Abstract
We identified 255 loci across the human genome that contain genomic imbalances among unrelated individuals. Twenty-four variants are present in > 10% of the individuals that we examined. Half of these regions overlap with genes, and many coincide with segmental duplications or gaps in the human genome assembly. This previously unappreciated heterogeneity may underlie certain human phenotypic variation and susceptibility to disease and argues for a more dynamic human genome structure.
Similar content being viewed by others
Main
Variation in the human genome is present in many forms, including single-nucleotide polymorphisms, small insertion-deletion polymorphisms, variable numbers of repetitive sequences, and genomic structural alterations1. Molecular genetic and cytogenetic analyses have catalogued many variations in the human genome, but little is known about large-scale copy-number variations (LCVs) that involve gains or losses of several kilobases to hundreds of kilobases of genomic DNA among phenotypically normal individuals. To investigate these LCVs in the human genome, we applied array-based comparative genomic hybridization (array CGH)2 to the genomes of 55 unrelated individuals. The arrays used contain selected large insert DNA fragments, distributed roughly every 1 Mb throughout the human genome. We compared genomic DNA samples isolated from 39 unrelated healthy control individuals with normal karyotypes and from 16 individuals with previously characterized chromosomal imbalances3 with pooled male or female genomic DNAs from karyotypically and phenotypically normal control individuals (Supplementary Methods online). Including samples from individuals with chromosomal imbalances allowed us to monitor the sensitivity and specificity of experiments, and we did detect all expected abnormalities (we excluded the regions of known imbalances from our analysis).
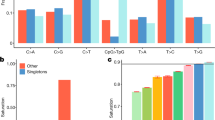
Our experiments identified 255 individual genomic clones that showed comparative gains or losses among the samples that we tested. Most of the clones seemed to be randomly distributed throughout the genome (Fig. 1 and Supplementary Table 1 online). On average, we observed 12.4 LCVs per individual. Most of these variants involve single large-insert genomic clones, suggesting that each of the identified LCVs may result from gains or losses involving as much as 2 Mb of DNA sequence (1 Mb to each flanking clone). We identified 102 LCVs (41%) that occurred in more than one individual and 24 LCVs that were present in > 10% of the individuals studied. The remaining 153 clones may represent LCVs that occur at lower frequencies. The genomic regions that we identified probably do not represent false positives, because control self-versus-self hybridization experiments indicated that there was less than 1 false positive clone for every two experiments (< 1 per 5,264 clones; Supplementary Fig. 1 online). Of the 102 recurring LCVs, 26 (25.5%) mapped to regions overlapping previously recognized segmental duplications4,5. This proportion is significantly higher than that observed for all clones on the array (7.3%; P < 0.0001). Moreover, 13 (12.7%) of the recurring LCVs reside within 100 kb of gaps in the current presentation of the human genome sequence; this proportion is also significantly higher than the expected 3.6% for all the clones of the array (P < 0.0001). This suggests that the presence of these LCVs may complicate the accurate assembly of sequences at these chromosomal loci3,5,6.
Circles to the right of each chromosome ideogram show the number of individuals with copy gains (blue) and losses (red) for each clone among 39 unrelated, healthy control individuals. Green circles to the left indicate known genome sequence gaps within 100 kb of the clone, or segmental duplications known to overlap the clone, as compared to the Human Recent Segmental Duplication Browser. Cytogenetic band positions are shown to the left.
We found that 142 of 255 (56%) polymorphic clones overlapped with known coding regions and that 67 clones encompassed one or more entire genes. This suggests that LCVs are not limited to intergenic or intronic regions. Fourteen LCVs were located near loci associated with human genetic syndromes or with cancer (Supplementary Table 2 online). Because these variants exist in the genomes of phenotypically normal individuals, they may not be a direct cause of genetic disease, but their presence could lead to chromosomal rearrangements that give rise to disease7,8,9 or more subtle phenotypic variation by influencing expression of specific genes10.
The most common LCV (identified in 49.1% of the individuals studied) encompassed the amylase alpha 1a and alpha 2a locus (AMY1A-AMY2A) at chromosome region 1p13.3 (ref. 11). We detected relative gains (in 23.6% of cases) and losses (in 25.5% of cases) at this locus and confirmed the array CGH results using metaphase-interphase fluorescence in situ hybridization (FISH), high-resolution fiber FISH and quantitative PCR (Fig. 2 and Supplementary Methods online). The length of this polymorphic region was estimated to range from 150 kb to 425 kb, and quantitative PCR results indicated that the length varied by a factor of 2.5 among the same individuals. Metaphase and interphase FISH data for this locus and 18 others indicated that each of the LCVs analyzed was confined to localized chromosomal regions (Supplementary Table 1 online). Therefore, the formation of these LCVs probably reflects, as with the amylase locus, tandem copy-number changes rather than duplication events involving other chromosomal loci. Eight identified LCVs map to regions previously described to exhibit variable copy numbers of genes or pseudogenes in the general population (Supplementary Table 2 online).
