Abstract
Cell-type diversity in multicellular organisms is created through a series of binary cell fate decisions. Lateral inhibition controlled by Delta–Notch signalling is the core mechanism for the choice of alternative cell types by homogeneous neighbouring cells. Here, we show that cells engineered with a Delta–Notch-dependent lateral inhibition circuit spontaneously bifurcate into Delta-positive and Notch-active cell populations. The synthetic lateral inhibition circuit comprises transcriptional repression of Delta and intracellular feedback of Lunatic fringe (Lfng). The Lfng-feedback subcircuit, even alone, causes the autonomous cell-type bifurcation. Furthermore, the ratio of two cell populations bifurcated by lateral inhibition is reproducible and robust against perturbation. The cell-type ratio is adjustable by the architecture of the lateral inhibition circuit as well as the degree of cell–cell attachment. Thus, the minimum lateral inhibition mechanism between adjacent cells not only serves as a binary cell-type switch of individual cells but also governs the cell-type ratio at the cell-population level.
Similar content being viewed by others
Introduction
Diverse cell types in a multicellular organism arise from a single fertilized egg. This fact indicates that cell-type diversification occurs repeatedly throughout development. An important mechanism for creating a difference among initially homogeneous cells is lateral inhibition, a mutual inhibition mechanism between adjacent cells that is mediated by Delta–Notch signalling1,2,3,4. Both the ligand Delta and its receptor Notch are transmembrane proteins, and ligand–receptor binding between adjacent cells causes activation and subsequent cleavage of Notch. The released Notch intracellular domain (NICD) acts as a transcriptional regulator, leading to the downregulation of Delta expression. Thus, production of Delta in one cell represses the transcription of Delta in the adjacent cells. The mutual transcriptional repression of Delta between adjacent cells is called lateral inhibition, which is equivalent to an intercellular (double-negative) positive feedback loop. As such, a cell–cell positive feedback loop has the ability to amplify a small initial difference between cells5,6,7,8, initially homogeneous adjacent cells can become radically different, eventually settling down in a pair of stable steady states: a Delta-on/Notch-off state and Delta-off/Notch-on state. Then, the cells with distinct Delta–Notch states adopt different cell fates. A prominent example of the binary cell fate decision through lateral inhibition is the specification of bristle precursor cells in initially homogeneous epithelial cells in Drosophila3,4,9,10,11. In vertebrates, the lateral inhibition mechanism also governs cell fate decisions between neurons and non-neuronal cells in the central nervous system3,12,13,14, secretory cells and absorptive cells in the intestine15,16,17,18, and hair cells and supporting cells in the inner ear3,19,20.
As these mechanisms and roles of lateral inhibition have been revealed mainly by disruption experiments, where the cell fate decision processes were disturbed by inhibition or overexpression of the components of Delta–Notch signaling, it has been difficult to identify sufficient conditions for lateral inhibition to create a stable difference between cells. For example, lateral inhibition by Delta–Notch signalling often accompanies asymmetric cell division (that is, unequal distribution of regulatory molecules between daughter cells)21,22. Whether lateral inhibition can create an enough difference without the help of asymmetric cell division has been unclear. In this study, inspired by the lateral inhibition mechanism in vivo, we aim to create a minimum genetic circuit that would cause spontaneous cell-type bifurcation between adjacent cells. We demonstrate that our synthetic lateral inhibition circuit, which comprises only four genetic components, not only gives rise to two distinct cell types but also maintains an appropriate ratio of the two cell types at the cell-population level without any external cues.
Results
Design and construction of lateral inhibition circuits
All of the genetic circuits were constructed in Chinese hamster ovary (CHO) cells, which do not have a native lateral inhibition mechanism. CHO cells do not express endogenous Delta and express only a small amount of endogenous Notch23. We first constructed a basic transcriptional repression (bTR) circuit (Fig. 1a). When Delta binds to weakly expressed endogenous Notch in the adjacent cells, an NICD-responsive promoter, PTP1, is activated24 (Supplementary Fig. 1a). Under the control of PTP1, the expression of an artificial transcriptional repressor, tTS, is induced. The induced tTS binds to a tTS-responsive promoter, PTetO, and then represses the expression of Delta, resulting in mutual transcriptional repression of Delta between adjacent cells (Fig. 1a). The expression levels of Delta and tTS were monitored by fluorescent reporter constructs joined by a self-cleaving 2A peptide25, Delta-2A-mCherry and tTS-2A-GFP. Thus, Delta-positive cells should exhibit red fluorescence, whereas Notch-active cells (that is, tTS-positive cells) should exhibit green fluorescence. However, cells engineered with the bTR circuit (hereafter called bTR cells) did not bifurcate into two distinct cell types but remained red (that is, a Delta-positive and tTS-negative state; Fig. 1b,c), indicating that the bTR circuit does not meet conditions for cell-type bifurcation.
(a) A scheme for the bTR circuit. Binding of Delta to endogenous Notch between adjacent cells induces the transcription of the tTS repressor from the TP1 promoter, PTP1. Then, tTS represses the transcription of Delta from the TetO promoter, PTetO. (b) A typical image of a bTR cell clone. The entire cell population was Delta-positive and tTS-negative. Scale bar, 100 μm. (c) A unimodal distribution of the bTR cells. The cells were analysed by FACS for GFP and mCherry fluorescence. (d) A scheme for the aTRLF circuit, which comprises the aTR circuit and the LF circuit. In the aTR circuit, an exogenous Notch gene was introduced. In addition, cell clones that showed a higher induction level of tTS were selected. In the LF circuit, activation of Notch induces the transcription of Lfng from the PTP1, and Lfng in turn enhances the activation of Notch. An additional negative effect of Lfng on Delta was shown in this study. (e) A typical image of an aTRLF cell clone. The cell population contained Delta-positive cells and tTS-positive cells. Scale bar, 100 μm. (f) A bimodal distribution of the aTRLF cells. The cells were analysed by FACS for GFP and mCherry fluorescence. (g) A scheme for cell-type bifurcation between adjacent cells. Initially, two cells have similar expression levels of the components of the lateral inhibition circuit, such as Delta, tTS and Lfng. If a slight difference in these expression levels arises from noise, the difference is self-amplified by the lateral inhibition circuit.
