Abstract
Spinocerebellar ataxia type 10 (SCA10; MIM 603516; refs 1,2) is an autosomal dominant disorder characterized by cerebellar ataxia and seizures. The gene SCA10 maps to a 3.8-cM interval on human chromosome 22q13–qter (refs 1,2). Because several other SCA subtypes show trinucleotide repeat expansions, we examined microsatellites in this region. We found an expansion of a pentanucleotide (ATTCT) repeat in intron 9 of SCA10 in all patients in five Mexican SCA10 families. There was an inverse correlation between the expansion size, up to 22.5 kb larger than the normal allele, and the age of onset (r2=0.34, P=0.018). Analysis of 562 chromosomes from unaffected individuals of various ethnic origins (including 242 chromosomes from Mexican persons) showed a range of 10 to 22 ATTCT repeats with no evidence of expansions. Our data indicate that the new SCA10 intronic ATTCT pentanucleotide repeat in SCA10 patients is unstable and represents the largest microsatellite expansion found so far in the human genome.
Similar content being viewed by others
Main
In addition to two previously described families with SCA10 (refs 1,2), we identified three additional Mexican families with an autosomal dominant disease characterized by cerebellar ataxia and seizures (Fig. 1a). The age of disease onset appeared to be earlier in successive generations in these families. This clinical phenomenon, known as ‘anticipation’, is common in dominantly inherited SCAs, in which expanded CAG repeats encoding polyglutamine tracts in the respective genes are unstable and have larger expansions in successive generations3,4,5,6,7,8. Here, we found neither expansion of the CAG repeats involved in SCA1–3, 6, 7 or 12 (refs 3–10) nor expansion of the CTG repeat at the SCA8 locus11.
a, Pedigrees of the four families studied for the SCA10 mutation. Family ‘a’ is of Mexican American descent1. Families ‘b’, ‘c’ and ‘d’ are newly identified Mexican families with relatively pure cerebellar ataxia and seizures. b, PCR analysis of the ATTCT pentanucleotide repeat. The ATTCT pentanucleotide repeat region was PCR-amplified from the genomic DNA samples of the family members indicated in (a) using primers attct-L and attct-R. All affected individuals showed a single allele of variable size in families a, b, c and d (note that each band is accompanied by a faint band underneath due to PCR artefact). In family a, two unaffected individuals (I-1 and III-2) are heterozygous and two spouses (II-1, III-5) are homozygous for the ATTCT repeat. In this family, affected individuals in the second generation (II-2 and II-3) failed to transmit their 12-repeat allele to their affected offspring (III-1, III-3, III-4, III-6, III-7, III-8 and III-9), whereas an unaffected offspring (III-2) received this allele from the affected father (II-2). The alleles of unaffected parents (I-1 and II-1) were passed on to their offspring in a pattern consistent with mendelian inheritance. These data suggest that the affected individuals are apparently hemizygous for the ATTCT repeat. c, Southern-blot analysis of expansion mutations of the ATTCT repeat region. Southern blots of genomic DNA samples revealed variably expanded alleles in affected members of the families shown above. All individuals examined have a normal allele (2.5 kb). The apparent variability of the normal allele size is attributable to gel-loading artefacts, as additional analyses using the same (EcoRI) and different (EcoRV, HindIII and BglI) restriction endonucleases did not show consistent variability of the normal allele size. The genotype of each individual is shown at the bottom, with an estimated number of pentanucleotide repeats for disease-associated chromosomes based on the fragment size.
Two recombination events narrowed the SCA10 region to a 2.7-cM region between D22S1140 and D22S1153 (refs 1,2,12). Although the DNA sequence of the entire euchromatic part of human chromosome 22 is now available, there are still 11 gaps that remain to be sequenced13. D22S1153 resides in one of these gaps. Consequently, we could not determine the exact physical size of the SCA10 candidate region. Nevertheless, two contigs composed of bacterial artificial chromosomes (BACs), phage P1-derived artificial chromosomes (PACs) and cosmids cover most of this region.
Because of the dominant inheritance of SCA10 and the suggestion of anticipation, we examined 14 trinucleotide repeats (>3 repeats in length) listed in the SCA10 candidate interval in the chromosome 22 genome database at the Sanger Centre13. None of these repeats showed expansions in the affected members of SCA10 families. Moreover, repeat expansion detection (RED) analysis14 of genomic DNA samples from affected family members also failed to show evidence of a CAG or CAA expansion (data not shown). Western-blot analysis of proteins obtained from patients' lymphoblastoid cells using a monoclonal antibody that recognizes expanded polyglutamine tracts15 failed to detect proteins (data not shown). These findings prompted us to extend our investigation to microsatellites other than trinucleotide repeats.
