Abstract
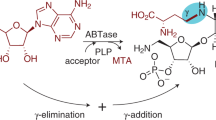
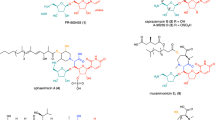
Caprazamycins (CPZs) belong to a group of liponucleoside antibiotics inhibiting the bacterial MraY translocase, an essential enzyme involved in peptidoglycan biosynthesis. We have recently identified analogs that are decorated with a sulfate group at the 2″-hydroxy of the aminoribosyl moiety, and we now report an unprecedented two-step sulfation mechanism during the biosynthesis of CPZs. A type III polyketide synthase (PKS) known as Cpz6 is used in the biosynthesis of a group of new triketide pyrones that are subsequently sulfated by an unusual 3′-phosphoadenosine-5′-phosphosulfate (PAPS)-dependent sulfotransferase (Cpz8) to yield phenolic sulfate esters, which serve as sulfate donors for a PAPS-independent arylsulfate sulfotransferase (Cpz4) to generate sulfated CPZs. This finding is to our knowledge the first demonstration of genuine sulfate donors for an arylsulfate sulfotransferase and the first report of a type III PKS to generate a chemical reagent in bacterial sulfate metabolism.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Chapman, E., Best, M.D., Hanson, S.R. & Wong, C.H. Sulfotransferases: structure, mechanism, biological activity, inhibition, and synthetic utility. Angew. Chem. Int. Edn Engl. 43, 3526–3548 (2004).
Malojčić, G. & Glockshuber, R. The PAPS-independent aryl sulfotransferase and the alternative disulfide bond formation system in pathogenic bacteria. Antioxid. Redox Signal. 13, 1247–1259 (2010).
Kobashi, K., Fukaya, Y., Kim, D.H., Akao, T. & Takebe, S. A novel type of aryl sulfotransferase obtained from an anaerobic bacterium of human intestine. Arch. Biochem. Biophys. 245, 537–539 (1986).
Kaysser, L. et al. Identification and manipulation of the caprazamycin gene cluster lead to new simplified liponucleoside antibiotics and give insights into the biosynthetic pathway. J. Biol. Chem. 284, 14987–14996 (2009).
Igarashi, M. et al. Caprazamycin B, a novel anti-tuberculosis antibiotic, from Streptomyces sp. J. Antibiot. (Tokyo) 56, 580–583 (2003).
Kaysser, L., Siebenberg, S., Kammerer, B. & Gust, B. Analysis of the liposidomycin gene cluster leads to the identification of new caprazamycin derivatives. ChemBioChem 11, 191–196 (2010).
Kaysser, L. et al. A new arylsulfate sulfotransferase involved in liponucleoside antibiotic biosynthesis in streptomycetes. J. Biol. Chem. 285, 12684–12694 (2010).
Gust, B., Challis, G.L., Fowler, K., Kieser, T. & Chater, K.F. PCR-targeted Streptomyces gene replacement identifies a protein domain needed for biosynthesis of the sesquiterpene soil odor geosmin. Proc. Natl. Acad. Sci. USA 100, 1541–1546 (2003).
Kelley, L.A. & Sternberg, M.J. Protein structure prediction on the Web: a case study using the Phyre server. Nat. Protoc. 4, 363–371 (2009).
Moon, A.F. et al. Structural analysis of the sulfotransferase (3-O-sulfotransferase isoform 3) involved in the biosynthesis of an entry receptor for herpes simplex virus 1. J. Biol. Chem. 279, 45185–45193 (2004).
Kakuta, Y., Pedersen, L.G., Pedersen, L.C. & Negishi, M. Conserved structural motifs in the sulfotransferase family. Trends Biochem. Sci. 23, 129–130 (1998).
Hatzios, S.K., Iavarone, A.T. & Bertozzi, C.R. Rv2131c from Mycobacterium tuberculosis is a CysQ 3′-phosphoadenosine-5′-phosphatase. Biochemistry 47, 5823–5831 (2008).
Hatzios, S.K. et al. The Mycobacterium tuberculosis CysQ phosphatase modulates the biosynthesis of sulfated glycolipids and bacterial growth. Bioorg. Med. Chem. Lett. 21, 4956–4959 (2011).
Mechold, U., Ogryzko, V., Ngo, S. & Danchin, A. Oligoribonuclease is a common downstream target of lithium-induced pAp accumulation in Escherichia coli and human cells. Nucleic Acids Res. 34, 2364–2373 (2006).
Pi, N. et al. Kinetic measurements and mechanism determination of Stf0 sulfotransferase using mass spectrometry. Anal. Biochem. 341, 94–104 (2005).
Song, L. et al. Type III polyketide synthase β-ketoacyl-ACP starter unit and ethylmalonyl-CoA extender unit selectivity discovered by Streptomyces coelicolor genome mining. J. Am. Chem. Soc. 128, 14754–14755 (2006).