Thus, we modified the bTR circuit and constructed an advanced transcriptional repression (aTR) circuit (Fig. 1d, green arrows). To increase the amount of tTS that is induced on activation of Notch, we first introduced an exogenous Notch gene expressed under the control of a strong constitutive promoter, PEF1a, because overexpression of the exogenous Notch increased the PTP1 activity in a dose-dependent manner (Supplementary Fig. 1b). The expression level of PEF1a-Notch was roughly comparable to the Notch expression level in neural stem cells that are known to have a native lateral inhibition mechanism3,12,13,14 (Supplementary Fig. 1c). Second, we selected a cell clone with a higher induction level of tTS. Considering the fact that more than 40 arbitrary fluorescence units (AFUs) of the tTS-2A-GFP signal are required to effectively repress the PTetO activity (Supplementary Fig. 1d), we selected cell clones where more than 100 AFU of tTS-2A-GFP were induced from the PTP1 (Supplementary Fig. 1e). To this end, the PTP1-tTS-2A-GFP construct was introduced by an efficient transposon vector, piggyBac26, while other constructs were introduced by a lentiviral vector. In fact, those cell clones that passed the 100 AFU threshold were found to contain more than 10 copies of the PTP1-tTS-2A-GFP construct, while containing only a single copy of the PEF1a-Notch and PTetO-Delta-2A-mCherry construct (Supplementary Fig. 2). As a result of these improvements, a PTetO-luciferase reporter construct showed an 18-fold decrease on activation of Notch (Supplementary Fig. 3a).
To further enhance the tTS induction and PTetO repression, we added a Lfng-feedback (LF) circuit (Fig. 1d, yellow arrows). Lfng is a glycosyltransferase that glycosylates Notch and thereby enhances the binding of Notch to Delta27,28,29,30. In turn, the activation of Notch is known to induce the expression of Lfng31,32. To mimic the intracellular positive feedback relationship of Notch and Lfng in vivo, we introduced a PTP1-Lfng construct. The PTP1-Lfng construct then enhanced the PTP1 activity (Supplementary Fig. 3b), suggesting that the induction of Lfng expression by Notch and the enhancement of Notch activity by Lfng constitute an intracellular positive feedback loop. Thus, the combined genetic circuit of the aTR circuit and LF circuit was named an aTRLF circuit (Fig. 1d).
Cell-type bifurcation among genetically identical cells
Cells engineered with the aTRLF circuit (hereafter called aTRLF cells) spontaneously bifurcated into red cells (that is, Delta-positive and tTS-negative cells) and green cells (that is, Delta-negative and tTS-positive cells) (Fig. 1e–g). Importantly, the aTRLF cells showed a bimodal distribution (Fig. 1f). Yellow cells (that is, Delta-positive and tTS-positive cells) or colourless cells (that is, Delta-negative and tTS-negative cells) were very rare at the steady state (Fig. 1e,f). The expression of Delta was detected only in the red cell population, whereas that of tTS or Lfng was detected only in the green cell population (Supplementary Fig. 3c). Pharmacological inhibition of Delta–Notch signalling converted the bimodal distribution into a unimodal distribution and made all the cells red (Supplementary Fig. 3d), confirming that the formation of the bimodal distribution was mediated by the Delta–Notch signaling. These results suggest that the aTRLF circuit is bistable, whereas the bTR circuit is monostable. Similar unimodal distributions were obtained for other clones of bTR cells (Supplementary Fig. 4a), whereas similar bimodal distributions for other aTRLF cell clones (Supplementary Fig. 4b). It is noteworthy that the ratio of the red cells to green cells varied to some extent among the 30 aTRLF cell clones we isolated, and that a few aTRLF cell clones failed to show a clear bimodal distribution (Supplementary Fig. 4b). We used the aTRLF cell clones that showed a clear bimodal distribution for further analyses.
Time-lapse imaging of proliferating aTRLF cells from a single cell revealed the cell-type bifurcation process in a single-cell colony (Fig. 2a, Supplementary Fig. 5 and Supplementary Movie 1). Red cells dominated as long as the cell number was low. When the cell number reached ~100 cells, some cells started to make a transition to the green state, eventually resulting in a salt-and-pepper pattern of the red cells and green cells (Fig. 2a and Supplementary Fig. 5b). These results indicate that the aTRLF circuit has the ability to cause cell-type bifurcation among a genetically identical cell population.
(a) Time-lapse imaging of cell-type bifurcation in an aTRLF cell colony. A single aTRLF cell, which happened to be green, was seeded. The cell proliferated and then changed the colour according to the total cell number. Scale bar, 100 μm. See Supplementary Movie 1. (b,c) The aTRLF cells were sorted into the red cell fraction and green cell fraction. The red cells and green cells were seeded separately at the same cell densities. (b) Three days after sorting, green cells were detected in the sorted red cell fraction (upper), whereas red cells were detected in the sorted green cell fraction (lower). Scale bar, 50 μm. See Supplementary Movie 2. (c) The sorted cells were re-seeded and analysed by FACS every other day. Six days after sorting, the original cell-type ratio was reproduced in each fraction culture. The ratio of red cells to green cells was defined by the broken grey line.
Robustness of the cell-type ratio in cell population
Using the aTRLF cells, we investigated whether the ratio of red cells to green cells is reproducible, or robust, as robustness of cell-type ratios should be pivotal for the development of multicellular organisms. The red cell fraction and green cell fraction were sorted and cultured separately (Fig. 2b,c). During 3 days of culture, a portion of the sorted red cells made a transition to the green state, and a portion of the sorted green cells made a transition to the red state (Fig. 2b and Supplementary Movie 2). Remarkably, after 6 days a bimodal distribution was fully reproduced in each fraction culture and the cell-type ratios in both cultures were quite similar to each other (Fig. 2c and Supplementary Fig. 6). In a typical aTRLF cell clone that showed a clear bimodal distribution, the cell-type ratio before sorting was red:green=38:62 and the recovered cell-type ratios were red:green=33:67 and 36:64 (Fig. 2c). These results indicate that the lateral inhibition mechanism between adjacent cells governs the ratio of two cell types at the cell-population level, and that the cell-type ratio is robust against temporal perturbation.
Static equilibrium of the cell-type ratio
A robust cell-type ratio at the steady state indicates two possibilities—(1) static equilibrium: at the steady state, the red cells remain red and the green cells remain green; (2) dynamic equilibrium: although the cell-type ratio remains constant, transitions are occurring from the red state to the green state, and vice versa. To discriminate between the two possibilities, we labelled the sorted red cells with a far-red fluorescent marker and mixed the labelled red cells and unlabelled green cells at different ratios (Fig. 3a). Whereas the labelled red cells showed transitions to the green state when the initial mix ratio (red:green) was 10:0, most of the labelled red cells remained red when the mix ratio was 4:6 or 2:8 (Fig. 3a, see the upper right fraction in each fluorescence-activated cellsorting (FACS) plot). Namely, the transition rate from the red state to the green state showed a correlation with the mix ratio of red cells (Fig. 3b, blue bars). Likewise, the transition rate from the green state to the red state showed a correlation with the mix ratio of green cells (Fig. 3b, pink bars). The total transition rate was at a minimum when the mix ratio was 4:6 (Fig. 3b), suggesting that the cells tend not to change their colours when the mix ratio is close to the cell-type ratio at the steady state (according to Fig. 2c, red:green=38:62). Essentially, the same results were obtained when the green cells were labelled instead of the red cells (Supplementary Fig. 7). These results support the static equilibrium model at the steady state, where most, if not all, of the cells tend to remain in their present colour states (Fig. 3c).
