Abstract
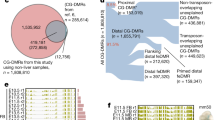
Mammalian development requires cytosine methylation, a heritable epigenetic mark of cellular memory believed to maintain a cell's unique gene expression pattern. However, it remains unclear how dynamic DNA methylation relates to cell type–specific gene expression and animal development. Here, by mapping base-resolution methylomes in 17 adult mouse tissues at shallow coverage, we identify 302,864 tissue-specific differentially methylated regions (tsDMRs) and estimate that >6.7% of the mouse genome is variably methylated. Supporting a prominent role for DNA methylation in gene regulation, most tsDMRs occur at distal cis-regulatory elements. Unexpectedly, some tsDMRs mark enhancers that are dormant in adult tissues but active in embryonic development. These 'vestigial' enhancers are hypomethylated and lack active histone modifications in adult tissues but nevertheless exhibit activity during embryonic development. Our results provide new insights into the role of DNA methylation at tissue-specific enhancers and suggest that epigenetic memory of embryonic development may be retained in adult tissues.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Accession codes
References
Feng, S., Jacobsen, S.E. & Reik, W. Epigenetic reprogramming in plant and animal development. Science 330, 622–627 (2010).
Jones, P.A. Functions of DNA methylation: islands, start sites, gene bodies and beyond. Nat. Rev. Genet. 13, 484–492 (2012).
Bird, A. DNA methylation patterns and epigenetic memory. Genes Dev. 16, 6–21 (2002).
Bonasio, R., Tu, S. & Reinberg, D. Molecular signals of epigenetic states. Science 330, 612–616 (2010).
Okano, M. & Li, E. Genetic analyses of DNA methyltransferase genes in mouse model system. J. Nutr. 132, 2462S–2465S (2002).
Meissner, A. Epigenetic modifications in pluripotent and differentiated cells. Nat. Biotechnol. 28, 1079–1088 (2010).
Polo, J.M. et al. A molecular roadmap of reprogramming somatic cells into iPS cells. Cell 151, 1617–1632 (2012).
Ball, M.P. et al. Targeted and genome-scale strategies reveal gene-body methylation signatures in human cells. Nat. Biotechnol. 27, 361–368 (2009).
Lister, R. et al. Human DNA methylomes at base resolution show widespread epigenomic differences. Nature 462, 315–322 (2009).
Stadler, M.B. et al. DNA-binding factors shape the mouse methylome at distal regulatory regions. Nature 480, 490–495 (2011).
Okano, M., Bell, D.W., Haber, D.A. & Li, E. DNA methyltransferases Dnmt3a and Dnmt3b are essential for de novo methylation and mammalian development. Cell 99, 247–257 (1999).
Hemberger, M., Dean, W. & Reik, W. Epigenetic dynamics of stem cells and cell lineage commitment: digging Waddington's canal. Nat. Rev. Mol. Cell Biol. 10, 526–537 (2009).
Irizarry, R.A. et al. The human colon cancer methylome shows similar hypo- and hypermethylation at conserved tissue-specific CpG island shores. Nat. Genet. 41, 178–186 (2009).
Hansen, K.D. et al. Increased methylation variation in epigenetic domains across cancer types. Nat. Genet. 43, 768–775 (2011).
Ehrlich, M. et al. Amount and distribution of 5-methylcytosine in human DNA from different types of tissues of cells. Nucleic Acids Res. 10, 2709–2721 (1982).
Novakovic, B. et al. DNA methylation–mediated down-regulation of DNA methyltransferase-1 (DNMT1) is coincident with, but not essential for, global hypomethylation in human placenta. J. Biol. Chem. 285, 9583–9593 (2010).
Schroeder, D.I. et al. The human placenta methylome. Proc. Natl. Acad. Sci. USA 110, 6037–6042 (2013).
Gilbert, S. Developmental Biology (Sinauer Associates, Sunderland, MA, 2000).
Lister, R. et al. Hotspots of aberrant epigenomic reprogramming in human induced pluripotent stem cells. Nature 471, 68–73 (2011).
Xie, W. et al. Base-resolution analyses of sequence and parent-of-origin dependent DNA methylation in the mouse genome. Cell 148, 816–831 (2012).
