Abstract
The continuous evolution and adaptation of HIV to its host has produced extensive global viral diversity. Understanding the kinetics and directions of this continuing adaptation and its impact on viral fitness, immunogenicity and pathogenicity will be crucial to the successful design of effective HIV vaccines. Here we discuss some potential scenarios of viral and host coevolution.
Similar content being viewed by others
Main
Since its introduction into human hosts, HIV has evolved into numerous distinct and diverse subtypes. The inherent infidelity of the viral reverse transcriptase, together with the absence of adequate repair mechanisms, favors generation of viral variants that adapt to environmental conditions, particularly virus-specific immune responses. Here we discuss emerging evidence suggesting that HIV has adapted to cellular and humoral immune surveillance in its human host and that immune surveillance has left population-dependent genetic imprints on the viral genome. As a consequence of this adaptation, HIV could become less immunogenic as the epidemic progresses, and continued evolution in the human host could result in either a more or less pathogenic virus in the future. These possible scenarios have significant implications in the quest for an effective HIV vaccine.
Viral diversity and AIDS virus infection in monkeys and humans
Phylogenetic and epidemiological studies suggest that HIV-1 and HIV-2 were introduced to the human population of West Africa around 1930 by cross-species transmissions from simian immunodeficiency virus (SIV)cpz-infected chimpanzees and SIVsmg-infected sooty mangabey monkeys, respectively1. None of the natural primate hosts of SIV show severe clinical symptoms upon SIV infection, but experimental SIV infection of captive macaques is characterized by a rapid and fatal outcome2,3. Because macaques acquired SIV only recently through experimental cross-species transmission of SIVsmg, they cannot be considered natural hosts for SIV4. This brings up the question of whether SIV (and HIV) will eventually adapt to more recent hosts, developing into infectious agents with reduced morbidity and mortality. This is likely to be accompanied by reduced immune reactivity and a reduced rate of adaptation to weaker immune pressure5. This has been observed in a comparison of natural hosts and rhesus macaques infected with SIVsmg, where the natural host showed a lower ratio of nonsynonymous compared with synonymous mutations, versus the non-natural host6. This strongly suggests that SIVsmg and sooty mangabeys have reached a balance between viral replication, intraspecies transmission efficiency and host health and survival.
Whether the introduction of SIV and HIV into the human population will lead to a similar adaptation, and if so, how long this process will take, is an open question. The potential for continuing HIV diversification still exists, although chemical mutagenesis studies indicate that mutation rates may be limited to prevent the virus from committing 'mutational suicide'7,8. The fact that only a small fraction of progenitor virus is replication-competent indicates that HIV may be close to its mutational limits9. Thus, in order to maintain overall viral fitness and to ensure efficient replication and transmission, adaptation must be limited by structural constraints. Mutations may only reach fixation if they provide superior advantages against other, balancing selection pressures, including those mediated by virus-specific cytotoxic T lymphocytes (CTLs).
Immune escape: HIV CTL epitope evolution over time
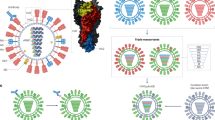
Although the subject has been controversial, recent data provide strong evidence of clinically relevant immune escape of HIV and SIV from virus-specific T-cell responses. Full-genome sequence analysis has shown the potential for sequence variability at almost every nucleotide in the HIV genome. This variability is observed, although to different degrees, in all optimal HIV CTL epitopes identified to date12,13,14. Epitope variation can often lead to reduced immunogenicity and impaired recognition by CTLs, especially when anchor positions required for high-affinity binding to human leukocyte antigen (HLA) class I molecules are substituted with residues that no longer confer strong binding10,15 (Fig. 1). One could therefore expect HIV to show a preference for mutation in these positions, but recent data from our lab suggest that this may not be apparent when one examines defined epitopes on a population basis.