(a) The aTRLF cells were sorted by their colours using FACS. The sorted red cells were labelled with a far-red fluorescent marker. The labelled red cells and unlabelled green cells were mixed and cocultured at the indicated ratios. Two days later, the cells were analysed by FACS for GFP and far-red fluorescence. (b) Quantification of the transition rates shown in a. The labelled green cells indicate transitions from the red state to the green state, whereas the unlabelled red cells indicate transitions from the green state to the red state. Data are means+s.e.m. (n=3). (c) A scheme for static equilibrium. When the mix ratio is far from the cell-type ratio at the steady state (according to Fig. 2c, red:green=38:62), the cells change their colours (left and right). Once the cell-type ratio at the steady state is achieved, the cells no longer change their colours (middle).
Temporal dynamics of cell-type bifurcation
Next, to investigate the dynamics of cell-type bifurcation, we created a lateral inhibition circuit with faster fluorescent reporters by fusing the protein destabilizing signal sequence, d2PEST, to green fluorescent protein (GFP) and mCherry (Fig. 4a). The half-life of d2PEST is reported to be 2 h (ref. 33), which is roughly comparable to the half-life of Delta (2.5 h) or that of tTS (0.5 h; Supplementary Fig. 8). As expected, the aTRLF cells with the d2PEST reporters (hereafter called aTRLFd2 cells) showed a bimodal distribution at the steady state (Fig. 4b). When aTRLFd2 cells were forced into the red state by the pharmacological inhibition of Delta–Notch signalling and then released from the inhibition, we observed a yellow cell population that was making a transition to the green state (Fig. 4c,d and Supplementary Movie 3). On the other hand, when the cell-type ratio was at the steady state, most cells did not change their colours even after cell divisions (Fig. 4e and Supplementary Movie 3). This is probably because the CHO cells quickly intermingle after cell divisions and meet multiple new neighbours, whose red/green ratio is roughly constant at the steady state. In such a situation, it should be easier for the cells to remain in the present colour state. We occasionally detected, however, colour flipping events (Fig. 4e, arrowheads), probably because the cell-type ratio of the neighbouring cells may deviate from the constant ratio accidentally and locally, and then the surrounded cell needs to make a transition to the other colour state. These results are consistent with our cell-labelling experiments (Fig. 3).
(a) A scheme for faster fluorescent reporters. A d2PEST sequence was fused to GFP and mCherry to make their half-lives shorter. A nuclear localization signal (NLS) was also fused to mCherry to make microscopy observation easier. (b) The aTRLFd2 cells were analysed by FACS. (c,d) The aTRLFd2 cells were treated with 3 μM DAPT for 2 days. After DAPT was washed out, the cells were seeded at a density of 8 × 105 cells per 35-mm dish. (c) The cells released from the DAPT treatment were analysed by FACS at indicated time points. (d) Time-lapse imaging of the cells released from the DAPT treatment. Arrowheads indicate the cells that are changing their colours. Scale bar, 50 μm. See Supplementary Movie 3, left. (e) Time-lapse imaging of the aTRLFd2 cells at the steady state. The aTRLFd2 cells were seeded at a density of 8 × 105 cells per 35-mm dish. Arrowheads indicate occasional colour flipping events. Scale bar, 100 μm. See Supplementary Movie 3, right. (e) Time-lapse imaging of cell-type bifurcation between two daughter aTRLFd2 cells. A mother cell was seeded on micropatterned cultureware, where a cell-substratum attachment area is a 25-μm square. Scale bar, 10 μm. See Supplementary Movie 4.
Furthermore, using the aTRLFd2 cells we investigated whether our lateral inhibition circuit is able to break ‘symmetry’ even between two daughter cells (Fig. 4f). As daughter cells are born from the same mother cell, they are expected to initially possess a similar gene expression profile. We enhanced cell–cell contacts between adjacent cells by using micropatterned cultureware, where the cell–substratum attachment area is restricted (a 25-μm square) such that the cells within the area cannot separate from each other. Under the condition, we successfully detected symmetry-breaking events: after a mother aTRLFd2 cell divided into two daughter cells, the two cells went through a ‘battling phase’ in which their colours were not decisive, and then one daughter cell became bright red, whereas the other cell became bright green (Fig. 4f, Supplementary Fig. 9 and Supplementary Movie 4). These results suggest that the aTRLF circuit has the ability to amplify a small initial difference between two cells.
The cell-type ratio is adjustable by cell–cell attachment
Thus far, we have shown that the cell-type ratio is constant and robust against perturbation (Figs 2 and 3). We then investigated what alters the cell-type ratio, because the ratio regulated by lateral inhibition often varies across species and tissues. As the time-lapse imaging of aTRLF cells suggested a relationship of the total cell number and the cell-type ratio (Fig. 2a), we altered the cell seeding density (Fig. 5a,b and Supplementary Fig. 10a). When the cells were seeded at a high density, the distribution of the cell population was bimodal and the ratio of red cells to green cells was 32:68 (Fig. 5b, 1.5 × 106 cells). In contrast, when the cell-seeding density was lower, the distribution became unimodal and the proportion of red cells dramatically increased. At the lowest cell density we examined, the proportion of red cells exceeded 90% (Fig. 5b, 5 × 103 cells). A plausible explanation for the ‘density effect’ on the cell-type ratio is that when cell density is low, direct cell–cell contacts are less frequent and, therefore, binding of Delta to Notch between adjacent cells should be less frequent. Without the Delta–Notch binding, PTP1-tTS-2A-GFP is not activated and PTetO-Delta-2A-mCherry is not repressed. As a result, the red cells should become dominant. In support of this hypothesis, pharmacological inhibition of Delta–Notch signalling produced a very similar transition of the cell-type ratio (Supplementary Fig. 10b,c; compare Fig. 5b with Supplementary Fig. 10c).
