1b). Line-scan analysis of the fluorescently labelled protein demonstrated that F-actin barbed ends were concentrated at the tip of the thin actin filament protrusions (Fig. 1c), consistent with parallel filaments in the protrusion. As expected, the presence of capping protein during assembly of the actin network inhibited formation of the protrusions. Further experiments with fluorescently labelled N-WASP and Arp2/3 complex also suggested that the protrusions lacked dendritic architecture along their length (see Supplementary Information).
Typically, the thin actin filament protrusions formed from the dense membrane-associated dendritic actin networks within the first few minutes after introduction of purified proteins and grew into the lumen of the GUVs. Elongation was tracked over time using epifluorescence microscopy of fluorescently labelled lipids and confirmed with phase-contrast microscopy (Fig. 1d). The thin actin filament protrusion in Fig. 1d initially grew at a rate of ∼1 μm min−1, gradually slowed, then halted at a terminal length of ∼24 μm. We found that the stationary length of thin actin filament protrusions ranged from 1 to 25 μm (n>1,000). During elongation, the protrusions remained straight with no visible lateral fluctuations. Using a kinetic model based on that of Mogilner and Rubinstein13, we found that the observed elongation dynamics were consistent with growth of ∼10 filaments within a single protrusion (see Supplementary Information).
To further examine the growth of the thin actin filament protrusions, we conducted a photobleaching experiment combined with confocal imaging to determine where free actin monomers were added (Fig. 2a). Photobleached spots within protrusions showed little recovery (see Supplementary Information) as filament tips elongated (Fig. 2b), indicating monomer addition does not occur along the thin actin filament protrusions. These observations confirmed that protrusions incorporate new monomers only at their tips.
a, Laser scanning confocal images of fluorescence recovery of a photobleached region along a protrusion. The red arrow denotes the tip of the protrusion. Scale bar, 1 μm. b, Trajectories of positions along the photobleached protrusion. Blue circles mark the tip of the filopodium-like protrusion. Black circles (white on the image) mark the edge of the photobleached spot that is proximal to the tip. Red circles mark the edge of the photobleached spot that is distal to the tip.
The observed actin-based protrusions show a striking resemblance to cellular filopodia in two respects: (1) elongation of the actin filaments occurs at the tip of the protrusion, and (2) the protrusions lack the dendritic architecture of the networks from which they emerge14,15. However, unlike in vivo filopodia, the formation and stability of the thin actin filament protrusions from dendritic actin networks did not require bundling proteins or tip-complex proteins. In fact, the spontaneous initiation and growth of the protrusions in our experiments would be considered unlikely owing to dendritic actin network geometry and resistance of the membrane to deformation14. We propose that our observation of thin protrusion growth and stability in the absence of accessory proteins can be explained by considering the interplay between actin network and membrane mechanics.
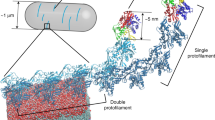
Using a mechanical model of actin-membrane configurations, we found that a deformable membrane can overcome the bending rigidity of actin filaments to both gather and bundle nearby filaments into a single tubular protrusion, even in the absence of tip-complex or bundling proteins. To illustrate how a deformable membrane can gather actin filaments, we computed the minimum energy configuration of an elastic sheet (representing the lipid bilayer) enveloping two semiflexible protrusions (Fig. 3a, see Supplementary Information for calculation details). In our calculations, the protruding filaments are anchored 100 nm (L0) below and are orthogonal to the unperturbed membrane. For this geometry, we determine the energetically optimal separation of the tips of filaments with contour length L and separation D at their bases. We find that, indeed, membrane elasticity is sufficient to bring together the tips of two nearby nascent protrusions. For a membrane tension of 0.005kBT nm−2 (ref. 16), the effective range of this membrane-mediated attraction extends beyond the filament thickness (∼10 nm) by only a few nanometres when the protrusion length is ∼80 nm, but the range of attraction grows rapidly with increasing protrusion length. Thus, if multiple filaments are gathered together by membrane-mediated attraction, their collective polymerization force can overcome the membrane’s resistance, resulting in formation of long tubular deformations.