Mutations within a CTL epitope can lead to inability to bind to the class I allele. Progressive accumulation and transmission of such mutations within viral peptides presented to the immune system may lead to evolution of viruses that are less well recognized. Alternatively, such mutations may make the virus less pathogenic by altering the replicative capacity of the virus.
We analyzed sequence variability in 44 well-described Gag CTL epitopes (typically 9–10 residues), from 20 individuals acutely infected with currently circulating viruses in the Boston, Massachusetts area. We did not observe a preference for changes affecting anchor positions compared with changes in other residues (unpublished observations). Although escape from antigen processing and T-cell receptor recognition may represent other important mechanisms for CTL escape, the above data suggest that HIV may have evolved away from many responses in the past by mutating in positions 'allowed' by viral structural constraints. This may have left us with epitopes that, due to the same functional constraints, can not escape. This interpretation is consistent with other data suggesting that epitopes restricted by common HLA class I alleles have been widely eliminated because population-wide selection pressure prevents reversion of escape mutations16,17,18. Other studies have shown that poorly targeted regions of the viral genome have an increased frequency of nonanchor residues, which again suggests that the targeted regions cannot escape easily and that extensive escape may have occurred in the nontargeted regions of the genome19,20. Alternatively, the epitopes may be targeted by functionally impaired CTLs, so there may be incentive for the virus to change in those regions. If this is the case, the 'common' epitopes identified to date may largely reflect ineffective epitopes, which may be less relevant for vaccine design. These observations and other recent data highlight the urgency to define the total CTL repertoire against HIV by comprehensive screenings, based wherever possible on autologous virus sequences21.
Insights from characterization of common HIV CTL epitopes?
The HIV immunology database at the Los Alamos National Laboratories provides annual updates of well-defined CTL epitopes and their restricting HLA alleles22. This selective list includes nearly 200 epitopes, restricted by >55 HLA class I alleles, and has often been the basis for the design of subunit vaccines and other immune studies13. However, most epitopes identified in the past have been found using antigen sources based on selected reference strains of the virus. Thus, these can differ significantly from the autologous virus and bias the identification of epitopes towards relatively conserved regions of the viral genome12,19,21,23. Although many current studies use consensus sequences built from alignments that include contemporary strains, there is still an almost unavoidable bias towards the detection of relatively conserved CTL epitopes12,23. Aside from preferences due to proteasomal antigen processing, epitope binding to HLA molecules and transporter associated with antigen processing (TAP) translocation efficacy, differences between autologous and test sequences may be a major explanation for the observed epitope clustering in certain regions of the viral protein sequences19,23,24. Together with natural limitations in the sensitivity of different assay systems, these studies suggest that identification of CTL epitopes may be heavily biased towards (i) epitopes located in more conserved regions, (ii) epitopes with conserved anchor residues and (iii) epitopes with high binding affinities for the restricting HLA molecules. In theory, these characteristics are welcome when considering vaccine candidates. However, the general inability of well-characterized CTL responses to control viral replication in vivo, the conflicting data correlating the breadth and strength of the total CTL responses with viral load, and the fact that CTL control and escape has only been shown for isolated individuals and certain epitopes suggest that not all CTL responses described to date contribute equally to virus control17,25,26,27,28.
Rapid CTL escape in SIV and HIV
Elegant studies have shown that rapid CTL-associated evolution of mutations leading to escape is a hallmark of acute SIV infection11,29. Although similar reports of CTL escape in acute HIV infection have arisen, the clinical consequences of this escape have been less clear30,31. This may reflect features of the SIV infection model, but aspects of host adaptation must be considered as well. The viruses used in the SIV model (often SIVMAC239 or SIVMAC251) are evolutionarily relatively new to the experimental host species (macaques), as they were derived from a strain isolated from naturally infected sooty mangabeys and have been passaged only once (SIVMAC251) or twice (SIVMAC239) in macaques, where the bulk of in vivo experiments are conducted29,32. Thus, the macaques are encountering a virus that has not been under prolonged immune selection pressure by macaque MHC-restricted CTLs, and are thus more likely to have retained epitopes in the 'wild-type' sequence. As a consequence, the virus may induce strong responses against multiple, still 'escapable' epitopes in SIV, providing us with a chance to observe acute CTL escape.