(a) Density effects on the cell-type ratio. The aTRLF cells were seeded at the indicated densities (cells per 35-mm dish). Two days later, the cells were re-seeded at the same densities to make sure that the system reaches a steady state. Two more days later, the images were taken and the cells were analysed by FACS. Scale bar, 100 μm. (b) The relationship between the cell density and the cell-type ratio. Quantification of the cell-type ratio of the FACS plots shown in (a). (c) Typical morphology of a cell colony of an aTRLF cell clone and that of the corresponding cell clone with N-cadherin overexpression (left). Scale bar, 100 μm. Quantification of the length of cell perimeter (right). Data are means+s.e.m. (n=30). *P<0.01 (Welch’s t-test). (d) The aTRLF cells without or with N-cadherin overexpression were seeded at a density of 8 × 105 cells per 35-mm dish. Two days later, the cells were re-seeded. Two more days later, the images were taken and the cell-type ratios were analysed by FACS. Scale bar, 100 μm.
Next, we attempted to do the opposite: increase the proportion of green cells. To enhance cell–cell adhesion, we introduced an adhesion molecule, N-cadherin, into the aTRLF cells (Supplementary Fig. 11a). Overexpression of N-cadherin under a constitutive promoter, PCAG34, changed CHO cell morphology; the cells became less elongated and the colony appeared more tightly packed (Fig. 5c). The N-cadherin overexpression increased the proportion of green cells from 62% to 73% (Fig. 5d and Supplementary Fig. 11b). Essentially, the same results were obtained when N-cadherin was introduced into other aTRLF cell clones (Supplementary Fig. 11c). We noticed that the cell perimeter was longer in the N-cadherin-expressing cells (Fig. 5c, right). In addition, the N-cadherin-expressing aTRLF cells tended to cluster together even at lower cell densities, while the control aTRLF cells tended to disperse (Supplementary Fig. 11d and Supplementary Movie 5). Thus, N-cadherin may contribute to the increased proportion of green cells (that is, Notch-active cells) by increasing the degree of cell–cell attachment and therefore enhancing the frequency of Delta–Notch signalling. These results indicate that the cell-type ratio in our system is finely adjustable by the degree of cell–cell attachment that is affected by the cell density and cell–cell adhesion.
Effects of the LF circuit on the cell-type ratio
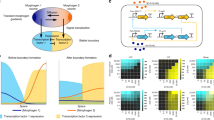
As another means to alter the cell-type ratio, we focused on the architecture of the lateral inhibition circuit, especially on the role of the LF circuit. We first attempted to clarify the effect of Lfng on Delta–Notch signalling, because Lfng has been reported to act as either a positive or negative modulator of the Delta–Notch signalling, depending on the context27,28,29,30,32,35. A series of coculture experiments revealed that when Lfng was expressed only in ‘Receptor cells’, which have Notch and its reporter construct, the presence of Lfng increased the reporter activity (Fig. 6a and Supplementary Fig. 12a). In contrast, when Lfng was expressed also in ‘Ligand cells’, which continuously produce large amounts of Delta, the presence of Lfng decreased the reporter activity of Receptor cells (Fig. 6b and Supplementary Fig. 12b). These results indicate that Lfng has a positive effect on the Notch receptor and a negative effect on the Delta ligand in our system (Fig. 6c). Thus, not only the aTR circuit but also the LF circuit serves as a lateral inhibition mechanism between adjacent cells (Fig. 6c). In addition, the negative effect of Lfng on Delta was dependent on the expression of Notch in the same Ligand cells (Supplementary Fig. 12b) and the positive effect of Lfng on Notch was more prominent in the presence of the Notch expression in the Ligand cells (Supplementary Fig. 12a), suggesting involvement of cis-inhibition (see Discussion).
(a) A positive effect of Lfng on the Notch receptor. CHO cells that have PEF1a-Notch, PTP1-luciferase and PCMV-Lfng constructs (Receptor cells) were cocultured with CHO cells that have PSV40-Delta and PEF1a-Notch constructs (Ligand cells) at a ratio of 1:9. The luciferase activities were measured in the absence or presence of Lfng in the Receptor cells. (b) A negative effect of Lfng on the Delta ligand. CHO cells that have PEF1a-Notch, PTP1-luciferase and PCMV-Lfng constructs (Receptor cells) were cocultured with CHO cells that have PSV40-Delta, PEF1a-Notch and PCMV-Lfng constructs (Ligand cells). The luciferase activities were measured in the absence or the presence of Lfng in the Ligand cells. (c) A scheme for the aTR circuit and a revised scheme for the LF circuit. Lfng has a positive effect on Notch and a negative effect on Delta. Both the aTR circuit and LF circuit constitute intercellular (double-negative) positive feedback loops. (d) Numerical simulations of the mathematical models for the aTR cells and aTRLF cells, with 400 cells in an 8 × 8 grid and a random cell arrangement. The cells within 0.5 radius were considered as the adjacent cells. Hill coefficient n=2. (e) Simulated cell-type ratios shown in d, t=100 h. (f) Experimental verification of ratio alteration by the LF circuit. A cell clone that has only the aTR circuit (aTR cells) was compared with the corresponding cell clone that has both the aTR and LF circuits (aTRLF cells). Scale bar, 100 μm.
We then built a mathematical model that incorporates the dual effects of Lfng (see Methods; Supplementary Table 1), to investigate how the LF circuit affects the cell-type ratio. Numeric simulations of the mathematical model predicted that the proportion of green cells in the aTRLF cells should be higher than that in the cells engineered only with the aTR circuit (hereafter called aTR cells; Fig. 6d,e). To test this predication, we compared an aTR cell clone with the corresponding clone that had been further engineered with the LF circuit (that is, an aTRLF cell clone). The aTR cells showed a bimodal distribution (Fig. 6f, upper), suggesting that the aTR circuit is also bistable. Importantly, the proportion of green cells in the aTRLF cells (68%) was substantially higher than that in the aTR cells (44%; Fig. 6f), in accordance with our prediction. Essentially, the same results were obtained when the LF circuit was added to another aTR cell clone (Supplementary Fig. 13a).
An intuitive explanation of the ratio alteration by the LF circuit is that as the aTRLF circuit contains two intercellular (double-negative) positive feedback loops (that is, the aTR circuit and LF circuit, see Fig. 6c), the aTRLF circuit should be able to create a stable difference among adjacent cells with an increased frequency. Note that the frequency of cell-type bifurcation is reflected in the proportion of green cells; without cell-type bifurcation, the cells remain red (see Fig. 5a, 5 × 103 cells and Supplementary Fig. 3d). Thus, addition of the LF circuit should increase the frequency of cell-type bifurcation and therefore increase the proportion of green cells. In fact, knockout or knockdown of Lfng has been reported to decrease the proportion of Notch-active cells (represented by the green cells in our study) in the developing vertebrate nervous system36,37.