PCR analysis of the region spanning a pentanucleotide (ATTCT) tandem repeat in intron 9 of SCA10 (previously designated E46; Fig. 3) showed repeat number polymorphism in normal individuals (Figs 1b and Fig. 2). Normal alleles contained from 10 to 20 repeats with 82.1% heterozygosity. Sequence analysis of the alleles obtained from 20 normal individuals showed tandem repeats of ATTCT without interruption. The distribution of the alleles was unimodal, with similar patterns among the European, Mexican and Japanese populations examined (Fig. 2). The allele distributions in each of three ethnically defined populations (including 127 persons from the Mexican population) were consistent with the Hardy-Weinberg equilibrium (P>0.05). In SCA10 families, PCR analysis demonstrated a uniform lack of heterozygosity in all affected individuals and carriers of the disease haplotype1,2, with the single amplified allele of the ATTCT repeat being shared by their unaffected parent. When the affected and unaffected parents of the patients carried distinct ATTCT repeat alleles, we found that the single allele amplified from the affected parent was never transmitted to any of these patients, suggesting that the affected parent was hemizygous and that only the allele on the wild-type (non-SCA10) chromosome is amplified (Fig. 1b). Two other sets of primers flanking the ATTCT repeat gave the same result, excluding the possibility that the apparent hemizygosity is due to a mutation within the primer binding site. To further investigate these observations, we hybridized Southern blots of genomic DNA digested with multiple restriction endonucleases with a non-repetitive probe obtained by PCR amplification of the region immediately upstream of the ATTCT repeat, using DNA from a PAC clone (RP1-37M3) as template (Fig. 3b). In addition to the expected normal allele, we detected a variably expanded allele in all affected individuals, whereas we saw only the wild-type allele in all unaffected family members, demonstrating that the ATTCT repeat region is expanded exclusively in SCA10 patients (Fig. 1c). So far, we have found SCA10 only in the Mexican population, where it is the second most common autosomal dominant cerebellar ataxia after SCA2 (A.S. et al., unpublished data). We did not detect the SCA10 expansion mutation in 17 probands of unrelated European American families with autosomal dominant cerebellar ataxia that have tested negative for the SCA1–3, -6, -7, -8 or -12 mutations. It remains unknown whether SCA10 exists at a low frequency in this population.
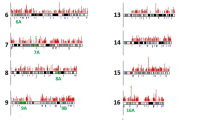
Shown is a histogram of the normal ATTCT repeat alleles in European (n=250), Japanese (n=100) and Mexican (n=254) chromosomes. The distribution of the alleles was unimodal with similar patterns among the European, Mexican and Japanese populations examined. There were, however, subtle differences between these populations. In the Mexican and Japanese populations, the 14-ATTCT-repeat allele was the modal allele with frequencies of 29% and 27%, respectively. In the European population, the 13-repeat allele was modal with a frequency of 30%. Larger samples, however, will be necessary to verify the statistical significance of these results.
a, A schematic presentation of the structure of SCA10. SCA10 consists of 12 exons. The ATTCT repeat is located in intron 9. The 12 exons of SCA10 span 172.8 kb of genomic DNA with an ORF of 1,428 bp, encoding 475 amino acids with no homology to known human proteins and one presumed mouse orthologue (82% identity, 91% similarity). Analysis of the SCA10 predicted amino acid sequence reveals a novel protein with no recognizable motifs or predicted structures. Sequence analysis of the large size of intron 9 (66,420 bp) produced no evidence for additional expressed sequences. The gap at the left of PAC 37M3 does not represent missing sequence, but was introduced to preserve scale. b, A restriction map of the ATTCT repeat region defined by flanking HindIII restriction sites (nt 17,023 and 34,567). The numbers are nucleotide positions in PAC37M3. ‘Probe’ indicates the position of the probe used (nt 25,222–26,021) to detect the 2.5-kb EcoRI fragment shown in Fig. 2c in the Southern-blot analysis. The ATTCT repeat is located downstream of the probe within the 2.5-kb EcoRI fragment. c, Nucleotide sequence of the ATTCT repeat (14 repeats; underlined, nt 26,101–26170) and the flanking regions (nt 25,981–26,281). Arrows underline PCR primer sequences (attct-L and attct-R) that were used for amplification of the ATTCT repeat region shown in Fig. 2b.