The situation in humans is quite different, particularly in geographically distinct populations with limited HLA diversity, where CTL responses will imprint their profile of immune selection pressure on the viral sequences10,16,33. However, if population-adapted HIV is transmitted into an individual of a significantly different genetic background, the previous CTL escape mutations will not provide an advantage to this virus. Thus, the 'foreign' individual (relative to the evolution of the virus) may mount more and qualitatively different CTL responses compared with the virus donor. Such 'xenoinfection' may actually have a better outcome than infections with a virus that has been passaged within a confined, genetically homogeneous population. There have been anecdotal data in humans to support this hypothesis: a few individuals in Europe infected with non-B-clade viruses from Africa fared surprisingly well and controlled the infection without the need for antiretroviral treatment (E. Bernasconi, personal communication; Lugano, Switzerland).
A recent report by Trachtenberg et al.34 provides more solid evidence for this observation. The authors hypothesized that transmission of CTL escape variants would put people expressing the most common HLA supertypes (groups of class I HLA-A and HLA-B alleles) at a disadvantage. Their report shows that HLA supertype frequency correlated with viral load, representing a 'rare-allele advantage' for people of African descent in the mostly white population of the Multicenter AIDS Cohort Study. The data may suggest that the virus circulating in the mostly white population early in the American epidemic was more effectively controlled in individuals of African descent. This may be because of conservation of specific CTL epitopes presented by HLA alleles relatively unique to African ethnicities. Because the epidemic may also impact human evolution and lead to shifts in HLA allele frequencies over time (by selecting for HLA class I alleles associated with slow disease progression), a specific rare-allele advantage may only be effective as long as the allele remains rare. Once an allele has provided a population-wide selective advantage, it will be too frequent and the virus will show host population imprinting in the targeted region. Similarly, if the epidemic spreads into additional ethnicities, rare-allele advantages may be lost after prolonged circulation. Furthermore, the current classification of HLA supertypes does not include HLA-C alleles, and cross-presentation of CTL epitopes among the alleles forming specific supertypes needs to be addressed in more detail. However, the data of Trachtenberg et al. indicate that transmission of CTL escape variants and gradual loss of effective CTL epitopes may have occurred in the past and will impact the immunogenicity of HIV in the future.
A recent study of HIV transmission and CTL escape in mother-child pairs also supports this scenario10. If an infant inherited the HLA class I allele that restricted an escaped CTL specificity in the mother, the virus did not revert back to wild type and the infant was unable to mount the CTL response. Thus, although an adapted (potentially less fit) virus was transmitted, the reduced immunogenicity could prevail and cause accelerated disease progression. These effects may be even more marked when HLA concordance between mother and child is more pronounced. However, increased HLA concordance also automatically increases the degree of HLA homogeneity in the infant, which is associated per se with faster disease progression, thus complicating the analyses35,36. The situation is also complicated by the infants' immature immune systems, which could further accelerate disease progression37. Thus, studies of HIV transmission between genetically-related adults will be required to assess this question satisfactorily and show that HLA allele frequency in a population is an important factor for viral escape mutations to reach fixation.
