Abstract
Individual B lymphocytes normally express immunoglobulin (Ig) proteins derived from single Ig heavy chain (H) and light chain (L) alleles. Allelic exclusion ensures monoallelic expression of Ig genes by each B cell to maintain single receptor specificity. Here we provide evidence that at later stages of B cell development, additional mechanisms may contribute to prioritizing expression of single IgH and IgL alleles. Fluorescent in situ hybridization analysis of primary splenic B cells isolated from normal and genetically manipulated mice showed that endogenous IgH, κ and λ alleles localized to different subnuclear environments after activation and had differential expression patterns. However, this differential recruitment and expression of Ig alleles was not typically seen among transformed B cell lines. These data raise the possibility that epigenetic factors help maintain the monoallelic expression of Ig.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Ohlsson, R., Tycko, B. & Sapienza, C. Monoallelic expression: 'there can only be one'. Trends Genet. 14, 435–438 (1998).
Honjo, T. & Alt, F. W. Immunoglobulin Genes (Academic Press, London, 1995).
Rajewsky, K. Clonal selection and learning in the antibody system. Nature 381, 751–758 (1996).
Rolink, A. & Melchers, F. Generation and regeneration of cells of the B-lymphocyte lineage. Curr. Opin. Immunol. 5, 207–217 (1993).
Schatz, D. G., Oettinger, M. A. & Baltimore, D. The V(D)J recombination activating gene, RAG-1. Cell 59, 1035–1048 (1989).
Barreto, B. & Cumano, A. Frequency and characterization of phenotypic Ig heavy chain allelically included IgM-expressing B cells in mice. J. Immunol. 164, 893–899 (2000).
Dernburg, A. F. et al. Perturbation of nuclear architecture by long-distance chromosome interactions. Cell 85, 745–759 (1996).
Andrulis, E. D., Neiman, A. M., Zappulla, D. C. & Sternglanz, R. Perinuclear localization of chromatin facilitates transcriptional silencing. Nature 394, 592–595 (1998).
Brown, K. E., Baxter, J., Graf, D., Merkenschlager, M. & Fisher, A. G. Dynamic repositioning of genes in the nucleus of lymphocytes preparing for cell division. Mol. Cell 3, 207–217 (1999).
Brown, K. E. et al. Association of transcriptionally silent genes with Ikaros complexes at centromeric heterochromatin. Cell 91, 845–854 (1997).
Gasser, S. M. Positions of potential: nuclear organization and gene expression. Cell 104, 639–642 (2001).
Grogan, J. L. et al. Early transcription and silencing of cytokine genes underlie polarization of T helper cell subsets. Immunity 14, 205–215 (2001).
Francastel, C., Walters, M. C., Groudine, M. & Martin, D. I. K. A functional enhancer suppresses silencing of a transgene and prevents its localization close to centromeric heterochromatin. Cell 99, 259–269 (1999).
Schübeler, D. et al. Nuclear localization and histone acetylation: a pathway for chromatin opening and transcriptional activation of the human β-globin locus. Genes Dev. 14, 940–950 (2000).
Lundgren, M. et al. Transcription factor dosage affects changes in higher order chromatin structure associated with activation of a heterochromatic gene. Cell 103, 733–743 (2000).
Georgopouos, K. et al. The Ikaros gene is required for the development of all lymphoid lineages. Cell. 79, 43–156 (1994).
Sabbattini, P. et al. Binding of Ikaros to the λ5 promoter silences transcription through a mechanism that does not require heterochromatin formation. EMBO J. 20, 2812–2822 (2001).
Elenich, L. A., Ford, C. S., Collins, J. T. & Dunnick, W. A. γ 1 heavy chain transgenes are responsive to IFN-γ repression and CD40 ligation. J. Immunol. 158, 4564–4573 (1997).
Ratcliffe, M. J. & Julius, M. H. H-2 restricted T-B cell interactions involved in polyspecific B cell responses mediated by soluble antigen. Eur. J. Immunol. 12, 634–641 (1982).
Hood, L., Grant, J. A. & Sox, H. C. Jr in Developmental Aspects of Antibody Formation and Structure (eds. Sterzl, J. & Hiha, I.) vol. 1, 283–295 (Academic Press, New York, 1969).
Bernier, G. M. & Cebra, J. J. Polypeptide chains of human γ-globulin: cellular localization by fluorescent antibody. Science 144, 1590–1591 (1964).
Zou, Y-R., Takeda, S. & Rajewsky, K. Gene targeting in the Igκ locus: efficient generation of λ chain-expressing B cells, independent of gene rearrangements in Igκ. EMBO J. 12, 811–820 (1993).
Esser, C. & Radbruch, A. Immunoglobulin class switching; molecular and cellular analysis. Annu. Rev. Immunol. 8, 717–735 (1990).
Fisher, A. G., Burdet, C., Bunce, C., Merkenschlager, M. & Ceredig, R. Lymphoproliferative disorders in IL-7 transgenic mice: expansion of immature B cells which retain macrophage potential. Int. Immunol. 7, 415–423 (1995).
Nelson, K. J., Haimovich, J. & Perry, R. P. Characterization of productive and sterile transcripts from the immunoglobulin heavy-chain locus: processing of μm and μs mRNA. Mol. Cell Biol. 3, 1317–1332 (1983).