In addition, bifurcation diagrams of the mathematical model indicated that addition of the LF circuit expands the parameter space for bistability (Supplementary Fig. 13b). Namely, the aTRLF circuit should attain bistability more easily than the aTR circuit. In accordance with the prediction, the aTRLF cells tended to show clearer separation of the two cell types in the bimodal distribution as compared with the aTR cells (Supplementary Fig. 13c).
Cell-type bifurcation without the aTR circuit
Although lateral inhibition usually refers to mutual transcriptional repression of Delta between adjacent cells (represented by the aTR circuit in our study), our results indicated that the LF circuit also serves as a lateral inhibition mechanism (Fig. 6c). Thus, we assessed whether the LF circuit alone is capable of cell-type bifurcation. To this end, the aTR circuit was specifically blocked by the tTS inhibitor, Doxycycline (Dox; Fig. 7a). Our mathematical model predicted that the LF circuit should be able to attain bistability even without the aTR circuit (Fig. 7b). In accordance with this prediction, the aTRLF cells showed a bimodal distribution even in the presence of Dox, whereas the aTR cells did not (Fig. 7c and Supplementary Fig. 13d). In the Dox-treated aTRLF cells, the expression of Lfng was detected only in the GFP-positive (that is, Notch-active) cell population (Fig. 7d). These results suggest importance of the LF circuit in cell-type bifurcation.
A scheme for inhibition of the aTR circuit with Dox. Dox inhibits the function of tTS. The LF circuit remains intact. (b) Bifurcation diagrams of the mathematical models for the aTR circuit and aTRLF circuit in the presence of Dox. (c) The aTR cells and aTRLF cells were treated with 30 ng ml−1 Dox. Two days later, the cells were re-seeded and again treated with Dox to make sure that the system reaches a steady state. Two more days later, the cells were analysed by FACS. The aTRLF cells bifurcated into the GFP-negative cells and GFP-positive cells. (d) The GFP-negative aTRLF cells and GFP-positive aTRLF cells shown in c, the upper right panel, were sorted by FACS. The expression level of Lfng was measured in each cell fraction by immunoblotting. The control, Tubulin, was run on a different gel.
Discussion
We previously constructed a genetic circuit for mutual activation of Delta–Notch signalling between adjacent cells to demonstrate signal propagation23. In this study, on the other hand, we aimed to construct a genetic circuit for lateral inhibition between adjacent cells that gives rise to cell-type bifurcation. Among different types of lateral inhibition circuits we constructed, the aTR circuit, LF circuit and aTRLF circuit made adjacent cells spontaneously bifurcate into two distinct cell types, while the bTR circuit was not sufficient for the cell-type bifurcation. The frequency of cell-type bifurcation (that is, the proportion of green cells) was highest in the aTRLF cells. The aTRLF circuit even showed the ability to amplify a small initial difference between two daughter cells.
Although a bistable system usually requires nonlinearity that is derived from cooperative regulatory processes in the genetic circuit, our mathematical model showed that the combination of the aTR circuit and LF circuit enables the aTRLF circuit to attain bistability without assuming any cooperativity (that is, Hill coefficient=1, or lower) in the transcriptional regulations of PTP1 and PTetO (Supplementary Fig. 13b). In fact, activation of PTP1 has been reported not to have significant cooperativity38 and repression of PTetO did not show significant cooperativity in our measurement (Supplementary Fig. 1d). Thus, the combination of the aTR circuit and LF circuit should be beneficial to increase the certainty of cell-type bifurcation.
The sufficiency of the LF circuit alone for cell-type bifurcation was surprising, because lateral inhibition usually refers to the transcriptional repression of Delta (that is, aTR circuit). We have found that Lfng has both positive and negative effects on Delta–Notch signalling in our system. Although the molecular mechanisms by which Lfng exerts the dual effects are unclear, the negative effect of Lfng on Delta was dependent on the expression of Notch in the same cells (Supplementary Fig. 12b). In addition, the two cell types bifurcated by the LF circuit did not differ in the transcription level of Delta (Fig. 7c, see the Delta-2A-mCherry fluorescence) but differed in the activation level of Notch (Fig. 7c, see the tTS-2A-GFP fluorescence). Thus, our working hypothesis is that the negative effect of Lfng may be mediated by cis-inhibition (Supplementary Fig. 14a), which is caused by deprivation of Delta through the cis-binding of Notch to Delta within the same cell11,38,39. Lfng may glycosylate Notch and thereby enhance the binding of Notch to Delta29,30 both in trans (that is, between the adjacent cells) and in cis (that is, within the same cell). The enhancement of trans-binding may cause the positive effect on the Notch receptor, whereas that of cis-binding may cause the negative effect on the Delta ligand.
Lateral inhibition controlled by Delta–Notch signalling is a mechanism that works locally between adjacent cells. In this study, however, the entire cell population engineered with the aTRLF circuit spontaneously bifurcated into two cell types with a particular ratio. Importantly, the cell-type ratio was robust: the original ratio was spontaneously reproduced even after the two cell types were separated and mixed at different ratios. Thus, the lateral inhibition mechanism has the ability to robustly maintain the cell-type ratio in a cell population (Supplementary Fig. 14b). The robust ratio regulation may be able to guarantee the appropriate balance of distinct cell types in tissues even when the tissue homeostasis is temporally disturbed by injuries or biological noise40,41.
The cell-type ratio regulated by lateral inhibition varies widely across species and tissues. The ratio of hair cells to supporting cells in the chick inner ear is ~1:2 (ref. 19), while the ratio of bristle precursor cells to surrounding epithelial cells in Drosophila is ~1:30 (ref. 10). Long-range activation of Delta–Notch signalling through filopodial contacts has been proposed to explain the different cell-type ratio10. Inhibition of Delta–Notch signalling has been shown to alter the ratio of secretory cells to absorptive cells in the mouse intestine, whose normal ratio is ~1:9 (refs 16, 42). In this study, we inquired whether other mechanisms would affect the cell-type ratio. We have shown that the cell-type ratio varies according to the degree of cell–cell attachment and the architecture of the genetic circuit. The proportion of green cells (that is, Notch-active cells) increased in proportion to the higher cell density and the stronger cell–cell adhesion. The LF circuit shifted the cell-type ratio to more Notch-active cells. In good agreement with our results, a knockout mouse of Lfng has been reported to show less Hes1-positive cells (that is, Notch-active cells) and more Ngn2-positive cells (that is, Delta-positive cells) in the developing nervous system36.