a, The total energy associated with membrane deformation and filament bending is calculated for two actin filaments anchored 100 nm below the membrane with protrusion length L−L0 and separation D. The lightly shaded region under the curve represents the set of thermodynamically accessible states that will lead to filament bundling by the membrane. The darkly shaded region represents a subset of these states that are kinetically accessible, determined by the branching distance of ∼100 nm (ref. 1). Inset schematic diagrams represent the merged and unmerged states (not to scale). b, Stability of thin actin filament protrusions against Euler buckling. Left: A schematic diagram illustrates a straight protrusion, the total energy of which is due to the membrane tube. Right: A hypothetical situation in which the filaments buckle under the load of the membrane, where buckling is energetically unfavourable (see text). c, Model of membrane-induced formation of a thin actin filament protrusion. Left: Small-amplitude local deformations of the membrane arise as actin filaments polymerize against the membrane (black arrows). Deformations that are within the range of attraction are able to merge to create a larger deformation that gathers extra filaments (curly brackets). Middle: Deformations that fail to gather extra filaments are stalled and diminish (grey arrows). Right: After bundling enough filaments to overcome the membrane resistance to tube formation, a ‘proto-filopodium’ can elongate without further physical constraint.
The force required to initiate a tubular membrane protrusion exceeds the force resisting elongation, which is independent of tube length17. Once formed, the thin actin filament protrusion should therefore grow in contour length under a constant force from the membrane. However, the load at which a semiflexible filament (or a collection of parallel filaments) undergoes Euler buckling decreases steadily with increasing contour length. It is therefore not immediately clear how the long protrusion shown in the overlay of Fig. 1a, with a buckling threshold of the order of femtonewtons13, withstands a membrane load of tens of piconewtons.
The balance of forces that suggests buckling of long protrusions neglects the fact that a membrane tube must bend with the deflections of the filaments it encloses. Although the extra bending energy from the curvature of the membrane tube is relatively small17, the steric constraint that it must follow the curvature of the enclosed actin filaments has important repercussions. For the case of a straight protrusion, the membrane energy is Emem=fmemLmem, where fmem is the membrane resistance force and Lmem is the length of the membrane tube, and the bending energy of the filaments, Ebundle, is zero (Fig. 3b, left). For the case of a buckled protrusion, the initial stages of buckling would involve membrane configurations conformal to the enclosed filaments. Because the membrane bends and does not shorten in these configurations, Emem exceeds fmemLmem and Ebundle is positive (Fig. 3b, right). Thus, the total energy of the deformation is lower for a straight protrusion than for a buckled protrusion (see Supplementary Information and ref. 18).
The resulting stability against lateral deflections is consistent with our observation of long, straight actin filament protrusions as well as other long filopodia observed in vivo19,20. This stabilization of membrane-enclosed filaments against buckling has in fact been observed for microtubules protruding from the interior of a liposome21. Strikingly, these microtubules bend substantially inside the liposome body, but remain straight where surrounded by membrane tubes.
Our elastic model for membrane-induced bundling points to the protrusion length of individual actin filaments as a key parameter regulating the incidence of the filopodium-like protrusions. According to our calculations, increasing the protrusion length while holding membrane tension fixed should increase both the range and strength of lateral attraction between filaments. The maximum protrusion length is typically thought to be determined by a balance between polymerization force and membrane resistance (Fig. 3a). The force-limited length LprotF resulting from this balance, while limiting on average, could be transiently exceeded owing to fluctuations in the network’s internal stress. But it is also possible that single-filament protrusions rarely reach this limit before branching disrupts membrane configurations that favour bundling. In this case, the branch-limited maximum protrusion length LprotB of a single filament is determined instead by the relative kinetics of branching and polymerization.