The variable size of expanded alleles among affected individuals suggests that the expanded ATTCT repeat is highly unstable. In the SCA10 families we studied, intergenerational changes of the expanded alleles were variable, including not only expansions but also contractions (Fig. 1c). Studies of additional SCA10 families should provide further details of the genotype-phenotype correlation and the patterns of instability of the expanded ATTCT repeats. Haplotype analysis using DNA markers flanking the ATTCT repeat showed a single disease haplotype shared by all SCA10 families studied (data not shown). Thus, our current haplotype data cannot exclude the possibility that the expanded repeat represents a rare polymorphism in linkage disequilibrium to the true pathogenic mutation. A notable observation in our SCA10 families was the weak inverse correlation between the size of the expanded allele and the age of disease onset (n=26, r2=0.34, P=0.018) (Fig. 4). There may be heterogeneity in expansion size among different tissues, and the correlation we found might have been stronger if we had compared patient brain tissues, but these were not available. Although direct evidence that the ATTCT repeat expansion causes SCA10 must await the elucidation of SCA10 function, the correlation argues against the possibility that the expansion represents a rare non-pathogenic polymorphism. Moreover, we found no evidence of the ATTCT expansion in 562 normal chromosomes, including 242 chromosomes from Mexicans individuals. On this basis, and the scale of the expansion observed, we propose that this unstable pentanucleotide repeat expansion is likely to be the disease-causing mutation in SCA10.
At present, we should consider both loss of function (haploinsufficiency) and gain of function as possible pathogenic mechanisms in SCA10. Sca10 is expressed widely in mammalian brain (Fig. 5), consistent with the phenotype of ataxia and epilepsy, and expressed sequence tags representing Sca10 are also detectable in several non-neuronal tissues. The ATTCT repeat is near the 3′ end of the large (>66 kb) intron in SCA10, so the large expansion might affect transcription or post-transcriptional processing of SCA10. We have documented transcription silencing by a large intronic repeat expansion in Friedreich ataxia, in which an expanded GAA repeat interferes with transcription of the gene FRDA (ref. 16). Our preliminary northern-blot data, however, show no obvious changes in the level of SCA10 mRNA in lymphoblastoid cell lines of SCA10 patients (data not shown). It will be important to examine SCA10 mRNA levels in affected tissues of SCA10 patients; however, these tissues are currently not available. The ATTCT expansion might also affect genes other than SCA10 in cis or in trans, similarly to DM1, in which an unstable CTG repeat expands up to several thousand copies in the 3′ untranslated region of the gene DMPK (ref. 17). The discovery of the pentanucleotide expansion in SCA10 patients defines a new class of mutations that will help determine basic mechanisms of microsatellite instability and their link to human diseases.
The Sca10 mRNA was detected by in situ hybridization of radiolabelled probes to horizontal sections of 4-month adult (a,b,c,d) and 10-day juvenile (e) mouse brain. The Sca10 expression pattern was similar to that of the cell density as determined by cresyl violet staining of the same sections (not shown). a–d, Dorsal to ventral progression; f, negative control for non-specific hybridization to an adult brain section.
Methods
DNA extraction and polymorphism analysis.
We extracted DNA from blood samples of all participating family members after obtaining informed consent. To define recombination events and construct haplotypes, we analysed genomic DNA by PCR amplification of polymorphic simple sequence repeat (SSR) markers using end-labelled primers. We determined the size of PCR products on 6% denaturing polyacrylamide gels.
RED analysis.
We performed the RED analysis as described14,18, using genomic DNA samples of SCA10 patients and control subjects devoid of CAG/CTG expansions at the ERDA1 (ref. 19) and SEF2.1 (ref. 20) loci. After 400 temperature cycles (95 °C for 10 s; 65 °C for 30 s) of genomic DNA (10 μg) and (CTG)10 or (TTG)10 oligonucleotides in the presence of Ampligase (5 U; Epicentre), the products were separated by electrophoresis and transferred to a nylon membrane. We hybridized the membrane with a 32P-end-labelled (CAG)10 or (CAA)10 oligonucleotide probe to detect tandemly ligated (CTG)10 or (TTG)10 oligonucleotides.
Western-blot analysis.
We separated proteins extracted from lymphoblastoid cell lines derived from SCA10 patients and control subjects by gel electrophoresis. After western-blot analysis, we used the monoclonal antibody 1C2, raised against the TATA-binding protein, to detect a long polyglutamine stretch as described15.