Our analyses of HLA-A3–restricted CTL epitopes strongly suggest that large proportions of the circulating viruses in the United States have mutations in the anchor positions of these epitopes10. When transmitted to an HLA-A3–negative host, the virus could potentially revert back to a more fit wild-type virus. However, this will be complicated when compensatory changes need to revert back to the wild-type. In addition, the infecting virus may be less fit at the time of transmission and, because it offers its full immunogenicity to the genetically unrelated host, may be controlled more effectively. These considerations also suggest that as the HIV epidemic continues, acute CTL escape may be restricted to epitopes presented by rare HLA alleles or epitopes that readily revert back to wild-type probably because of high fitness constraints associated with the escape mutation. Accordingly, the few instances in which acute CTL escape has been shown30,31 may represent cases where a relatively unadapted virus was transmitted and a rare HLA allele advantage was effective. In all cases where the restricting HLA class I allele for the escaped CTL response was properly determined, the frequency of the restricting HLA class I alleles in the local population (HLA-A29, HLA-A68, HLA-B8, HLA-B14, HLA-B44, HLA-B57 and HLA-C12) was low. This would give the virus ample opportunity to be passaged in individuals not expressing these alleles and to revert the epitope back to wild type. Such reversion would occur more frequently the more rare the restricting HLA allele is in a given population and the stronger the structural fitness constraints are. There is a need for more detailed analyses of acute CTL escape and broader evolution studies of viruses introduced into ethnicities to which they have not been previously adapted. It is intriguing that no acute CTL escape has been observed for some HLA alleles such as HLA-A*0201 and other frequently studied alleles. This may have to do with the specific HLA allele (which may not generate high-affinity responses) or could indicate that HLA-A*0201 restricted epitopes may have been deleted shortly after entering the HLA-A*0201-rich population that was studied16,18.
Does adaptation impact viral fitness?
In addition to the implications for vaccine development and in vitro immune studies, the above considerations have bearing on an important question: will continuous adaptation to the human host reduce HIV immunogenicity and, as a consequence, will this impair viral replication fitness and lead to a virus with markedly reduced pathogenicity? There is some indirect evidence affirming this hypothesis, stemming from viral fitness measurements of drug-resistant HIV and SIV. The data suggest that adaptation to drug selection pressure is not free of costs to the virus, and that the virus actively seeks to overcome this handicap by generating compensatory mutations38. This has been described for CTL-mediated selection pressure as well: escape from the immunodominant HLA-B27 epitope in HIV Gag includes complex events such as compensatory changes within and outside the targeted epitope. This suggests that the virus tries to compensate for losses in viral fitness as a consequence of CTL escape15. Well-controlled viral fitness analyses are clearly needed to compare contemporary and older viral isolates, as well as isolates with single point mutations and compensatory changes introduced by site-directed mutagenesis, for their replication competence
Conclusions
Emerging data suggest that HIV and SIV have adapted to their animal hosts since their introduction into these species, and are likely to continue eliminating important targets of the host cellular immune response. Finding genetic traces of these past responses to HIV infection may contribute to the identification of true immune correlates of controlled HIV infection today. Because such viral adaptation to the human host may take its progressive toll on the virus, reduced immunogenicity may be accompanied by reduced viral fitness. By inference from other host-pathogen models, including SIV, there are reasons to believe that such adaptation may be associated with reduced pathogenicity. The best arguments for this are SIV infections in their respective natural hosts. Although there are some profound differences between HIV transmission and experimental SIV models, the latter can teach us what we might learn if we could test individuals infected early in the epidemic and compare their responses to autologous virus sequences with responses of individuals infected today. Whether these approaches will indeed support the scenario drawn here needs to be determined before one can predict to which side the balance between immunogenicity/escape and adaptation/viral fitness will swing.
References
Gao, F. et al. Origin of HIV-1 in the chimpanzee Pan troglodytes troglodytes. Nature 397, 436–441 (1999).
Norley, S., Beer, B., Holzammer, S., zur Megede, J. & Kurth, R. Why are the natural hosts of SIV resistant to AIDS? Immunol. Lett. 66, 47–52 (1999).
Rey-Cuille, M.A. et al. Simian immunodeficiency virus replicates to high levels in sooty mangabeys without inducing disease. J. Virol. 72, 3872–3886 (1998).