Arrows on the right side of chromosome 1 array CGH profiles indicate relative losses (a), normal ratios (b) or relative gains (c) for the LCV represented by BAC RP11-259N12 in three unrelated healthy individuals. FISH analyses with a Spectrum Green–labeled RP11-259N12 probe on metaphase chromosome preparations from these same individuals showed a normal hybridization pattern of one signal per chromatid. Likewise, two signals (or pairs of signals) were observed in 50 scored interphase nuclei from these same individuals. Quantification of signal intensities showed a correlation of probe hybridization with respective array CGH gain and loss data (data not shown). High-resolution fiber FISH was done on stretched DNA fibers from the same cases using the RP11-259N12 probe cohybridized with a 5′ amylase gene probe (green) and a 3′ amylase gene probe (red). The case with a relative loss showed 6 gene signals (a), the case with a normal signal showed 9 gene signals (b), and the case with a gain showed 12 signals (c). The approximate length of the polymorphic region was estimated to vary from ∼150 kb (a) to ∼425 kb (c). Quantitative PCR (d) done on DNA from the same individuals was consistent with the array CGH and FISH findings.
We described more than 200 LCVs in the human genome. Twenty-four of these variants are present in > 10% of the individuals that we studied, and six of these variants are present at a frequency of > 20%. Because the array platform used for these studies comprises only 12% of the total human genome sequence, denser arrays12 will probably detect additional new genomic variations. In contrast to most small-scale genetic polymorphisms, the LCVs described here may have an important functional effect on the evolution of the human genome. To catalog this large-scale genomic variation, we collated all the available information into a database (the Genome Variation Database), which will be a crucial resource in correlating these genomic variations with experimental findings and clinical outcomes.
URLs. The Genome Variation Database is available at http://projects.tcag.ca/variation/. The Human Recent Segmental Duplication Browser is available at http://projects.tcag.ca/humandup/.
Note: Supplementary information is available on the Nature Genetics website.
References
Wright, A.F. Nature Encyclopedia of the Human Genome vol. 2, 959–968 (Nature Publishing Group, London, 2003).
Albertson, D.G. & Pinkel, D. Hum. Mol. Genet. 12, R145–R152 (2003).
Scherer, S.W. et al. Science 300, 767–772 (2003).
Bailey, J.A. et al. Science 297, 1003–1007 (2002).
Cheung, J. et al. Genome Biol. 4, R25 (2003).
Eichler, E.E., Clark, R.A. & She, X. Nat. Rev. Genet. 5, 345–354 (2004).
Shaw, C.J. & Lupski, J.R. Hum. Mol. Genet. 13, R57–R64 (2004).
Giglio, S. et al. Am. J. Hum. Genet. 68, 874–883 (2001).
Osborne, L.R. et al. Nat. Genet. 29, 321–325 (2001).
Hollox, E.J., Armour, J.A. & Barber, J.C. Am. J. Hum. Genet. 73, 591–600 (2003).
Groot, P.C., Mager, W.H. & Frants, R.R. Genomics 10, 779–785 (1991).
Ishkanian, A.S. et al. Nat. Genet. 36, 299–303 (2004).
Acknowledgements
We thank J. Kim for biostatistical assistance. This work was supported in part by a grant from the Friends of the Dana-Farber Cancer Institute (C.L.), the Brigham and Women's Hospital Pathology Department training grant (A.J.I.), Genome Canada (S.W.S.) and an National Institute of Health program project grant (P.K.D.). L.F. is supported by The Swedish Medical Research Council, and S.W.S. is an Investigator of the Canadian Institutes of Health Research and an International Scholar of the Howard Hughes Medical Institute.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Fig. 1
Genome-wide array CGH plots for two representative self vs. self hybridization experiments. (PDF 162 kb)
Supplementary Table 1
Comprehensive description of large-scale copy number variations detected by array CGH. (PDF 141 kb)
Supplementary Table 2
Large-scale copy number variations near known polymorphic and disease loci. (PDF 86 kb)
Rights and permissions
About this article
Cite this article
Iafrate, A., Feuk, L., Rivera, M. et al. Detection of large-scale variation in the human genome. Nat Genet 36, 949–951 (2004). https://doi.org/10.1038/ng1416
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng1416
This article is cited by
-
Ethnic and functional differentiation of copy number polymorphisms in Tunisian and HapMap population unveils insights on genome organizational plasticity
Scientific Reports (2024)
-
Nexus between genome-wide copy number variations and autism spectrum disorder in Northeast Han Chinese population
BMC Psychiatry (2023)
-
Exploring quantitative traits-associated copy number deletions through reanalysis of UK10K consortium whole genome sequencing cohorts
BMC Genomics (2023)
-
Frequent copy number variants in a cohort of Mexican-Mestizo individuals
Molecular Cytogenetics (2023)
-
Whole-genome sequencing reveals an association between small genomic deletions and an increased risk of developing Parkinson’s disease
Experimental & Molecular Medicine (2023)