Abstract
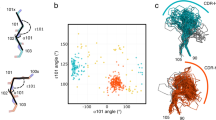
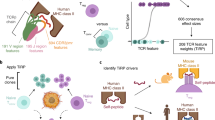
All complexes of T cell receptors (TCRs) bound to peptide–major histocompatibility complex (pMHC) molecules assume a stereotyped binding 'polarity', despite wide variations in TCR-pMHC docking angles. However, existing TCR-pMHC crystal structures have failed to show broadly conserved pairwise interaction motifs. Here we determined the crystal structures of two TCRs encoded by the variable β-chain 8.2 (Vβ8.2), each bound to the MHC class II molecule I-Au, and did energetic mapping of Vα and Vβ contacts with I-Au. Together with two previously solved structures of Vβ8.2-containing TCR-MHC complexes, we found four TCR–I-A complexes with structurally superimposable interactions between the Vβ loops and the I-A α-helix. This examination of a narrow 'slice' of the TCR-MHC repertoire demonstrates what is probably one of many germline-derived TCR-MHC interaction 'codons'.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
References
Rudolph, M.G., Stanfield, R.L. & Wilson, I.A. How TCRs bind MHCs, peptides, and coreceptors. Annu. Rev. Immunol. 24, 419–466 (2006).
Buslepp, J., Wang, H., Biddison, W.E., Appella, E. & Collins, E.J. A correlation between TCR Vα docking on MHC and CD8 dependence: implications for T cell selection. Immunity 19, 595–606 (2003).
Jerne, N.K. The somatic generation of immune recognition. Eur. J. Immunol. 1, 1–9 (1971).
Blackman, M. et al. The T cell repertoire may be biased in favor of MHC recognition. Cell 47, 349–357 (1986).
Zerrahn, J., Held, W. & Raulet, D.H. The MHC reactivity of the T cell repertoire prior to positive and negative selection. Cell 88, 627–636 (1997).
Matsui, K. et al. Low affinity interaction of peptide-MHC complexes with T cell receptors. Science 254, 1788–1791 (1991).
Krogsgaard, M. & Davis, M.M. How T cells 'see' antigen. Nat. Immunol. 6, 239–245 (2005).
Wu, L.C., Tuot, D.S., Lyons, D.S., Garcia, K.C. & Davis, M.M. Two-step binding mechanism for T-cell receptor recognition of peptide MHC. Nature 418, 552–556 (2002).
Huseby, E.S. et al. How the T cell repertoire becomes peptide and MHC specific. Cell 122, 247–260 (2005).
Huseby, E.S., Crawford, F., White, J., Marrack, P. & Kappler, J.W. Interface-disrupting amino acids establish specificity between T cell receptors and complexes of major histocompatibility complex and peptide. Nat. Immunol. 7, 1191–1199 (2006).
Zamvil, S.S. & Steinman, L. The T lymphocyte in experimental allergic encephalomyelitis. Annu. Rev. Immunol. 8, 579–621 (1990).
Zamvil, S.S. et al. T cell specificity for class II (I-A) and the encephalitogenic N- terminal epitope of the autoantigen myelin basic protein. J. Immunol. 139, 1075–1079 (1987).
Goverman, J. Tolerance and autoimmunity in TCR transgenic mice specific for myelin basic protein. Immunol. Rev. 169, 147–159 (1999).
Acha-Orbea, H. et al. Limited heterogeneity of T cell receptors from lymphocytes mediating autoimmune encephalomyelitis allows specific immune intervention. Cell 54, 263–273 (1988).
Urban, J.L. et al. Restricted use of T cell receptor V genes in murine autoimmune encephalomyelitis raises possibilities for antibody therapy. Cell 54, 577–592 (1988).
Maynard, J. et al. Structure of an autoimmune T cell receptor complexed with class II peptide-MHC: insights into MHC bias and antigen specificity. Immunity 22, 81–92 (2005).