If protrusion lengths are branch limited, our model would predict that shorter protrusion lengths created by increased branching would reduce the likelihood of the thin actin filament protrusions. Indeed, when Arp2/3 complex concentration was increased threefold while actin concentration was held constant, we observed a ∼60% reduction in the frequency of filopodium formation, consistent with expectations from the model and confirming that filament protrusion lengths are limited by kinetics rather than polymerization force. Conversely, the frequency of filopodium-like structures did not decrease when Arp2/3 complex concentration was decreased to as low as 10 nM (data not shown).
A second key parameter in the elastic model is membrane tension σ. Our calculations indicate that decreasing σ, while keeping LprotB fixed, should reduce the membrane’s ability to gather filaments. Consistent with this prediction, we found that decreasing membrane tension by increasing solution osmolarity, while keeping concentrations of Arp2/3 complex and free actin monomer fixed, indeed lowered the frequency of forming thin actin filament protrusions by ∼15% (see Supplementary Information).
On the basis of in vitro experiments and electron microscopy studies, Svitkina et al. proposed the convergent elongation model to describe the formation of filopodia from lamellipodia14. This model describes the transition from branched to parallel filament architecture in two stages: (1) elongation and lateral association of filaments by tip-complex proteins with anti-capping and barbed-end clustering activities, followed by (2) stabilization of the thin array of actin filaments by bundling proteins. Recent studies have identified extra filopodium regulators22,23, and other studies have challenged whether an underlying dendritic network is necessary for filopodium formation24,25. This debate has focused solely on the role of actin-binding proteins26, overlooking the question of whether and how the membrane could play a role in filament organization during filopodium formation. The unexpected formation and stability of the filopodium-like protrusions in our in vitro system suggests that parallel actin-based structures are a natural result of the interaction between growing actin networks and an elastic membrane. Our results demonstrate that the lipid bilayer alone can facilitate free barbed-end clustering and alignment of filaments that can continue to elongate without buckling under the membrane load.
Formation of filopodium-like structures without bundling proteins or tip-complex proteins suggests that membrane-induced alignment of actin filaments could facilitate formation of filopodia in cells (Fig. 3c). We propose a model in which actin filaments are routinely brought together by elastic interactions with the membrane, forming a ‘proto-filopodium’, which sets the stage for the formation of a mature filopodium through the participation of actin-binding proteins. Specifically, actin filaments initially gathered by the membrane would be stabilized by tip-complex proteins that associate with the filament ends and by bundling proteins such as fascin that reinforce parallel filament organization during membrane protrusion. In this way, accessory proteins exploit membrane-induced organization of actin filaments to provide regulatory control of protrusions but are not required for their initial formation. The minimal system for generating thin actin filament protrusions presented here would serve as an excellent platform for investigating and evaluating the role of various proteins required for filopodium formation in vivo; however, electron microscopy images of the actin filaments in these protrusions will be required to definitively confirm their organization. In a broader context, this work suggests that the mechanical properties of the plasma membrane may have a critical role in organizing the cytoskeletal filaments that drive many forms of cellular shape change.
Methods
Protein expression, purification and fluorescent labelling
Actin was purified from rabbit muscle acetone powder as described by Spudich and Watt27. Fluorescently labelled actin was prepared by reacting actin with AF350, AF488 or AF555 maleimide (Molecular Probes). His6-hemaglutinnin-tagged N-WASP (without Ena/VASP homology domain 1) was over-expressed in Escherichia coli and purified using nickel-affinity chromatography and an anion-exchange HiTRAP Q column (Amersham)28. Arp2/3 complex was purified from bovine brains29. AF488 N-WASP and AF488 Arp2/3 complex were labelled with AF488 maleimide. Protein purities were checked by sodium dodecyl sulphate polyacrylamide gel electrophoresis. Protein concentrations were determined by protein absorbance and bicinchoninic acid assays. N-WASP and Arp2/3 complex activities were confirmed by using in vitro bead motility assays with N-WASP-coated microspheres30. AF488 capping protein, Cdc42/Rho GDP dissociation inhibitor (RhoGDI), and intersectin DH-PH were expressed in E. coli and were gifts from Jack Taunton.