PCR assay of SCA10 repeats.
We amplified the ATTCT repeat region from genomic DNA by PCR, using the primers attct-L (5′–AGAAAACAGATGGCAGAATGA–3′) and attct-R (5′–GCCTGGGCAACATAGAGAGA–3′) in HotStarTaq Master Mix (Qiagen) including 10% DMSO. The PCR conditions consisted of an initial denaturing at 95 °C for 15 min, 30 PCR cycles (94 °C for 30 s, 54 °C for 30 s, 72 °C for 45 s), and additional extension at 72 °C for 7 min. We estimated the number of the pentanucleotide repeats based on ladders of cloned segments of normal DNA containing 9–20 repeats, which had been sequenced.
Southern-blot analysis.
EcoRI-digested genomic DNA (10 μg) was separated on a 0.8% agarose gel and transferred to a Hybond N+ membrane (Amersham). We generated an 800-bp SCA10 probe located upstream to the pentanucleotide repeat by PCR, using DNA of a PAC clone (RP1-37M3) as the template and primers DanL (5′–TCCTTCCTCAGTCTTTCTGG–3′) and DanR (5′–TGCCATCTGTTTTCTATTTG–3′). Using that probe, random-prime-labelled with 32P-α-dCTP (Amersham), we hybridized the membrane in Church buffer (0.1 mM EDTA at pH 8.0, 0.5 M sodium phosphate at pH 7.2, 7% SDS) at 60 °C overnight, washed in 2×SSC (60 °C for 5 min), 1×SSC (60 °C for 20 min) and 0.5×SSC (60 °C for 20 min) with 0.1% SDS and analysed by autoradiography.
Sequencing.
We amplified the ATTCT repeat region from genomic DNA obtained from normal individuals by PCR using primers attct-L and attct-R (Fig. 3c), then sequenced the PCR products using an Applied Biosystems (ABI) automated sequencer with fluororescent dideoxynucleotides.
In situ hybridization.
We fixed horizontal brain sections (12 μm) from male C57BL/6J mice in 4% paraformaldehyde in PBS followed by dehydration. We end-labelled antisense oligonucleotide probes using terminal deoxynucleotidyl transferase (Promega) and [α-35S] dATP (1250 Ci/mmol; NEN) to a specific activity of ∼109 dpm/μg. Hybridization solution contained formamide (50%, v/v), 4×SSC, sodium phosphate (25 mM), sodium pyrophosphate (1 mM), dextran sulphate (10%, w/v), 5×Denhardt's solution, sonicated herring sperm DNA (200 μg/ml; Promega), polyadenylic acid [5′] (100 μg/ml; Sigma-Aldrich) and [α-35S] dATP-labelled probe (5×102 d.p.m.). To monitor background hybridization, we added a 100-fold excess of unlabelled oligonucleotide to control sections. The sequences of the 45-mer probes were as follows: mE46A, 5′–CTGTTGTCTTCAGTGAGATTTCGCACAGCATACACCACCCACTGC–3′; mE46B, 5′–CACTGCAGAGATGAGAGGTCCGTGAGATGGAATCTGAATGTGTTC–3′. We hybridized the sections with the probes overnight at 42 °C, washed in 1×SSC (22 °C, 20 min), 0.3×SSC (55 °C, 40 min) and 2×SSC (22 °C, 5 min), then dehydrated and exposed to a Kodak BioMax MR film for 1 week.
GenBank accession numbers.
SCA10, 9956851; E46, AL050282; sequences of the 45-mer in situ hybridization probes, mE46A and mE46B, X61506 and AI836854, respectively; PAC37M3, Z84478.
References
Matsuura, T. et al. Mapping of the gene for a novel spinocerebellar ataxia with pure cerebellar signs and epilepsy. Ann. Neurol. 45, 407–411 (1999).
Zu, L., Figueroa, K.P., Grewal, R. & Pulst, S.M. Mapping of a new autosomal dominant spinocerebellar ataxia to chromosome 22. Am. J. Hum. Genet. 64, 594–599 (1999).
Orr, H.T. et al. Expansion of an unstable trinucleotide CAG repeat in spinocerebellar ataxia type 1. Nature Genet. 4, 221–226 (1993).
Pulst, S.M. et al. Moderate expansion of a normally biallelic trinucleotide repeat in spinocerebellar ataxia type 2. Nature Genet. 14, 269–276 (1996).