Finally, the lateral inhibition circuit described in this study has a potential to recapitulate two other developmentally important phenomena: regular cell pattern formation and synchronized oscillation. Although the spatial arrangement of red cells and green cells shown in this study appeared random, the lateral inhibition mechanism is known to create a more regular salt-and-pepper pattern in several epithelial cell tissues, such as the inner ear1,19,20,39. In fact, our numerical simulation of the aTRLF model with a hexagonal cell arrangement resulted in a regular salt-and-pepper pattern (Supplementary Fig. 14c). On the other hand, the lateral inhibition mechanism is also known to regulate synchronized oscillation in segmentation clock43,44. Thus, the next challenge will be reconstituting a regular salt-and-pepper pattern as well as synchronized oscillation by using the lateral inhibition circuit to understand these self-organization mechanisms of development.
Methods
DNA constructs
The genetic constructs used in this study are listed in Supplementary Table 2. Each promoter and gene were subcloned into pDONR to construct entry vectors according to the manufacturer’s protocol (Invitrogen). These entry clones were recombined with pLenti6.3 (Invitrogen) or piggyBac vector, using the Multisite Gateway technology (Invitrogen)23.
Culture protocols
CHO cells, provided by the RIKEN BRC through the National Bio-Resource Project of the MEXT, Japan, were maintained in DMEM/F12 medium containing 10% fetal bovine serum at 37 °C with 5% CO2. Mouse neural stem cells were purchased from R&D Systems.
Creation of stable cell clones
Viruses for each pLenti construct were created according to the manufacturer’s protocols (Invitrogen). CHO cells were infected with the viruses and maintained in the basal medium plus 10 μg ml−1 blasticidin, 1 μg ml−1 puromycin, 200 μg ml−1 hygromycin or 50 μg ml−1 G418 for the selection of drug-resistant clones23. CHO cells were also transfected with the piggyBac construct and piggyBac transposase plasmid26. To construct the aTRLF circuit, the PEF1a-Notch construct was first introduced by lentiviruses and one cell clone that showed high Notch expression was isolated by limiting dilution. Next, the PTP1-tTS-2A-GFP construct was introduced by piggyBac and the cells that showed a high induction rate of GFP (>100 AFU) on Notch activation were sorted by FACS. This step is the most important criterion in our clone screening. In fact, <5% cells passed this criterion. Finally, the PTetO-Delta-2A-mCherry and PTP1-Lfng constructs were introduced by lentiviruses and cell clones were isolated by limiting dilution.
Luciferase reporter assay
For real-time luciferase assays, CHO cells engineered with each genetic construct were cocultured with Ligand cells at a ratio of 1:9 at a total density of 1 × 106 cells per 35-mm dish. Four hours later, luciferase substrate D-luciferin (0.1 mM) was added to the culture medium. Luciferase activity was recorded once every hour using Kronos Dio Luminometer (Atto). For non-real-time luciferase assays, the cells were cocultured with Ligand cells at a ratio of 1:9 at a density of 3 × 105 cells per well on a 12-well plate. After 40 h of coculture, luciferase activity was measured using a luciferase reporter assay system (Promega).
Image and data analysis
The cells were seeded at a density of 1 × 106 cells per 35-mm dish unless indicated otherwise. Two days later, the cells were re-seeded at the same densities to make sure that the system reaches a steady state. Two more days later, the cells were fixed with 3.7% formaldehyde and images were taken with Axio Observer A1 (Carl Zeiss) or IX73 (Olympus). Time-lapse imaging was performed with an incubator microscope system, LCV110 (Olympus). Data collection and analysis were performed with MetaMorph software (Molecular Devices). To enhance cell–cell contacts between daughter cells, cells were seeded on micropatterned cultureware, CytoGraph (DNP).
FACS analysis and cell sorting
For FACS analyses, the cells were seeded at a density of 1 × 106 cells per 35-mm dish unless indicated otherwise. Two days later, the cells were re-seeded at the same densities to make sure that the system reaches a steady state. Two more days later, the cells were quantified with JSAN (Bay bioscience) or FACSCalibur (BD) and analysed with FlowJo software. Cell sorting was performed with JSAN. The sorted mCherry-positive cells and GFP-positive cells were seeded separately at a density of 6 × 105 cells per 35-mm dish for time-lapse imaging and at a density of 4 × 105 cells per well on a 12-well plate for FACS analyses. For immunoblotting, 1 × 105 cells were sorted.
Cell labelling and transition assessment
The sorted cells were labelled with a far-red fluorescent marker, CellVue Maroon (POL), according to the manufacturer’s protocols. The labelled and unlabelled cells were cocultured at a density of 1.5 × 105 cells per well on a 24-well plate. Two days later, the cells were analysed by FACSCalibur or JSAN for GFP and far-red fluorescence.
Immunoblotting
Immunoblotting was performed by means of standard protocols. The blot was probed with rabbit anti-Myc (A14, Santa Cruz; 1/2,000), mouse anti-α-Tubulin (DM1A, Sigma; 1/20,000), goat anti-Delta (C-20, Santa Cruz; 1/2,000), rat anti-N-cadherin (NCD-2, gift from M. Takeichi; 1/1,000), rabbit anti-Tet-Repressor (TET01, MoBiTec; 1/2,000), or mouse anti-V5 antibody (46-0705, Invitrogen; 1/2,000). Original scans of immunoblottings are shown in Supplementary Fig. 15.
Southern blotting
Southern blotting was performed according to the DIG Application Manual for Filter Hybridization (Roche) using the PCR DIG Probe Synthesis kit (Roche). The probe sequences are available in Supplementary Table 2.
Quantittive RT–PCR
Total RNA was extracted with the RNeasy kit (Qiagen) and reverse transcribed with the QuantiTect Reverse Transcription kit (Qiagen). Real-time PCR was performed using LightCycler 480 II (Roche). The measured value was normalized to glyceraldehyde 3-phosphate dehydrogenase. Primers were designed in the messenger RNA sequence where the parts of the sequences are identical between mouse and hamster. The primer sequences are listed in Supplementary Table 2.
Degradation assay
CHO cells engineered with each genetic construct were treated with cycloheximide (20 μg ml−1). The cells were frozen in liquid nitrogen at the indicated time points. Cell extracts were resolved by immunoblotting. Curve fitting was performed with Excel.
Models
We constructed simple mathematical models that represented the lateral inhibition circuits.