Reinherz, E.L. et al. The crystal structure of a T cell receptor in complex with peptide and MHC class II. Science 286, 1913–1921 (1999).
Pearson, C.I., van Ewijk, W. & McDevitt, H.O. Induction of apoptosis and T helper 2 (Th2) responses correlates with peptide affinity for the major histocompatibility complex in self- reactive T cell receptor transgenic mice. J. Exp. Med. 185, 583–599 (1997).
Lafaille, J.J., Nagashima, K., Katsuki, M. & Tonegawa, S. High incidence of spontaneous autoimmune encephalomyelitis in immunodeficient anti-myelin basic protein T cell receptor transgenic mice. Cell 78, 399–408 (1994).
Goverman, J. et al. Transgenic mice that express a myelin basic protein-specific T cell receptor develop spontaneous autoimmunity. Cell 72, 551–560 (1993).
He, X.L. et al. Structural snapshot of aberrant antigen presentation linked to autoimmunity: the immunodominant epitope of MBP complexed with I-Au. Immunity 17, 83–94 (2002).
Maynard, J. et al. High-level bacterial secretion of single-chain αβ T-cell receptors. J. Immunol. Methods 306, 51–67 (2005).
Garcia, K.C. et al. Structural basis of plasticity in T cell receptor recognition of a self peptide-MHC antigen. Science 279, 1166–1172 (1998).
Garcia, K.C., Radu, C.G., Ho, J., Ober, R.J. & Ward, E.S. Kinetics and thermodynamics of T cell receptor-autoantigen interactions in murine experimental autoimmune encephalomyelitis. Proc. Natl. Acad. Sci. USA 98, 6818–6823 (2001).
Cunningham, B.C. & Wells, J.A. High-resolution epitope mapping of hGH-receptor interactions by alanine-scanning mutagenesis. Science 244, 1081–1085 (1989).
Manning, T.C. et al. Alanine scanning mutagenesis of an αβ T cell receptor: mapping the energy of antigen recognition. Immunity 8, 413–425 (1998).
Lee, P.U., Churchill, H.R., Daniels, M., Jameson, S.C. & Kranz, D.M. Role of 2CT cell receptor residues in the binding of self- and allo-major histocompatibility complexes. J. Exp. Med. 191, 1355–1364 (2000).
Baker, B.M., Turner, R.V., Gagnon, S.J., Wiley, D.C. & Biddison, W.E. Identification of a crucial energetic footprint on the alpha1 helix of human histocompatibility leukocyte antigen (HLA)-A2 that provides functional interactions for recognition by tax peptide/HLA-A2-specific T cell receptors. J. Exp. Med. 193, 551–562 (2001).
Borg, N.A. et al. The CDR3 regions of an immunodominant T cell receptor dictate the 'energetic landscape' of peptide-MHC recognition. Nat. Immunol. 6, 171–180 (2005).
Colf, L.A. et al. How a single T cell receptor recognizes both self and foreign MHC. Cell 129, 135–146 (2007).
Hennecke, J., Carfi, A. & Wiley, D.C. Structure of a covalently stabilized complex of a human αβ T-cell receptor, influenza HA peptide and MHC class II molecule, HLA-DR1. EMBO J. 19, 5611–5624 (2000).
Reiser, J.B. et al. Crystal structure of a T cell receptor bound to an allogeneic MHC molecule. Nat. Immunol. 1, 291–297 (2000).
Reiser, J.B. et al. A T cell receptor CDR3β loop undergoes conformational changes of unprecedented magnitude upon binding to a peptide/MHC class I complex. Immunity 16, 345–354 (2002).
Garboczi, D.N. et al. Structure of the complex between human T-cell receptor, viral peptide and HLA-A2. Nature 384, 134–141 (1996).
Ding, Y.H. et al. Two human T cell receptors bind in a similar diagonal mode to the HLA- A2/Tax peptide complex using different TCR amino acids. Immunity 8, 403–411 (1998).