GUV formation
GUVs were prepared in sucrose solution from synthetic and natural lipids as described previously11. All lipids used were purchased from Avanti Polar Lipids unless otherwise stated. Three compositions were used primarily and all nucleated actin equally well. For canonical N-WASP activation with Cdc42 and PIP2, GUVs containing 75% egg phosphatidylcholine, 20% dioleoyl-phosphatidylserine and 5% brain PIP2 were used12. For some experiments, phase-separating GUVs containing 24.3% dioleoyl-phosphatidylcholine, 35% dipalmitoyl-phosphatidylcholine, 40% cholesterol and 0.7% BODIPY TMR-PIP2 were used where actin nucleation occurs robustly with BODIPY TMR-PIP2 alone (in the absence of Cdc42). DOGS-Ni-NTA-containing vesicles, which binds His-tagged N-WASP, have 95% egg phosphatidylcholine and 5% DOGS-Ni-NTA. Refer to Supplementary Information for the specific conditions used in each experiment. BODIPY TMR PIP2 was purchased from Echelon Biosciences. For experiments where visualization of membrane was desired, either fluorescent lipid fluorescein-DOPE or C5 C8 BODIPY was used.
In vitro reconstitution of thin actin filament protrusions
GUVs were pre-incubated with 4 μM of N-WASP for a minimum of 10 min before initiating the reaction. For experiments requiring Cdc42, 0.75 μM of Cdc42/RhoGDI complex, 0.18 μM DH-PH and 0.5 μM GTPγS were included during the pre-incubation step. Assembly of actin on GUVs was initiated by diluting the vesicle solution 10-fold in actin and Arp2/3 complex. The final mixture contains ∼8.5 μM G-actin, ∼160 nM Arp2/3 complex and ∼400 nM N-WASP in actin polymerization buffer. Osmolarity of the final protein mixtures was matched to the vesicles to within 5%. Large osmolarity differences are otherwise adjusted by varying the final salt concentration of the polymerization buffer (between 1× and 2×), which has no effect on the extent of actin polymerization. The protein vesicle mixture was added to a home-made chamber (0.1 mm×5 mm×20 mm) consisting of a glass coverslip adhered to a glass slide using double-sided tape, then sealed with 1:1:1 of vaseline/lanoline/paraffin before imaging.
Microscopy
Actin-associated vesicles were imaged by phase-contrast and epifluorescence microscopy using a ×100 1.25 numerical aperture objective for most studies on an inverted microscope (Zeiss Axiovert 200) equipped with motorized stage control and a back-illuminated cooled charge-coupled device (CCD) camera. All light-microscopy operations were driven by Metamorph imaging software. Laser scanning confocal microscopy was used for protein localization and photobleaching experiments (Zeiss LSM 510). A small region along a single proto-filopodium was bleached using a ×63 objective 1.25 numerical aperture with an 18 mW argon laser at 488 and 458 nm. Fluorescence recovery was monitored over 10 min. Spinning–disc confocal microscopy was also used to image actin networks on GUVs.
Data analysis and modelling
Line scans of AF555 actin, AF488 Arp2/3 complex and AF488 capping proteins along proto-filopodia were carried out in Metamorph. Raw fluorescence intensity was corrected for background fluorescence and normalized with respect to the base of proto-filopodia (dendritic networks) for actin. For Arp2/3 complex and capping proteins, the line scans were further divided by the intensity traces for actin to represent the relative localization of actin-binding proteins to actin.
For analysis of the photobleaching experiment, fluorescence intensity across the length of the proto-filopodium was tracked over time. Regions of the fluorescence line scan were fitted to a sigmoidal function. The inflection point is identified and used as a metric to track movements of the marked positions.
