Abstract
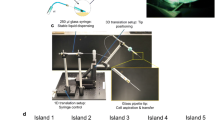
We present a sectioning and bonding technology to make ultrahigh density microarrays of solid samples, cutting edge matrix assembly (CEMA). Maximized array density is achieved by a scaffold-free, self-supporting construction with rectangular array features that are incrementally scalable. This platform technology facilitates arrays of >10,000 tissue features on a standard glass slide, inclusion of unique sample identifiers for improved manual or automated tracking, and oriented arraying of stratified or polarized samples.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Kononen, J. et al. Nat. Med. 4, 844–847 (1998).
Sauter, G., Simon, R. & Hillan, K. Nat. Rev. Drug Discov. 2, 962–972 (2003).
Perrone, E.E. et al. J. Natl. Cancer Inst. 92, 937–939 (2000).
Camp, R.L., Dolled-Filhart, M., King, B.L. & Rimm, D.L. Cancer Res. 63, 1445–1448 (2003).
Nevalainen, M.T. et al. J. Clin. Oncol. 22, 2053–2060 (2004).
Galil, K.A., Schofield, I.D. & Wright, G.Z. J. Biomed. Mater. Res. 18, 609–616 (1984).
Grimley, P.M., Dong, F. & Rui, H. Cytokine Growth Factor Rev. 10, 131–157 (1999).
Acknowledgements
Animal studies were approved by Georgetown University Animal Care and Use Committee. Supported by US National Institutes of Health grants R01-DK52013 and R01-CA101841 (to H.R.) and T32-CA009686-08 (to M.J.L.). We thank Lombardi Comprehensive Cancer Center shared resources of Histopathology, Microscopy and Imaging, and Tissue Culture (supported in part by NIH 1P30-CA-51008), and K. Catlin-LeBaron for help with graphic design.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
Intellectual property arising from this work belongs to Georgetown University (GU). A patent application, which covers the work described in this manuscript, naming H.R and M.J.L. as inventors has been filed and is managed by GU Office of Technology Licensing according to institutional guidelines.
Supplementary information
Supplementary Fig. 1
Ultrahigh density microarraying of solid samples using CEMA. (PDF 761 kb)
Supplementary Fig. 2
CEMA arrays of stratified tissues and 3D cell culture samples. (PDF 1453 kb)
Supplementary Fig. 3
Frozen tissue array of skeletal muscle. (PDF 496 kb)
Rights and permissions
About this article
Cite this article
LeBaron, M., Crismon, H., Utama, F. et al. Ultrahigh density microarrays of solid samples. Nat Methods 2, 511–513 (2005). https://doi.org/10.1038/nmeth772
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth772
This article is cited by
-
Prolactin-Stat5 signaling in breast cancer is potently disrupted by acidosis within the tumor microenvironment
Breast Cancer Research (2013)
-
Low levels of Stat5a protein in breast cancer are associated with tumor progression and unfavorable clinical outcomes
Breast Cancer Research (2012)
-
Differential expression of arrestins is a predictor of breast cancer progression and survival
Breast Cancer Research and Treatment (2011)
-
The adult human brain in preclinical drug development
Nature Reviews Drug Discovery (2008)
-
The construction of high-density paraffin tissue microarrays with 0.43-mm-diameter paraffin tissue core biopsies is technically feasible
Virchows Archiv (2008)