Sanpei, K. et al. Identification of the spinocerebellar ataxia type 2 gene using a direct identification of repeat expansion and cloning technique, DIRECT. Nature Genet. 14, 277–284 (1996).
Imbert, G. et al. Cloning of the gene for spinocerebellar ataxia 2 reveals a locus with high sensitivity to expanded CAG/glutamine repeats. Nature Genet. 14, 285–291 (1996).
Kawaguchi, Y. et al. CAG expansions in a novel gene for Machado-Joseph disease at chromosome 14q32.1. Nature Genet. 8, 221–228 (1994).
David, G. et al. Cloning of the SCA7 gene reveals a highly unstable CAG repeat expansion. Nature Genet. 17, 65–70 (1997).
Zhuchenko, O. et al. Autosomal dominant cerebellar ataxia (SCA6) associated with small polyglutamine expansions in the α 1A-voltage-dependent calcium channel. Nature Genet. 15, 62–69 (1997).
Holmes, S.E. et al. Expansion of a novel CAG trinucleotide repeat in the 5′ region of P2R2B is associated with SCA12. Nature Genet. 23, 391–392 (1999).
Koob, M.D. et al. An untranslated CTG expansion causes a novel form of spinocerebellar ataxia (SCA8). Nature Genet. 21, 379–384 (1999).
Matsuura, T., Watase, K., Nagamitsu, S., Zoghbi, H.Y. & Ashizawa, T. Fine mapping of the spinocerebellar ataxia type 10 region and search for a polyglutamine expansion. Ann. Neurol. 46, 480 (1999).
Dunham, I. et al. The DNA sequence of human chromosome 22. Nature 402, 489–495 (1999).
Schalling, M., Hudson, T.J., Buetow, K.H. & Housman, D.E. Direct detection of novel expanded trinucleotide repeats in the human genome. Nature Genet. 4, 135–139 (1993).
Trottier, Y. et al. Polyglutamine expansion as a pathological epitope in Huntington's disease and four dominant cerebellar ataxias. Nature 378, 403–406 (1995).
Bidichandani, S.I., Ashizawa, T. & Patel, P.I. The GAA triplet-repeat expansion in Friedreich ataxia interferes with transcription and may be associated with an unusual DNA structure. Am. J. Hum. Genet. 62, 111–121 (1998).
Wells, R.D. & Warren, S.T. Genetic Instability and Hereditary Neurological Diseases (Academic, San Diego, 1998).
Koob, M.D. et al. Rapid cloning of expanded trinucleotide repeat sequences from genomic DNA. Nature Genet. 18, 72–75 (1998).
Ikeuchi, T. et al. A novel long and unstable CAG/CTG trinucleotide repeat on chromosome 17q. Genomics 49, 321–326 (1998).
Breschel, T.S. et al. A novel, heritable, expanding CTG repeat in an intron of the SEF2-1 gene on chromosome 18q21.1. Hum. Mol. Genet. 6, 1855–1863 (1997).
Acknowledgements
We thank the patients for cooperation; L.P.W. Ranum for confirming RED analysis results; and The Sanger Centre for BAC, PAC and cosmid clones located in the chromosome 22 region of interest. This work was supported by grants from the Oxnard Foundation/National Ataxia Foundation (T.A.), National Ataxia Foundation (S.M.P.), Howard Hughes Medical Institute (H.Y.Z.), the National Institute of Health/NICHD HD29256 (D.L.N.), MRRC HD24064, NINDS NS27699 and GCRCM01RR00188 (H.Y.Z.), NS29709 (J.L.N.), NS33123 and NS37883 (S.M.P.), and K12-AG0052-01 (R.P.G.). The fellowship of T.M. was partly supported by the Cell Science Research Foundation, the Yamanouchi Foundation for Research on Metabolic Disorders and the Nakayama Foundation for Human Science, Japan. D.L.B. was supported by an American Epilepsy Society Fellowship.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Matsuura, T., Yamagata, T., Burgess, D. et al. Large expansion of the ATTCT pentanucleotide repeat in spinocerebellar ataxia type 10. Nat Genet 26, 191–194 (2000). https://doi.org/10.1038/79911
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/79911
This article is cited by
-
Consensus Paper: Strengths and Weaknesses of Animal Models of Spinocerebellar Ataxias and Their Clinical Implications
The Cerebellum (2022)
-
Spinocerebellar Ataxia Type 10 with Atypical Clinical Manifestation in Han Chinese
The Cerebellum (2022)
-
Intellectual disability genomics: current state, pitfalls and future challenges
BMC Genomics (2021)