Hon, G.C. et al. Global DNA hypomethylation coupled to repressive chromatin domain formation and gene silencing in breast cancer. Genome Res. 22, 246–258 (2012).
Berman, B.P. et al. Regions of focal DNA hypermethylation and long-range hypomethylation in colorectal cancer coincide with nuclear lamina–associated domains. Nat. Genet. 44, 40–46 (2012).
Peric-Hupkes, D. et al. Molecular maps of the reorganization of genome–nuclear lamina interactions during differentiation. Mol. Cell 38, 603–613 (2010).
Hiratani, I. et al. Genome-wide dynamics of replication timing revealed by in vitro models of mouse embryogenesis. Genome Res. 20, 155–169 (2010).
Heintzman, N.D. et al. Histone modifications at human enhancers reflect global cell-type-specific gene expression. Nature 459, 108–112 (2009).
Heintzman, N.D. et al. Distinct and predictive chromatin signatures of transcriptional promoters and enhancers in the human genome. Nat. Genet. 39, 311–318 (2007).
Rada-Iglesias, A. et al. A unique chromatin signature uncovers early developmental enhancers in humans. Nature 470, 279–283 (2011).
Shen, Y. et al. A map of the cis-regulatory sequences in the mouse genome. Nature 488, 116–120 (2012).
Hawkins, R.D. et al. Dynamic chromatin states in human ES cells reveal potential regulatory sequences and genes involved in pluripotency. Cell Res. 21, 1393–1409 (2011).
Creyghton, M.P. et al. Histone H3K27ac separates active from poised enhancers and predicts developmental state. Proc. Natl. Acad. Sci. USA 107, 21931–21936 (2010).
Siepel, A. et al. Evolutionarily conserved elements in vertebrate, insect, worm, and yeast genomes. Genome Res. 15, 1034–1050 (2005).
Brawand, D. et al. The evolution of gene expression levels in mammalian organs. Nature 478, 343–348 (2011).
Heinz, S. et al. Simple combinations of lineage-determining transcription factors prime cis-regulatory elements required for macrophage and B cell identities. Mol. Cell 38, 576–589 (2010).
Visel, A. et al. ChIP-seq accurately predicts tissue-specific activity of enhancers. Nature 457, 854–858 (2009).
McLean, C.Y. et al. GREAT improves functional interpretation of cis-regulatory regions. Nat. Biotechnol. 28, 495–501 (2010).
Reichert, H. Evolutionary conservation of mechanisms for neural regionalization, proliferation and interconnection in brain development. Biol. Lett. 5, 112–116 (2009).
Blow, M.J. et al. ChIP-Seq identification of weakly conserved heart enhancers. Nat. Genet. 42, 806–810 (2010).
Taberlay, P.C. et al. Polycomb-repressed genes have permissive enhancers that initiate reprogramming. Cell 147, 1283–1294 (2011).
Visel, A., Minovitsky, S., Dubchak, I. & Pennacchio, L.A. VISTA Enhancer Browser—a database of tissue-specific human enhancers. Nucleic Acids Res. 35, D88–D92 (2007).
Hallonet, M., Hollemann, T., Pieler, T. & Gruss, P. Vax1, a novel homeobox-containing gene, directs development of the basal forebrain and visual system. Genes Dev. 13, 3106–3114 (1999).
Muzio, L. et al. Emx2 and Pax6 control regionalization of the pre-neuronogenic cortical primordium. Cereb. Cortex 12, 129–139 (2002).
ENCODE Project Consortium. A user's guide to the encyclopedia of DNA elements (ENCODE). PLoS Biol. 9, e1001046 (2011).
Thurman, R.E. et al. The accessible chromatin landscape of the human genome. Nature 489, 75–82 (2012).
Wu, H. et al. Dnmt3a-dependent nonpromoter DNA methylation facilitates transcription of neurogenic genes. Science 329, 444–448 (2010).
Brinkman, A.B. et al. Sequential ChIP–bisulfite sequencing enables direct genome-scale investigation of chromatin and DNA methylation cross-talk. Genome Res. 22, 1128–1138 (2012).
Bocker, M.T. et al. Genome-wide promoter DNA methylation dynamics of human hematopoietic progenitor cells during differentiation and aging. Blood 117, e182–e189 (2011).
