Abstract
Histamine, a neurotransmitter and neuroregulatory compound in diverse species1, serves as the neurotransmitter of photoreceptors in insects and other arthropods by directly activating a chloride channel2. By systematic expression screening of novel putative ligand-gated anion channels predicted from the Drosophila genome project, we identified two cDNAs (DM-HisCl-α1 and -α2) coding for putative histamine-gated chloride channels by functional expression in Xenopus laevis oocytes. DM-HisCl-α1 mRNA localizes in the lamina region of the Drosophila eye, supporting the idea that DM-HisCl-α1 may be a neurotransmitter receptor for histamine in the visual system.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Schwartz, J. C., Arrang, J. M., Garbarg, M., Pollard, H. & Ruat, M. Physiol. Rev. 71, 1–51 (1991).
Hardie, R. C. Nature 339, 704–706 (1989).
Ortells, M. O. Trends Neurosci. 18, 121–127 (1995).
Ffrench-Constant, R. H., Mortlock, D. P., Shaffer, C. D., MacIntyre, R. J. & Roush, R. T. Proc. Natl. Acad. Sci. USA 88, 7209–7213 (1991).
Cully, D. F., Paress, P. S., Liu, K. K., Schaeffer, J. M. & Arena, J.P. J. Biol. Chem. 271, 20187–20191 (1996).
Ranganathan, R., Cannon, S. C. & Horvitz, H. R. Nature 408, 470–475 (2000).
Littleton, J. T. & Ganetzky, B. Neuron 26, 35–43 (2000).
Wetzel, C. H., et al. J. Neurosci. 19, 7426–7433 (1999).
van der Goot, H. & Timmerman, H. Eur. J. Med. Chem. 35, 5–20 (2000).
Hardie, R. C. J. Exp. Biol. 138, 221–241 (1988).
Ache, B. W. & McClintock, T. S. Proc. Natl. Acad. Sci. USA 86, 8137–8141 (1989).
Hashemzadeh-Gargari, H. & Freschi, J. E. J. Neurophysiol. 68, 9–15 (1992).
Stuart, A. E. Neuron 22, 431–433 (1999).
Melzig, J., Burg, M., Gruhn, M., Pak, W. L. & Buchner, E. J. Neurosci. 18, 7160–7166 (1998).
Skingsley, D. R., Laughlin, S. B. & Hardie, R. C. J. Comp. Physiol. A 176, 611–623 (1995).
Acknowledgements
We thank S. Wagner, A. Stoeck, T. Sobik, M. Bathen and R. Kolarow for technical assistance, and B. Ache and K.F. Störtkuhl for comments on the manuscript.
Author information
Authors and Affiliations
Corresponding author
Supplementary information
Supplementary Figure 1.Amino acid alignment.
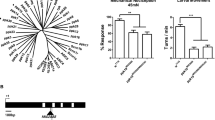
(a)Tree structure of putative ligand-gated ion channel proteins predicted by the data of the genome sequencing project (all accession numbers beginning with AAF) and already known channel proteins (Rdl: M69057, LCCH3: S62717, Grd: X78349, GluCl: AF297500). The tree was constructed by the program megalign of the Lasergene program package. (JPG 40 kb)
(b)
Amino-acid alignment with DM-HisCl-α1 (AF435469), DM-HisCl-α2 (AF435470), GABA receptor Rdl (M69057), glutamate receptor GluCl (AF297500), the C. elegans serotonin receptor MOD-1 (AF303088) and the murine glycine receptor α1 subunit (C49970). The putative DM-HisCl-α1 protein contains hydrophobic N-terminal residues with sequences highly predictive of a signal cleavage site (http:\\www.cbs.dtu.dk/services/SignalP) which would result in a mature protein beginning at amino-acid residue 22.The putative signal cleavage site in the DM-HisCl-α2 polypeptide is located at pos. 23 and 24. The DM-HisCl-α proteins contain consensus sites for protein kinase C and casein kinase II located between M3 and M4 as well as N-linked glycosylation sites in the proposed extracellular domain (http:\\www.expasy.ch/prosite/). The conserved motifs characteristic of the 'cys-brigde' family of channels, such as a large N-terminal extracellular domain and four hydrophobic transmembrane domains M1-M4, also occur in DM-HisCl-α. The main difference between DM-HisCl-α1 and DM-HisCl-α2 is the large intracellular loop between M3 and M4, which is considerably shorter in DM-HisCl-α2. The protein alignment shows that the DM-HisCl-α proteins are related to Drosophila glutamate- and GABA-gated chloride channels. Similarity to the Drosophila GABA- or glutamate-gated chloride channels Rdl {4942} and GluCl {1677} ranges from 19 to 27% of identical amino acids. Labels above the lines: putative signal sequence, M1-M4 regions, Cys-Cys bridge; ♦ N-glycosylation, ▪ PKC. (PDF 25 kb)
(c)
Alignment of the M2 regions of DM-HisCl-α1/-α2 with the ligand-gated chloride channels Rdl, GluCl, MOD-1 and Gly-alpha1 and the serotonin- and acetylcholine-gated cation channels GP-5HT3As (AF006462) and rat-nAChRαM (X03986). The putative pore-forming M2 regions of HisCl-α are more similar to the M2 regions of other ligand-gated chloride channels from Drosophlia, such as Rdl and GluCl, than to those of ligand-gated cation channels, suggesting that the DM-HisCl-α pore may also be chloride-selective. (PDF 10 kb)
Rights and permissions
About this article
Cite this article
Gisselmann, G., Pusch, H., Hovemann, B. et al. Two cDNAs coding for histamine-gated ion channels in D. melanogaster. Nat Neurosci 5, 11–12 (2002). https://doi.org/10.1038/nn787
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nn787
This article is cited by
-
Alkaline taste sensation through the alkaliphile chloride channel in Drosophila
Nature Metabolism (2023)
-
A single photoreceptor splits perception and entrainment by cotransmission
Nature (2023)
-
Blood-feeding adaptations and virome assessment of the poultry red mite Dermanyssus gallinae guided by RNA-seq
Communications Biology (2023)
-
In silico identification and assessment of insecticide target sites in the genome of the small hive beetle, Aethina tumida
BMC Genomics (2020)
-
The HisCl1 histamine receptor acts in photoreceptors to synchronize Drosophila behavioral rhythms with light-dark cycles
Nature Communications (2019)