1. Model for aTR cells:






2. Model for aTRLF cells:








The variables are N (Notch), G (glycosylated Notch), D (Delta), S (released NICD), R (tTS repressor), L (Lfng), F (GFP) and C (mCherry). The cis-inhibition model developed by Sprinzak et al.39 was used as a reference of the model for aTR cells. Then, the effect of the LF circuit was added to construct the model for aTRLF cells. The expression of Lfng is induced by the same promoter as tTS (equation (12)). The induced Lfng gylcosylates Notch (equations (7 and 8)). The binding of glycosylated Notch to Delta (both in trans and in cis) is kG-fold stronger than that of non-glycosylated Notch (equations (8, 9 and 10)). The parameter values are listed in Supplementary Table 1. The symbol ‹› denotes the sum of the expression levels in adjacent cells divided by 6. Namely,

where j and j′ are the indices of each cell and its adjacent cells, respectively. In the two cell models (Fig. 7b and Supplementary Fig. 13b), the ‘adjacent’ cell means the other cell. In the multiple cell models with a random cell arrangement (Fig. 6d,e), the adjacent cells mean the cells that are located within a radius of 0.5. In the multiple cell model with a hexagonal cell arrangement (Supplementary Fig. 14c), the adjacent cells mean the six immediate neighbours. The sum of the expression levels in adjacent cells was divided by 6, so that it would be easier to compare the random cell arrangement (Fig. 6d,e) with the hexagonal cell arrangement (Supplementary Fig. 14c). As for the model for the aTR cells and aTRLF cells that were treated with Dox (Fig. 7b), the following equations were used instead of equations (2, 6, 9 and 14).




Bifurcation diagrams of the models with two cells were produced by XPPAUT. Numerical simulations of the models were performed by using Matlab's ode45 solver until the system reaches a steady state.
Additional information
How to cite this article: Matsuda, M. et al. Synthetic lateral inhibition governs cell-type bifurcation with robust ratios. Nat. Commun. 6:6195 doi: 10.1038/ncomms7195 (2015).
References
Collier, J. R., Monk, N. A., Maini, P. K. & Lewis, J. H. Pattern formation by lateral inhibition with feedback: a mathematical model of delta-notch intercellular signalling. J. Theor. Biol. 183, 429–446 (1996).
Bray, S. J. Notch signalling: a simple pathway becomes complex. Nat. Rev. Mol. Cell. Biol. 7, 678–689 (2006).
Lewis, J. Notch signalling and the control of cell fate choices in vertebrates. Semin. Cell Dev. Biol. 9, 583–589 (1998).
Artavanis-Tsakonas, S., Rand, M. D. & Lake, R. J. Notch signaling: cell fate control and signal integration in development. Science 284, 770–776 (1999).
Gardner, T., Cantor, C. & Collins, J. Construction of a genetic toggle switch in Escherichia coli. Nature 403, 339–342 (2000).
Ferrell, J. E. Jr Self-perpetuating states in signal transduction: positive feedback, double-negative feedback and bistability. Curr. Opin. Cell Biol. 14, 140–148 (2002).
Ferrell, J. E. Jr Bistability, bifurcations, and Waddington's epigenetic landscape. Curr. Biol. 22, R458–R466 (2012).
Chau, A. H., Walter, J. M., Gerardin, J., Tang, C. & Lim, W. A. Designing synthetic regulatory networks capable of self-organizing cell polarization. Cell 151, 320–332 (2012).
Heitzler, P. & Simpson, P. The choice of cell fate in the epidermis of Drosophila. Cell 64, 1083–1092 (1991).
Cohen, M., Georgiou, M., Stevenson, N. L., Miodownik, M. & Baum, B. Dynamic filopodia transmit intermittent Delta-Notch signaling to drive pattern refinement during lateral inhibition. Dev. Cell 19, 78–89 (2010).
Barad, O., Rosin, D., Hornstein, E. & Barkai, N. Error minimization in lateral inhibition circuits. Sci. Signal. 3, ra51 (2010).
Kawaguchi, D., Yoshimatsu, T., Hozumi, K. & Gotoh, Y. Selection of differentiating cells by different levels of delta-like 1 among neural precursor cells in the developing mouse telencephalon. Development 135, 3849–3858 (2008).
Pierfelice, T., Alberi, L. & Gaiano, N. Notch in the vertebrate nervous system: an old dog with new tricks. Neuron 69, 840–855 (2011).
Kageyama, R., Ohtsuka, T., Shimojo, H. & Imayoshi, I. Dynamic Notch signaling in neural progenitor cells and a revised view of lateral inhibition. Nat. Neurosci. 11, 1247–1251 (2008).
van Es, J. H. & Clevers, H. Notch and Wnt inhibitors as potential new drugs for intestinal neoplastic disease. Trends Mol. Med. 11, 496–502 (2005).
Vooijs, M., Liu, Z. & Kopan, R. Notch: architect, landscaper, and guardian of the intestine. Gastroenterology 141, 448–459 (2011).
Jensen, J. et al. Control of endodermal endocrine development by Hes-1. Nat. Genet. 24, 36–44 (2000).
Yang, Q., Bermingham, N. A., Finegold, M. J. & Zoghbi, H. Y. Requirement of Math1 for secretory cell lineage commitment in the mouse intestine. Science 294, 2155–2158 (2001).
Goodyear, R. & Richardson, G. Pattern formation in the basilar papilla: evidence for cell rearrangement. J. Neurosci. 17, 6289–6301 (1997).
Chrysostomou, E., Gale, J. E. & Daudet, N. Delta-like 1 and lateral inhibition during hair cell formation in the chicken inner ear: evidence against cis-inhibition. Development 139, 3764–3774 (2012).
Knoblich, J. A. Mechanisms of asymmetric stem cell division. Cell 132, 583–597 (2008).
Zhong, W., Feder, J. N., Jiang, M. M., Jan, L. Y. & Jan, Y. N. Asymmetric localization of a mammalian numb homolog during mouse cortical neurogenesis. Neuron 17, 43–53 (1996).
Matsuda, M., Koga, M., Nishida, E. & Ebisuya, M. Synthetic signal propagation through direct cell-cell interaction. Sci. Signal. 5, ra31 (2012).
Kurooka, H., Kuroda, K. & Honjo, T. Roles of the ankyrin repeats and C-terminal region of the mouse notch1 intracellular region. Nucleic Acids Res. 26, 5448–5455 (1998).
Ryan, M. D. & Drew, J. Foot-and-mouth disease virus 2A oligopeptide mediated cleavage of an artificial polyprotein. EMBO J. 13, 928–933 (1994).
