Abstract
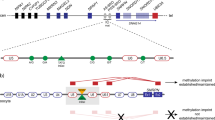
Imprinting in the 15q11–q13 region involves an ‘imprinting centre’ (IC), mapping in part to the promoter and first exon of SNRPN. Deletion of this IC abolishes local paternally derived gene expression and results in Prader-Willi syndrome (PWS). We have created two deletion mutations in mice to understand PWS and the mechanism of this IC. Mice harbouring an intragenic deletion in Snrpn are phenotypically normal, suggesting that mutations of SNRPN are not sufficient to induce PWS. Mice with a larger deletion involving both Snrpn and the putative PWS-IC lack expression of the imprinted genes Zfp127 (mouse homologue of ZNF127), Ndn and Ipw, and manifest several phenotypes common to PWS infants. These data demonstrate that both the position of the IC and its role in the coordinate expression of genes is conserved between mouse and human, and indicate that the mouse is a suitable model system in which to investigate the molecular mechanisms of imprinting in this region of the genome.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Holm, V.A. et al. Prader-Willi syndrome: consensus diagnostic criteria. Pediatrics 91, 398–402 (1993).
Clayton-Smith, J. & Pembrey, M.E. Angelman syndrome. Med. Genet. 29, 412–415 (1992).
Nicholls, R.D. et al. Restriction fragment length polymorphisms within proximal 15q and their use in molecular cytogenetics and the Prader-Willi syndrome. Am. J. Med. Genet. 33, 66–77 (1989).
Robinson, W.P. et al. Molecular, cytogenetic, and clinical investigations of Prader-Willi syndrome patients. Am. J. Hum. Genet. 49, 1219–1234 (1991).
Mascari, M.J. et al. The frequency of uniparental disomy in Prader-Willi syndrome. Implications for molecular diagnosis. N. Engl. J. Med. 326, 1599–1607 (1992).
Nicholls, R.D., Knoll, J.H., Butler, M.G., Karam, S. & Lalande, M. Genetic imprinting suggested by maternal heterodisomy in nondeletion Prader-Willi syndrome. Nature 342, 281–285 (1989).
Knoll, J.H. et al. Angelman and Prader-Willi syndromes share a common chromosome 15 deletion but differ in parental origin of the deletion. Am. J. Med. Genet. 32, 285–290 (1989).
Magenis, R.E. et al. Comparison of the 15q deletions in Prader-Willi and Angelman syndromes: specific regions, extent of deletions, parental origin, and clinical consequences. Am. J. Med. Genet. 35, 333–349 (1990).
Williams, C.A. et al. Maternal origin of 15q11-13 deletions in Angelman syndrome suggests a role for genomic imprinting. Am. J. Med. Genet. 35, 350–353 (1990).
Malcolm, S. et al. Uniparental paternal disomy in Angelman's syndrome. Lancet 337, 694–697 (1991).
Nicholls, R.D., Pai, G.S., Gottlieb, W. & Cantu, E.S. Paternal uniparental disomy of chromosome 15 in a child with Angelman syndrome. Ann. Neurol. 32, 512–518 (1992).
Kishino, T., Lalande, M. & Wagstaff, J. UBE3A/E6-AP mutations cause Angelman syndrome. Nature Genet. 15, 70–73 (1997).
Matsuura, T. et al. De novo truncating mutations in E6-AP ubiquitin-protein ligase gene (UBE3A) in Angelman syndrome. Nature Genet. 15, 74–77 (1997).
Nicholls, R.D. New insights reveal complex mechanisms involved in genomic imprinting. Am. J. Hum. Genet. 54, 733–740 (1994).
Ozcelik, T. et al. Small nuclear ribonucleoprotein polypeptide N (SNRPN), an expressed gene in the Prader-Willi syndrome critical region. Nature Genet. 2, 265–269 (1992).
Glenn, C.C., Porter, K.A., Jong, M.T., Nicholls, R.D. & Driscoll, D.J. Functional imprinting and epigenetic modification of the human SNRPN gene. Hum. Mol. Genet. 2, 2001–2005 (1993).
Reed, M.L. & Leff, S.E. Maternal imprinting of human SNRPN, a gene deleted in Prader-Willi syndrome. Nature Genet. 6, 163–167 (1994).
MacDonald, H.R. & Wevrick, R. The necdin gene is deleted in Prader-Willi syndrome and is imprinted in human and mouse. Hum. Mol. Genet. 6, 1873–1878 (1997).
Wevrick, R., Kerns, J.A. & Francke, U. Identification of a novel paternally expressed gene in the Prader-Willi syndrome region. Hum. Mol. Genet. 3, 1877–1882 (1994).
Sutcliffe, J.S. et al. Deletions of a differentially methylated CpG island at the SNRPN gene define a putative imprinting control region. Nature Genet. 8, 52–58 (1994).
Reis, A. et al. Imprinting mutations suggested by abnormal DNA methylation patterns in familial Angelman and Prader-Willi syndromes. Am. J. Hum. Genet. 54, 741–747 (1994).
Buiting, K. et al. Inherited microdeletions in the Angelman and Prader-Willi syndromes define an imprinting centre on human chromosome 15. Nature Genet. 9, 395–400 (1995).
Saitoh, S. et al. Minimal definition of the imprinting center and fixation of chromosome 15q11-q13 epigenotype by imprinting mutations. Proc. Natl. Acad. Sci. USA 93, 7811–7815 (1996).