Mazza, C. et al. How much can a T-cell antigen receptor adapt to structurally distinct antigenic peptides? EMBO J. 26, 1972–1983 (2007).
Tynan, F.E. et al. T cell receptor recognition of a 'super-bulged' major histocompatibility complex class I–bound peptide. Nat. Immunol. 6, 1114–1122 (2005).
Kjer-Nielsen, L. et al. A structural basis for the selection of dominant alphabeta T cell receptors in antiviral immunity. Immunity 18, 53–64 (2003).
Chothia, C. & Lesk, A.M. Canonical structures for the hypervariable regions of immunoglobulins. J. Mol. Biol. 196, 901–917 (1987).
Mian, I.S., Bradwell, A.R. & Olson, A.J. Structure, function and properties of antibody binding sites. J. Mol. Biol. 217, 133–151 (1991).
Fellouse, F.A. et al. Molecular recognition by a binary code. J. Mol. Biol. 348, 1153–1162 (2005).
Al-Lazikani, B., Lesk, A.M. & Chothia, C. Canonical structures for the hypervariable regions of T cell αβ receptors. J. Mol. Biol. 295, 979–995 (2000).
Otwinowski, Z., Minor, W. & Carter, C.W., Jr. in Methods in Enzymology Vol. 276 (eds. Abelson, J.N., Simon, M.I., Carter, C.W. Jr. & Sweet, R.M.) 307–326 (Academic, New York, 1997).
Vagin, A. & Teplyakov, A. MOLREP: an automated program for molecular replacement. J. Appl. Crystallogr. 30, 1022–1025 (1997).
Li, H., Lebedeva, M.I., Ward, E.S. & Mariuzza, R.A. Dual conformations of a T cell receptor Vα homodimer: implications for variability in VαVβ domain association. J. Mol. Biol. 269, 385–394 (1997).
Emsley, P. & Cowtan, K. COOT: model-building tools for molecular graphics. Acta Crystallogr. D Biol. Crystallogr. 60, 2126–2132 (2004).
Brunger, A.T. et al. Crystallography & NMR system: A new software suite for macromolecular structure determination. Acta Crystallogr. D Biol. Crystallogr. 54, 905–921 (1998).
Read, R. Pushing the boundaries of molecular replacement with maximum likelihood. Acta Crystallographica D Biol. Crystallogr. 57, 1373–1382 (2001).
DeLano, W.L. The PyMol molecular graphics system. (DeLano Scientific, San Carlos, California, 2002).
Acknowledgements
We acknowledge M. Davis for discussions and access to a BIAcore 3000. Supported by the National Institutes of Health (AI48540 to K.C.G.), the Howard Hughes Medical Institute (K.C.G.) and the National Health and Medical Research Council of Australia (CJ Martin Fellowship to L.K.E.).
Author information
Authors and Affiliations
Contributions
D.F., X-ray crystallographic analyses; D.F., C.J.B, L.K.E. and J.M., biochemical and biophysical studies; and K.C.G., project direction.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figure 1 and Tables 1–2 (PDF 1737 kb)
Rights and permissions
About this article
Cite this article
Feng, D., Bond, C., Ely, L. et al. Structural evidence for a germline-encoded T cell receptor–major histocompatibility complex interaction 'codon'. Nat Immunol 8, 975–983 (2007). https://doi.org/10.1038/ni1502
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ni1502
This article is cited by
-
NetTCR-2.0 enables accurate prediction of TCR-peptide binding by using paired TCRα and β sequence data
Communications Biology (2021)
-
Structural understanding of T cell receptor triggering
Cellular & Molecular Immunology (2020)
-
Structural basis for oligoclonal T cell recognition of a shared p53 cancer neoantigen
Nature Communications (2020)
-
DynaDom: structure-based prediction of T cell receptor inter-domain and T cell receptor-peptide-MHC (class I) association angles
BMC Structural Biology (2018)
-
Germline bias dictates cross-serotype reactivity in a common dengue-virus-specific CD8+ T cell response
Nature Immunology (2017)