Doi, A. et al. Differential methylation of tissue- and cancer-specific CpG island shores distinguishes human induced pluripotent stem cells, embryonic stem cells and fibroblasts. Nat. Genet. 41, 1350–1353 (2009).
Feinberg, A.P. & Vogelstein, B. Hypomethylation distinguishes genes of some human cancers from their normal counterparts. Nature 301, 89–92 (1983).
Monk, M., Boubelik, M. & Lehnert, S. Temporal and regional changes in DNA methylation in the embryonic, extraembryonic and germ cell lineages during mouse embryo development. Development 99, 371–382 (1987).
Gama-Sosa, M.A. et al. The 5-methylcytosine content of DNA from human tumors. Nucleic Acids Res. 11, 6883–6894 (1983).
Herman, J.G. et al. Silencing of the VHL tumor-suppressor gene by DNA methylation in renal carcinoma. Proc. Natl. Acad. Sci. USA 91, 9700–9704 (1994).
ENCODE Project Consortium. An integrated encyclopedia of DNA elements in the human genome. Nature 489, 57–74 (2012).
Bestor, T., Laudano, A., Mattaliano, R. & Ingram, V. Cloning and sequencing of a cDNA encoding DNA methyltransferase of mouse cells. The carboxyl-terminal domain of the mammalian enzymes is related to bacterial restriction methyltransferases. J. Mol. Biol. 203, 971–983 (1988).
Langmead, B., Trapnell, C., Pop, M. & Salzberg, S.L. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol. 10, R25 (2009).
Rajagopal, N. et al. RFECS: a random-forest based algorithm for enhancer identification from chromatin state. PLoS Comput. Biol. 9, e1002968 (2013).
Maurano, M.T. et al. Systematic localization of common disease-associated variation in regulatory DNA. Science 337, 1190–1195 (2012).
Meyer, L.R. et al. The UCSC Genome Browser database: extensions and updates 2013. Nucleic Acids Res. 41, D64–D69 (2013).
Dixon, J.R. et al. Topological domains in mammalian genomes identified by analysis of chromatin interactions. Nature 485, 376–380 (2012).
Schug, J. et al. Promoter features related to tissue specificity as measured by Shannon entropy. Genome Biol. 6, R33 (2005).
Acknowledgements
We thank S. Kuan, Z. Ye and L. Edsall for their assistance in sequencing and the initial processing of sequencing reads. This work is funded by the Ludwig Institute for Cancer Research and the US National Institutes of Health (R01 HG003991 and ES017166).
Author information
Authors and Affiliations
Contributions
G.C.H., N.R. and F.Y. performed bioinformatics analysis. G.C.H., Y.S., D.F.M. and M.D.D. performed experiments. G.C.H. and B.R. prepared the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–8 and Supplementary Note (PDF 1743 kb)
Supplementary Table 1
tsDMRs in mouse tissues (XLSX 16827 kb)
Supplementary Table 2
Motif analysis of tsDMRs not recovered by known regulatory elements (XLSX 20 kb)
Supplementary Table 3
AD-A and AD-I tsDMRs in mouse tissues (XLSX 12898 kb)
Supplementary Table 4
Overlap of embryonic enhancers with adult AD-I tsDMRs (XLSX 11 kb)
Supplementary Table 5
Predicted enhancers (XLSX 16979 kb)
Supplementary Table 6
Known motifs used in analysis (XLSX 15 kb)
Rights and permissions
About this article
Cite this article
Hon, G., Rajagopal, N., Shen, Y. et al. Epigenetic memory at embryonic enhancers identified in DNA methylation maps from adult mouse tissues. Nat Genet 45, 1198–1206 (2013). https://doi.org/10.1038/ng.2746
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.2746
This article is cited by
-
DNA methylation haplotype block signatures responding to Staphylococcus aureus subclinical mastitis and association with production and health traits
BMC Biology (2024)
-
DNA methylation restricts coordinated germline and neural fates in embryonic stem cell differentiation
Nature Structural & Molecular Biology (2024)
-
Cross-tissue patterns of DNA hypomethylation reveal genetically distinct histories of cell development
BMC Genomics (2023)
-
Co-effects of m6A and chromatin accessibility dynamics in the regulation of cardiomyocyte differentiation
Epigenetics & Chromatin (2023)
-
The methylome and cell-free DNA: current applications in medicine and pediatric disease
Pediatric Research (2023)