Hermann, H. et al. snRNP Sm proteins share two evolutionary conserved sequence motifs whiah are involved in Sm protein-protein interactions. EMBO J. 14, 2076–2088 (1995).
Szabo, P. & Mann, J.R. Expression and methylation of imprinted genes during in vitro differentiation of mouse parthenogenetic and androgenetic embryonic stem cell lines. Development 120, 1651–1660 (1994).
Hogan, B., Beddington, R., Constantini, F. & Lacy, E. Manipulating the mouse embryo. A laboratory manual. (Cold Spring Harbor Laboratory Press, Cold Spring Harbor, 1994).
Steitz, J.A. et al. Function of the abundant U-snRNPs. In Structure and Function of Major and Minor Small Nuclear Ribonuclear Ribonucleoprotein Particles (ed. Birnstiel, M.L.) 115–154 (Springer-Verlag, New York, 1990).
Lerner, M.C. & Steitz, J.A. Antibodies to small nuclear RNAs complexed with proteins are produced by patients with systemic lupus erythematosis. Proc. Natl. Acad. Sci. USA 76, 5495–5499 (1979).
McAllister, G., Amara, S.G. & Lerner, M.R. Tissue-specific expression and cDNA cloning of small nuclear ribonucleoprotein-associated polypeptide N. Proc. Natl. Acad. Sci. USA 85, 5296–5300 (1988).
McAllister, G., Roby-Shemkovitz, A., Amara, S.G. & Lerner, M.R. cDNA sequence of the rat U snRNP-associated protein N: Description of a potential Sm epitope. EMBO J. 8, 1177–1181 (1989).
Li, S., Klein, E.S., Russo, A.F., Simmons, D.M. & Rosenfeld, M.G. Isolation of cDNA clones encoding small nuclear ribonucleoparticle-associated proteins with different tissue specificities. Proc. Natl. Acad. Sci. USA 86, 9778–9782 (1989).
Schmauss, C. & Lerner, M.R. The closely related small nuclear ribonucleoprotein polypeptides N and B/B′ are distinguishable by antibodies as well as by differences in their mRNAs and gene structure. J. Biol. Chem. 265, 10733–10739 (1990).
Schmauss, C., Brines, M.L. & Lerner, M.R. The gene encoding the small nuclear ribonucleoprotein-associated protein N is expressed at high levels in neurons. J. Biol. Chem. 267, 8521–8529 (1992).
Sun, Y. et al. Breakage in the Snrpn locus in a balanced 46,XY,t(15;19) Prader-Willi syndrome patient. Hum. Mol. Genet. 5, 517–524 (1996).
Maruyama, K., Usami, M., Aizawa, T. & Yoshikawa, K. A novel brain-specific mRNA encoding nuclear protein (Necdin) expressed in neurally differentiated embryonal carcinoma cells. Biochem. Biophys. Res. Comm. 178, 291–296 (1991).
Uetsuki, T., Takagi, K., Sugiura, H. & Yoshikawa, K. Structure and expression of the mouse Necdin gene. J. Biol. Chem. 271, 918–924 (1996).
Cattanach, B.M. et al. A candidate mouse model for Prader-Willi syndrome which shows an absence of Snrpn expression. Nature Genet. 2, 270–274 (1992).
Dittrich, B. et al. Imprint switching on human chromosome 15 may involve alternative transcripts of the SNRPN gene. Nature Genet. 14, 163–170 (1996).
Swiatek, P.J. & Gridley, T. Perinatal lethality and defects in hindbrain development in mice homozygous for a targeted mutation of the zinc finger gene Krox20. Genes Dev. 7, 2071–2084 (1993).
Robertson, E.J. Embryo-derived stem cells. In Teratocarcinomas and Embryonic Stem Cells. (ed. Robertson, E.J.) 71–112 (IRL Press, Oxford, 1987).
Laird, P.W. et al. Simplified mammalian DNA isolation procedure. Nucleic Acids Res. 19, 4293–4294 (1991).
Church, G.M. & Gilbert, W. The genomic sequencing technique. Prog. Clin. Biol. Res. 177, 17–21 (1985).
Rubenstein, J.L. et al. Subtractive hybridization system using single-stranded phagemids with directional inserts. Nucleic Acids Res. 18, 4833–4842 (1990).
Wevrick, R. & Francke, U. An imprinted mouse transcript homologous to the human Imprinted in Prader-Willi syndrome (IPW) gene. Hum. Mol. Genet. 6, 325–332 (1997).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Yang, T., Adamson, T., Resnick, J. et al. A mouse model for Prader-Willi syndrome imprinting-centre mutations. Nat Genet 19, 25–31 (1998). https://doi.org/10.1038/ng0598-25
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/ng0598-25
This article is cited by
-
Novel epigenetic molecular therapies for imprinting disorders
Molecular Psychiatry (2023)
-
Genomic imprinted genes in reciprocal hybrid endosperm of Brassica napus
BMC Plant Biology (2021)
-
The diverse roles of DNA methylation in mammalian development and disease
Nature Reviews Molecular Cell Biology (2019)
-
Increased alternate splicing of Htr2c in a mouse model for Prader-Willi syndrome leads disruption of 5HT2C receptor mediated appetite
Molecular Brain (2016)
-
Maintaining memory of silencing at imprinted differentially methylated regions
Cellular and Molecular Life Sciences (2016)