The growth rate of filopodium-like protrusions was measured from phase-contrast images. Integrated fluorescence actin intensity associated with the GUV was collected simultaneously with proto-filopodium elongation. After correcting for photobleaching and background fluorescence, the actin ‘polymerization curve’ was fitted to a sigmoid and time-dependent actin concentrations were interpolated (described in detail in Supplementary Information).
Image processing
All images were processed using Adobe Photoshop. Any brightness or contrast adjustments were uniformly applied to the entire image field and stack.
References
Pollard, T. D. & Borisy, G. G. Cellular motility driven by assembly and disassembly of actin filaments. Cell 112, 453–465 (2003).
Yeung, T., Ozdamar, B., Paroutis, P. & Grinstein, S. Lipid metabolism and dynamics during phagocytosis. Curr. Opin. Cell Biol. 18, 429–437 (2006).
Smythe, E. & Ayscough, K. R. Actin regulation in endocytosis. J. Cell Sci. 119, 4589–4598 (2006).
Eggert, U. S., Mitchison, T. J. & Field, C. M. Animal cytokinesis: From parts list to mechanisms. Annu. Rev. Biochem. 75, 543–566 (2006).
McMahon, H. T. & Gallop, J. L. Membrane curvature and mechanisms of dynamic cell membrane remodelling. Nature 438, 590–596 (2005).
Mullins, R. D., Heuser, J. A. & Pollard, T. D. The interaction of Arp2/3 complex with actin: Nucleation, high affinity pointed end capping, and formation of branching networks of filaments. Proc. Natl Acad. Sci. USA 95, 6181–6186 (1998).
Loisel, T. P., Boujemaa, R., Pantaloni, D. & Carlier, M. F. Reconstitution of actin-based motility of Listeria and Shigella using pure proteins. Nature 401, 613–616 (1999).
Bernheim-Groswasser, A., Wiesner, S., Golsteyn, R. M., Carlier, M. F. & Sykes, C. The dynamics of actin-based motility depend on surface parameters. Nature 417, 308–311 (2002).
Giardini, P. A., Fletcher, D. A. & Theriot, J. A. Compression forces generated by actin comet tails on lipid vesicles. Proc. Natl Acad. Sci. USA 100, 6493–6498 (2003).
Upadhyaya, A., Chabot, J. R., Andreeva, A., Samadani, A. & van Oudenaarden, A. Probing polymerization forces by using actin-propelled lipid vesicles. Proc. Natl Acad. Sci. USA 100, 4521–4526 (2003).
Liu, A. P. & Fletcher, D. A. Actin polymerization serves as a membrane domain switch in model lipid bilayers. Biophys. J. 91, 4064–4070 (2006).
Co, C., Wong, D. T., Gierke, S., Chang, V. & Taunton, J. Mechanism of actin network attachment to moving membranes: Barbed end capture by N-WASP WH2 domains. Cell 128, 901–913 (2007).
Mogilner, A. & Rubinstein, B. The physics of filopodial protrusion. Biophys. J. 89, 782–795 (2005).
Svitkina, T. M. et al. Mechanism of filopodia initiation by reorganization of a dendritic network. J. Cell Biol. 160, 409–421 (2003).
Faix, J. & Rottner, K. The making of filopodia. Curr. Opin. Cell Biol. 18, 18–25 (2006).
Atilgan, E., Wirtz, D. & Sun, S. X. Mechanics and dynamics of actin-driven thin membrane protrusions. Biophys. J. 90, 65–76 (2006).
Derenyi, I., Julicher, F. & Prost, J. Formation and interaction of membrane tubes. Phys. Rev. Lett. 88, 238101 (2002).
Pronk, S., Geissler, P. L. & Fletcher, D. A. Limits of filopodium stability. Phys. Rev. Lett. 100, 258102 (2008).