Holmes, E.C. On the origin and evolution of the human immunodeficiency virus (HIV). Biol. Rev. Camb. Philos. Soc. 76, 239–254 (2001).
Kaur, A. et al. Emergence of cytotoxic T lymphocyte escape mutations in nonpathogenic simian immunodeficiency virus infection. Eur. J. Immunol. 31, 3207–3217 (2001).
Courgnaud, V., Saurin, W., Villinger, F. & Sonigo, P. Different evolution of simian immunodeficiency virus in a natural host and a new host. Virology 247, 41–50 (1998).
Pelletier, E. & Wain, H.S. AIDS is not caused by the extreme genetic variability of HIV. J. NIH Res. 8, 45–49 (1996).
Loeb, L.A. et al. Lethal mutagenesis of HIV with mutagenic nucleoside analogs. Proc. Natl. Acad. Sci. USA 96, 1492–1497 (1999).
Browning, M.J., Huang, A.S. & Reiss, C.S. Cytolytic T lymphocytes from the BALB/c-H-2dm2 mouse recognize the vesicular stomatitis virus glycoprotein and are restricted by class II MHC antigens. J. Immunol. 145, 985–994 (1990).
Goulder, P.J. et al. Evolution and transmission of stable CTL escape mutations in HIV infection. Nature 412, 334–338 (2001).
Allen, T.M. et al. Tat-specific cytotoxic T lymphocytes select for SIV escape variants during resolution of primary viraemia. Nature 407, 386–390 (2000).
Gaschen, B. et al. Diversity considerations in HIV-1 vaccine selection. Science 296, 2354–2360 (2002).
Frahm, N., Goulder, P. & Brander, C. Total assessment of HIV specific CTL responses: epitope clustering, processing preferences and the impact of HIV sequence heterogeneity. in HIV Molecular Immunology Database 2002 (eds. Korber, B. et al.) I.1–I.22 Theoretical Biology and Biophysics, Los Alamos National Laboratory, (2002).
Korber, B. et al. HIV Molecular Immunology Database 2001, (Eds. Korber, B et al.) II.A.1–I.A.271 (Theoretical Biology and Biophysics Group, Los Alamas National Laboratory, 2001).
Kelleher, A.D. et al. Clustered mutations in HIV-1 Gag are consistently required for escape from HLA-B27 restricted CTL responses. J. Exp. Med. 193, 375–386 (2001).
Moore, C.B. et al. Evidence of HIV-1 adaptation to HLA-restricted immune responses at a population level. Science 296, 1439–1443 (2002).
Brander, C. et al. Lack of strong immune selection pressure by the immunodominant, HLA-A*0201 restricted CTL response in chronic HIV-1 infection. J. Clin. Invest. 101, 2559–2566 (1998).
Shankar, P., Fabry, J.A., Fong, D.M. & Lieberman, J. Three regions of HIV-1 gp160 contain clusters of immunodominant CTL epitopes. Immunol. Lett. 52, 23–30 (1996).
Yusim, K. et al. Clustering patterns of cytotoxic T-lymphocyte epitopes in human immunodeficiency virus type 1 (HIV-1) proteins reveal imprints of immune evasion on HIV-1 global variation. J. Virol. 76, 8757–8768 (2002).
Calef, C. et al. PeptGen: designing peptides for immunological studies and application to HIV consensus sequences. in HIV Molecular Immunology Database 2000 (eds. Korber, B. et al. I.63–I.100) (Theoretical Biology and Biophysics, Los Alamos National Laboratory, 2000).
Altfeld, M. et al. Enhanced detection of HIV-1-specific T cell responses to highly variable regions using peptides based on autologous virus sequences. J. Virol. 77, 7330–7340 (2003).
Brander, C. & Goulder, P. The identification of optimal HIV derived CTL epitopes in diverse populations using HIV clade specific consensus. in HIV Molecular Immunology Database 2001 (eds. Korber, B. et al.) I.1–I.20 (Theoretical Biology and Biophysics Group, Los Alamos National Laboratory, 2001).