Woltjen, K. et al. piggyBac transposition reprograms fibroblasts to induced pluripotent stem cells. Nature 458, 766–770 (2009).
Moloney, D. J. et al. Fringe is a glycosyltransferase that modifies Notch. Nature 406, 369–375 (2000).
Hicks, C. et al. Fringe differentially modulates Jagged1 and Delta1 signalling through Notch1 and Notch2. Nat. Cell Biol. 2, 515–520 (2000).
Yang, L. T. et al. Fringe glycosyltransferases differentially modulate Notch1 proteolysis induced by Delta1 and Jagged1. Mol. Biol. Cell 16, 927–942 (2005).
Visan, I. et al. Regulation of T lymphopoiesis by Notch1 and Lunatic fringe-mediated competition for intrathymic niches. Nat. Immunol. 7, 634–643 (2006).
Cole, S. E., Levorse, J. M., Tilghman, S. M. & Vogt, T. F. Clock regulatory elements control cyclic expression of Lunatic fringe during somitogenesis. Dev. Cell 3, 75–84 (2002).
Dale, J. K. et al. Periodic notch inhibition by lunatic fringe underlies the chick segmentation clock. Nature 421, 275–278 (2003).
Li, X. et al. Generation of destabilized green fluorescent protein as a transcription reporter. J. Biol. Chem. 273, 34970–34975 (1998).
Niwa, H., Yamamura, K. & Miyazaki, J. Efficient selection for high-expression transfectants with a novel eukaryotic vector. Gene 108, 193–199 (1991).
Okubo, Y. et al. Lfng regulates the synchronized oscillation of the mouse segmentation clock via trans-repression of Notch signalling. Nat. Commun. 3, 1141 (2012).
Kato, T. M., Kawaguchi, A., Kosodo, Y., Niwa, H. & Matsuzaki, F. Lunatic fringe potentiates Notch signaling in the developing brain. Mol. Cell Neurosci. 45, 12–25 (2010).
Nikolaou, N. et al. Lunatic fringe promotes the lateral inhibition of neurogenesis. Development 136, 2523–2533 (2009).
Sprinzak, D. et al. Cis-interactions between Notch and Delta generate mutually exclusive signalling states. Nature 465, 86–90 (2010).
Sprinzak, D., Lakhanpal, A., LeBon, L., Garcia-Ojalvo, J. & Elowitz, M. B. Mutual inactivation of Notch receptors and ligands facilitates developmental patterning. PLoS Comput. Biol. 7, e1002069 (2011).
Balazsi, G., van Oudenaarden, A. & Collins, J. J. Cellular decision making and biological noise: from microbes to mammals. Cell 144, 910–925 (2011).
Eldar, A. & Elowitz, M. B. Functional roles for noise in genetic circuits. Nature 467, 167–173 (2010).
van Es, J. H. et al. Notch/gamma-secretase inhibition turns proliferative cells in intestinal crypts and adenomas into goblet cells. Nature 435, 959–963 (2005).
Lewis, J. Autoinhibition with transcriptional delay: a simple mechanism for the zebrafish somitogenesis oscillator. Curr. Biol. 13, 1398–1408 (2003).
Horikawa, K., Ishimatsu, K., Yoshimoto, E., Kondo, S. & Takeda, H. Noise-resistant and synchronized oscillation of the segmentation clock. Nature 441, 719–723 (2006).
Acknowledgements
We thank J. Miyazaki for the PCAG construct. We also thank I. Wada, T. Shibata, Y. Mukumoto and T. Abe for technical advice, Y. Sakaide for technical assistance and S. Hayashi for critical reading of the manuscript. This work was supported by the following grants to M.E.: the funding from Precursory Research for Embryonic Science and Technology (PRESTO), the Funding Program for Next Generation World-Leading Researchers (NEXT Program), the Special Coordination Fund for Promoting Science and Technology and Grant-in-Aid for Scientific Research on Innovative Areas from Japan Ministry of Education, Culture, Sports, Science and Technology (MEXT).
Author information
Authors and Affiliations
Contributions
M.M. and M.E. designed the study. M.M., M.K. and M.E. performed the experiments and analysed the data. K.W. provided the piggyBac system. M.M., E.N. and M.E. wrote the manuscript. All authors discussed the results and commented on the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
Supplementary Figures 1-15. (PDF 7894 kb)
Supplementary Movie 1
Generation of a salt-and-pepper pattern of red cells and green cells Left: phase, Right: fluorescence. Interval = 1 hour, total time = 205 hours. (MOV 2656 kb)
Supplementary Movie 2
Cell-type bifurcation in sorted cell populations Left: sorted red cell population, Right: sorted green cell population. Interval = 1 hour, total time = 73 hours. (MOV 1035 kb)
Supplementary Movie 3
Dynamics of cell state transitions Left: DAPT washout, Right: w/o DAPT (steady state). Arrowheads indicate transitions. Interval = 15 min, total time = 74 hours 45 min. (MOV 4476 kb)
Supplementary Movie 4
Cell-type bifurcation between two daughter cells Montage of 3 fields. Interval = 0.5 hour, total time = 52 hours. (MOV 1465 kb)
Supplementary Movie 5
Clustering of N-cadherin-expressing cells Left: control CHO cells, Right: N-cadherin-expressing cells. Interval = 15 min, total time = 20 hours. (MOV 1112 kb)
Supplementary Dataset 1
Model parameters (XLSX 12 kb)
Supplementary Dataset 2
List of genetic constructs (XLSX 18 kb)
Rights and permissions
About this article
Cite this article
Matsuda, M., Koga, M., Woltjen, K. et al. Synthetic lateral inhibition governs cell-type bifurcation with robust ratios. Nat Commun 6, 6195 (2015). https://doi.org/10.1038/ncomms7195
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/ncomms7195
This article is cited by
-
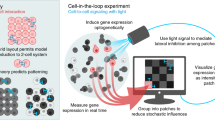
Cell-in-the-loop pattern formation with optogenetically emulated cell-to-cell signaling
Nature Communications (2020)
-
Improvement of the memory function of a mutual repression network in a stochastic environment by negative autoregulation
BMC Bioinformatics (2019)
-
Notch Signaling Mediates Astrocyte Abnormality in Spinal Muscular Atrophy Model Systems
Scientific Reports (2019)
-
Synthetic mammalian pattern formation driven by differential diffusivity of Nodal and Lefty
Nature Communications (2018)
-
Faster embryonic segmentation through elevated Delta-Notch signalling
Nature Communications (2016)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.