Funabashi, M., Funa, N. & Horinouchi, S. Phenolic lipids synthesized by type III polyketide synthase confer penicillin resistance on Streptomyces griseus. J. Biol. Chem. 283, 13983–13991 (2008).
Surup, F. et al. The iromycins, a new family of pyridone metabolites from Streptomyces sp. I. Structure, NOS inhibitory activity, and biosynthesis. J. Org. Chem. 72, 5085–5090 (2007).
Funabashi, M. et al. The biosynthesis of liposidomycin-like A-90289 antibiotics featuring a new type of sulfotransferase. ChemBioChem 11, 184–190 (2010).
Moore, B.S. et al. Plant-like biosynthetic pathways in bacteria: from benzoic acid to chalcone. J. Nat. Prod. 65, 1956–1962 (2002).
Austin, M.B. & Noel, J.P. The chalcone synthase superfamily of type III polyketide synthases. Nat. Prod. Rep. 20, 79–110 (2003).
Watanabe, K., Praseuth, A.P. & Wang, C.C. A comprehensive and engaging overview of the type III family of polyketide synthases. Curr. Opin. Chem. Biol. 11, 279–286 (2007).
Abe, I. & Morita, H. Structure and function of the chalcone synthase superfamily of plant type III polyketide synthases. Nat. Prod. Rep. 27, 809–838 (2010).
Yu, D., Xu, F., Zeng, J. & Zhan, J. Type III polyketide synthases in natural product biosynthesis. IUBMB Life 64, 285–295 (2012).
Katsuyama, Y. & Ohnishi, Y. Type III polyketide synthases in microorganisms. Methods Enzymol. 515, 359–377 (2012).
Funa, N. et al. A new pathway for polyketide synthesis in microorganisms. Nature 400, 897–899 (1999).
Funa, N., Ozawa, H., Hirata, A. & Horinouchi, S. Phenolic lipid synthesis by type III polyketide synthases is essential for cyst formation in Azotobacter vinelandii. Proc. Natl. Acad. Sci. USA 103, 6356–6361 (2006).
Aoki, Y., Matsumoto, D., Kawaide, H. & Natsume, M. Physiological role of germicidins in spore germination and hyphal elongation in Streptomyces coelicolor A3(2). J. Antibiot. (Tokyo) 64, 607–611 (2011).
Flett, F., Mersinias, V. & Smith, C.P. High efficiency intergeneric conjugal transfer of plasmid DNA from Escherichia coli to methyl DNA-restricting streptomycetes. FEMS Microbiol. Lett. 155, 223–229 (1997).
Doumith, M. et al. Analysis of genes involved in 6-deoxyhexose biosynthesis and transfer in Saccharopolyspora erythraea. Mol. Gen. Genet. 264, 477–485 (2000).
Acknowledgements
The authors thank G. Challis (University of Warwick) for providing S. coelicolor M145/Δsco7221 and germicidin A. We are grateful to R. Machinek and C. Zolke (Institute of Organic Chemistry, University of Göttingen) for carrying out NMR measurements. We also thank A. Jones for reviewing the manuscript. This work was supported by a grant from the Deutsche Forschungsgemeinschaft (SFB766) to K.E. and X.T., a grant from the Graduate School 'Promotionsverbund Antibakterielle Wirkstoffe' of the University of Tuebingen to X.T. and by the European Commission (IP005224, ActinoGen) to L.K.
Author information
Authors and Affiliations
Contributions
X.T., L.K. and B.G. designed the research. X.T. and L.K. generated and analyzed the mutants. X.T. performed the biochemical experiments and purified presulficidins. X.T. and K.E. purified hydroxyacylcaprazol and purified the proteins. X.T., K.E., L.K. and B.G. analyzed the data. A.K. performed MS analysis. S.G. elucidated the structure of presulficidins. X.T., S.G. and B.G. wrote the manuscript. B.G. supervised the project.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Results, Supplementary Figures 1–15, Supplementary Note and Supplementary Tables 1–3. (PDF 2162 kb)
Rights and permissions
About this article
Cite this article
Tang, X., Eitel, K., Kaysser, L. et al. A two-step sulfation in antibiotic biosynthesis requires a type III polyketide synthase. Nat Chem Biol 9, 610–615 (2013). https://doi.org/10.1038/nchembio.1310
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchembio.1310
This article is cited by
-
β-Hydroxylation of α-amino-β-hydroxylbutanoyl-glycyluridine catalyzed by a nonheme hydroxylase ensures the maturation of caprazamycin
Communications Chemistry (2022)
-
Identification and engineering of 32 membered antifungal macrolactone notonesomycins
Microbial Cell Factories (2020)
-
Production of zosteric acid and other sulfated phenolic biochemicals in microbial cell factories
Nature Communications (2019)
-
Novel Type III Polyketide Synthases Biosynthesize Methylated Polyketides in Mycobacterium marinum
Scientific Reports (2018)
-
Activation of a plasmid-situated type III PKS gene cluster by deletion of a wbl gene in deepsea-derived Streptomyces somaliensis SCSIO ZH66
Microbial Cell Factories (2016)