Abstract
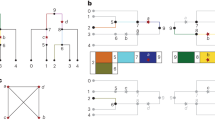
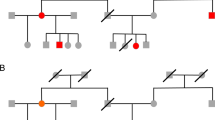
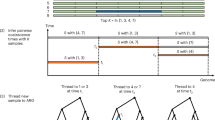
Efforts to find disease genes using high-density single-nucleotide polymorphism (SNP) maps will produce data sets that exceed the limitations of current computational tools. Here we describe a new, efficient method for the analysis of dense genetic maps in pedigree data that provides extremely fast solutions to common problems such as allele-sharing analyses and haplotyping. We show that sparse binary trees represent patterns of gene flow in general pedigrees in a parsimonious manner, and derive a family of related algorithms for pedigree traversal. With these trees, exact likelihood calculations can be carried out efficiently for single markers or for multiple linked markers. Using an approximate multipoint calculation that ignores the unlikely possibility of a large number of recombinants further improves speed and provides accurate solutions in dense maps with thousands of markers. Our multipoint engine for rapid likelihood inference (Merlin) is a computer program that uses sparse inheritance trees for pedigree analysis; it performs rapid haplotyping, genotype error detection and affected pair linkage analyses and can handle more markers than other pedigree analysis packages.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Mullikin, J.C. et al. An SNP map of human chromosome 22. Nature 407, 516–520 (2000).
Altshuler, D. et al. An SNP map of the human genome generated by reduced representation shotgun sequencing. Nature 407, 513–516 (2000).
Lathrop, G.M., Lalouel, J.M., Julier, C. & Ott, J. Multilocus linkage analysis in humans: detection of linkage and estimation of recombination. Am. J. Hum. Genet. 37, 482–498 (1985).
Kruglyak, L. & Lander, E.S. Complete multipoint sib-pair analysis of qualitative and quantitative traits. Am. J. Hum. Genet. 57, 439–454 (1995).
O'Connell, J.R. & Weeks, D.E. The VITESSE algorithm for rapid exact multilocus linkage analysis via genotype set-recoding and fuzzy inheritance. Nature Genet. 11, 402–408 (1995).
Cottingham, R.W. Jr, Idury, R.M. & Schaffer, A.A. Faster sequential genetic linkage computations. Am. J. Hum. Genet. 53, 252–263 (1993).
Sobel, E. & Lange, K. Descent graphs in pedigree analysis: applications to haplotyping, location scores, and marker-sharing statistics. Am. J. Hum. Genet. 58, 1323–1337 (1996).
Heath, S.C. Markov chain Monte Carlo segregation and linkage analysis for oligogenic models. Am. J. Hum. Genet. 61, 748–760 (1997).
Gudbjartsson, D.F., Jonasson, K., Frigge, M.L. & Kong, A. Allegro, a new computer program for multipoint linkage analysis. Nature Genet. 25, 12–13 (2000).
Elston, R.C. & Stewart, J. A general model for the genetic analysis of pedigree data. Hum. Hered. 21, 523–542 (1971).
Lander, E.S. & Green, P. Construction of multilocus genetic linkage maps in humans. Proc. Natl Acad. Sci. USA 84, 2363–2367 (1987).
Guo, S.W. & Thompson, E.A. A Monte Carlo method for combined segregation and linkage analysis. Am. J. Hum. Genet. 51, 1111–1126 (1992).
Douglas, J.A., Boehnke, M. & Lange, K. A multipoint method for detecting genotyping errors and mutations in sibling-pair linkage data. Am. J. Hum. Genet. 66, 1287–1297 (2000).
Abecasis, G.R., Cherny, S.S. & Cardon, L.R. The impact of genotyping error on linkage and association analysis of quantitative traits. Eur. J. Hum. Genet. 9, 130–134 (2001).
Gordon, D., Heath, S.C. & Ott, J. True pedigree errors more frequent than apparent errors for single nucleotide polymorphisms. Hum. Hered. 49, 65–70 (1999).
Markianos, K., Daly, M.J. & Kruglyak, L. Efficient multipoint linkage analysis through reduction of inheritance space. Am. J. Hum. Genet. 68, 963–977 (2001).
Press, W.H., Teukolsky, S.A., Vetterling, W.T. & Flannery, B.P. Numerical Recipes in C. (Cambridge University Press, New York, 1992).
Idury, R.M. & Elston, R.C. A faster and more general hidden Markov model algorithm for multipoint likelihood calculations. Hum. Hered. 47, 197–202 (1997).
Excoffier, L. & Slatkin, M. Maximum-likelihood estimation of molecular haplotype frequencies in a diploid population. Mol. Biol. Evol. 12, 921–927 (1995).
Keavney, B. et al. Measured haplotype analysis of the angiotensin-I converting enzyme gene. Hum. Mol. Genet. 7, 1745–1751 (1998).
Gray, F. Pulse Code Communication. in Patent 2,632,058 (USA, 1953).
Acknowledgements
This work was supported by the Wellcome Trust through a Prize Studentship (G.R.A.) Senior Research Fellowship (W.O.C.) and a Principal Research Fellowship (L.R.C.), and by the National Eye Institute (S.S.C. and L.R.C.).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Abecasis, G., Cherny, S., Cookson, W. et al. Merlin—rapid analysis of dense genetic maps using sparse gene flow trees. Nat Genet 30, 97–101 (2002). https://doi.org/10.1038/ng786
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng786
This article is cited by
-
Improved computations for relationship inference using low-coverage sequencing data
BMC Bioinformatics (2023)
-
Clustering of Juvenile Canavan disease in an Indian community due to population bottleneck and isolation: genomic signatures of a founder event
European Journal of Human Genetics (2023)
-
Common genetic risk factors in ASD and ADHD co-occurring families
Human Genetics (2023)
-
Variants in SART3 cause a spliceosomopathy characterised by failure of testis development and neuronal defects
Nature Communications (2023)
-
Genome-wide linkage search for cancer susceptibility loci in a cohort of non BRCA1/2 families in Sri Lanka
BMC Research Notes (2022)