Wood, W. et al. Wound healing recapitulates morphogenesis in Drosophila embryos. Nature Cell Biol. 4, 907–912 (2002).
Onfelt, B. et al. Structurally distinct membrane nanotubes between human macrophages support long-distance vesicular traffic or surfing of bacteria. J. Immunol. 177, 8476–8483 (2006).
Fygenson, D. K., Elbaum, M., Shraiman, B. & Libchaber, A. Microtubules and vesicles under controlled tension. Phys. Rev. E 55, 850–859 (1997).
Sigal, Y. J., Quintero, O. A., Cheney, R. E. & Morris, A. J. Cdc42 and ARP2/3-independent regulation of filopodia by an integral membrane lipid-phosphatase-related protein. J. Cell Sci. 120, 340–352 (2007).
Mattila, P. K. et al. Missing-in-metastasis and IRSp53 deform PI(4,5)P2-rich membranes by an inverse BAR domain-like mechanism. J. Cell Biol. 176, 953–964 (2007).
Snapper, S. B. et al. N-WASP deficiency reveals distinct pathways for cell surface projections and microbial actin-based motility. Nature Cell Biol. 3, 897–904 (2001).
Steffen, A. et al. Filopodia formation in the absence of functional WAVE- and Arp2/3-complexes. Mol. Biol. Cell 17, 2581–2591 (2006).
Gupton, S. L. & Gertler, F. B. Filopodia: The fingers that do the walking. Sci. STKE 2007, re5 (2007).
Spudich, J. A. & Watt, S. The regulation of rabbit skeletal muscle contraction. I. Biochemical studies of the interaction of the tropomyosin-troponin complex with actin and the proteolytic fragments of myosin. J. Biol. Chem. 246, 4866–4871 (1971).
Papayannopoulos, V. et al. A polybasic motif allows N-WASP to act as a sensor of PIP(2) density. Mol. Cell 17, 181–191 (2005).
Egile, C. et al. Activation of the CDC42 effector N-WASP by the Shigella flexneri IcsA protein promotes actin nucleation by Arp2/3 complex and bacterial actin-based motility. J. Cell Biol. 146, 1319–1332 (1999).
Wiesner, S. et al. A biomimetic motility assay provides insight into the mechanism of actin-based motility. J. Cell Biol. 160, 387–398 (2003).
Acknowledgements
We thank D. Drubin and E. Nogales, as well as M. van Duijn, R. Rounsvell, S. Parekh and other members of the Fletcher laboratory for helpful discussions. We thank V. Risca for her help with confocal microscopy, and J. Taunton, D. Wong and C. Co for providing various proteins. We are grateful to C. Fromme for assisting us with fast protein liquid chromatography. This work is supported in part by a fellowship from the Natural Sciences and Engineering Research Council of Canada (A.P.L. and D.L.R.) and by funding from the NIH for the Cell Propulsion Laboratory as part of the Nanomedicine Roadmap Initiative (D.A.F.).
Author information
Authors and Affiliations
Corresponding author
Supplementary information
Supplementary Information
Supplementary Notes and Supplementary Figures 1–7 (PDF 306 kb)
Rights and permissions
About this article
Cite this article
Liu, A., Richmond, D., Maibaum, L. et al. Membrane-induced bundling of actin filaments. Nature Phys 4, 789–793 (2008). https://doi.org/10.1038/nphys1071
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nphys1071
This article is cited by
-
Reconstitution of contractile actomyosin rings in vesicles
Nature Communications (2021)
-
A self-organized synthetic morphogenic liposome responds with shape changes to local light cues
Nature Communications (2021)
-
Vesicle shape transformations driven by confined active filaments
Nature Communications (2021)
-
Actin crosslinker competition and sorting drive emergent GUV size-dependent actin network architecture
Communications Biology (2021)
-
A hierarchy of protein patterns robustly decodes cell shape information
Nature Physics (2021)