Lennon, G. G. & Perry, R. P. Cμ-containing transcripts initiate heterogeneously within the IgH enhancer region and contain a novel 5′-nontranslatable exon. Nature 318, 475–478 (1985).
Lennon, G. G. & Perry, R. P. Identification of a defective mouse immunoglobulin D (diversity) element which can undergo DJH, but not VHD, recombination. Immunogenetics 30, 383–386 (1989).
Nelson, K. J., Mather, E. L. & Perry, R. P. Lipopolysaccharide-induced transcription of the κ immunoglobulin locus occurs on both alleles and is independent of methylation status. Nucleic Acids Res. 12, 1911–1923 (1984).
Bridger, J. M., Boyle, S., Kill, I. R. & Bickmore, W. A. Re-modelling of nuclear architecture in quiescent and senescent human fibroblasts. Curr. Biol. 10, 149–152 (2000).
Nemazee, D. Receptor selection in B and T Lymphocytes. Annu. Rev. Immunol. 18, 19–51 (2000).
Kelley, D. E. & Perry, R. P. Transcriptional and posttranscriptional control of immunoglobulin mRNA production during B lymphocyte development. Nucleic Acids Res. 14, 5431–5447 (1986).
Kim, K. J., Kanellopoulos-Langevin, C., Merwin, R. M., Sachs, D. H. & Asofsky, R. Establishment and characterization of BALB/c lymphoma lines with B cell properties. J. Immunol. 122, 549–554 (1979).
Weiss, S. & Wu, G. E. Somatic point mutations in unrearranged immunoglobulin gene segments encoding the variable region of λ light chains. EMBO J. 6, 927–932 (1987).
Shimizu, A., Takahashi, N., Yaoita, Y. & Honjo, T. Organisation of the constant-region gene family of the mouse immunoglobulin heavy chain. Cell 28, 499–506 (1982).
Mainville, C. A. et al. Deletion mapping of fifteen mouse VH gene families reveals a common organization for three Igh haplotypes. J. Immunol. 3, 1038–46 (1996).
Liu, C. P., Tucker, P. W., Mushinski, J. F. & Blattner, F.R. Mapping of heavy chain genes for mouse immunoglobulins M and D. Science 209, 1348–1353 (1980).
Hahm, K. et al. The lymphoid transcription factor LyF-1 is encoded by specific, alternatively spliced mRNAs derived from the Ikaros gene. Mol. Cell. Biol. 14, 7111–7123 (1994).
Gu, H., Kitamura, D. & Rajewsky, K. B cell development regulated by gene rearrangement: arrest of maturation by membrane-bound Dμ protein and selection of DH element reading frames. Cell 65, 47–54 (1991).
Acknowledgements
We thank our colleagues for supplying mouse genomic probes; S. Smale for anti-Ikaros sera; G. Klaus for anti-IgM; J. Roes for Cμ-Cδ probes; P. Brodeur for VHJ558 and VHS107 probes; E. Severinson, L. Ström, V. Buckle and N. Brockdorff for helpful discussions; and G. Reed, R. Newton and I. Devonish for photographic and secretarial assistance. Supported by the Medical Research Council, the Wellcome Trust (J. S. and D. G.) and the Royal Society (K. E. B.).
Author information
Authors and Affiliations
Corresponding author
Supplementary information
Web Figure 1.
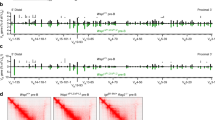
Measurement of allelic distances between IgH loci and γ-satellite DNA. (a) Sequential images (ordered from the top left to right, middle left to right, lower left to right) of optical sections through the nucleus of a 4-day-activated mouse B cell nucleus (Z series, 0.2 µm separation) in which the location of IgH alleles (green) and γ-satellite DNA (red) is shown. Data from 54 individual cells was subjected to 3D reconstruction (using Metamorph Imaging series 4.5, Universal Imaging Corporation) to allow 3-D distances between pixels in X, Y and Z planes to be estimated. (b,c) Intranuclear distances (in mm) between the centroid (signal intensity weighted center) of the nearest γ-satellite signal and the center of each IgH allele (maximum intensity). (b) Distribution analysis in which the distance of the nearest (allele 1) and furthest (allele 2) IgH allele from centromeric DNA, is given. (c) Distances were plotted for each individual cell. (GIF 55 kb)
Rights and permissions
About this article
Cite this article
Skok, J., Brown, K., Azuara, V. et al. Nonequivalent nuclear location of immunoglobulin alleles in B lymphocytes. Nat Immunol 2, 848–854 (2001). https://doi.org/10.1038/ni0901-848
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/ni0901-848
This article is cited by
-
Nuclear positioning and pairing of X-chromosome inactivation centers are not primary determinants during initiation of random X-inactivation
Nature Genetics (2019)
-
The importance of the nuclear positioning of the PPARG gene for its expression during porcine in vitro adipogenesis
Chromosome Research (2019)
-
Lamin B1 regulates somatic mutations and progression of B-cell malignancies
Leukemia (2018)
-
CRISPR-dCas9 and sgRNA scaffolds enable dual-colour live imaging of satellite sequences and repeat-enriched individual loci
Nature Communications (2016)
-
Random monoallelic expression of autosomal genes: stochastic transcription and allele-level regulation
Nature Reviews Genetics (2015)