Draenert, R. et al. Comparison of overlapping peptide sets for detection of antiviral CD4 and CD8 T cell responses. J. Immunol. Methods 275, 19–29 (2003).
Goulder, P.J. et al. Differential narrow focusing of immunodominant human immunodeficiency virus gag-specific cytotoxic T-lymphocyte responses in infected african and caucasoid adults and children. J. Virol. 74, 5679–5690 (2000).
Betts, M.R. et al. Analysis of total human immunodeficiency virus (HIV)-specific CD4(+) and CD8(+) T-cell responses: relationship to viral load in untreated HIV infection. J. Virol. 75, 11983–11991 (2001).
Goulder, P.J. et al. Functionally inert HIV-specific cytotoxic T lymphocytes do not play a major role in chronically infected adults and children. J. Exp. Med. 192, 1819–1832 (2000).
Migueles, S.A. et al. HLA B*5701 is highly associated with restriction of virus replication in a subgroup of HIV-infected long term nonprogressors. Proc. Natl. Acad. Sci. USA 97, 2709–2714 (2000).
Addo, M. et al. Comprehensive epitope analysis of HIV-1-specific T cell responses directed against the entire expressed HIV-1 genome demonstrate broadly directed responses, but no correlation to viral load. J. Virol. 77, 2081–2092 (2003).
O'Connor, D.H., Allen, T.M. & Watkins, D.I. Where have all the monkeys gone? Evaluating SIV-specific CTL in the post-Mamu-A*01 era., (eds. Korber, B. et al.) I.26–I39 (Theoretical Biology and Biophysics Group, Los Alamos National Laboratory, 2001).
Borrow, P. et al. Antiviral pressure exerted by HIV-1-specific cytotoxic T lymphocytes (CTLs) during primary infection demonstrated by rapid selection of CTL escape virus. Nat. Med. 3, 205–211 (1997).
Price, D.A. et al. Positive selection of HIV-1 cytotoxic T lymphocyte escape variants during primary infection. Proc. Nat. Acad. Sci. USA 94, 1890–1895 (1997).
Desrosiers, R.C. & Ringler, D.J. Use of simian immunodeficiency viruses for AIDS research. Intervirology 30, 301–312 (1989).
de Campos-Lima, P.O. et al. HLA-A11 epitope loss isolates of Epstein-Barr virus from a highly A11+ population. Science 260, 98–100 (1993).
Trachtenberg, E. et al. Advantage of rare HLA supertype in HIV disease progression. Nat. Med. 9, 928–993 (2003).
Carrington, M. et al. HLA and HIV-1: heterozygote advantage and B*35-Cw04 disadvantage. Science 283, 1748–1752 (1999).
MacDonald, K.S. et al. Mother-child class I HLA concordance increases perinatal human immunodeficiency virus type 1 transmission. J. Infect. Dis. 177, 551–556 (1998).
Brander, C. et al. Persistent HIV-1 specific CTL clonal expansion despite high viral burden post in utero HIV-1 infection. J. Immunol. 162, 4796–4800 (1999).
Molla, A. et al. Ordered accumulation of mutations in HIV protease confers resistance to ritonavir. Nat. Med. 2, 760–766 (1996).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Brander, C., Walker, B. Gradual adaptation of HIV to human host populations: good or bad news?. Nat Med 9, 1359–1362 (2003). https://doi.org/10.1038/nm941
Published:
Issue Date:
DOI: https://doi.org/10.1038/nm941
This article is cited by
-
Therapeutic immunization for HIV
Springer Seminars in Immunopathology (2006)
-
Control of human immunodeficiency virus replication by cytotoxic T lymphocytes targeting subdominant epitopes
Nature Immunology (2006)
-
Conflicting selective forces affect T cell receptor contacts in an immunodominant human immunodeficiency virus epitope
Nature Immunology (2006)