Abstract
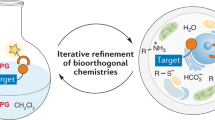
Functional tools are needed to understand complex biological systems. Here we review how chemical reporters in conjunction with bioorthogonal labeling methods can be used to image and retrieve nucleic acids, proteins, glycans, lipids and other metabolites in vitro, in cells as well as in whole organisms. By tagging these biomolecules, researchers can now monitor their dynamics in living systems and discover specific substrates of cellular pathways. These advances in chemical biology are thus providing important tools to characterize biological pathways and are poised to facilitate our understanding of human diseases.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Change history
19 December 2013
In the version of this Review Article initially published, a double bond was missing from the chemical structure of the cyclooctene on the left side of the reaction arrow in Figure 1e. The chemical structure has been corrected in the HTML and PDF versions of the article.
References
Prescher, J.A. & Bertozzi, C.R. Chemistry in living systems. Nat. Chem. Biol. 1, 13–21 (2005).
Sletten, E.M. & Bertozzi, C.R. From mechanism to mouse: a tale of two bioorthogonal reactions. Acc. Chem. Res. 44, 666–676 (2011).
Nomura, D.K., Dix, M.M. & Cravatt, B.F. Activity-based protein profiling for biochemical pathway discovery in cancer. Nat. Rev. Cancer 10, 630–638 (2010).
Salic, A. & Mitchison, T.J. A chemical method for fast and sensitive detection of DNA synthesis in vivo. Proc. Natl. Acad. Sci. USA 105, 2415–2420 (2008). This study describes the first alkyne chemical reporter for labeling nucleic acids in cells and animals.
Jao, C.Y. & Salic, A. Exploring RNA transcription and turnover in vivo by using click chemistry. Proc. Natl. Acad. Sci. USA 105, 15779–15784 (2008).
Neef, A.B. & Luedtke, N.W. Dynamic metabolic labeling of DNA in vivo with arabinosyl nucleosides. Proc. Natl. Acad. Sci. USA 108, 20404–20409 (2011).
Guan, L., van der Heijden, G.W., Bortvin, A. & Greenberg, M.M. Intracellular detection of cytosine incorporation in genomic DNA by using 5-ethynyl-2′-deoxycytidine. ChemBioChem 12, 2184–2190 (2011).
Grammel, M., Hang, H. & Conrad, N.K. Chemical reporters for monitoring RNA synthesis and poly(A) tail dynamics. ChemBioChem 13, 1112–1115 (2012).
Yamakoshi, H. et al. Imaging of EdU, an alkyne-tagged cell proliferation probe, by Raman microscopy. J. Am. Chem. Soc. 27, 6102–6105 (2011). This study describes the direct imaging of alkyne-tagged DNA by Raman spectroscopy.
Song, C.X. et al. Selective chemical labeling reveals the genome-wide distribution of 5-hydroxymethylcytosine. Nat. Biotechnol. 29, 68–72 (2011). This study describes a chemoenzymatic method for bioorthogonal detection and enrichment of 5-hmc modification of nucleic acids.
He, Y.F. et al. Tet-mediated formation of 5-carboxylcytosine and its excision by TDG in mammalian DNA. Science 333, 1303–1307 (2011).
Ito, S. et al. Tet proteins can convert 5-methylcytosine to 5-formylcytosine and 5-carboxylcytosine. Science 333, 1300–1303 (2011).
Liu, C.C. & Schultz, P.G. Adding new chemistries to the genetic code. Annu. Rev. Biochem. 79, 413–444 (2010).
Greiss, S. & Chin, J.W. Expanding the genetic code of an animal. J. Am. Chem. Soc. 133, 14196–14199 (2011).
Bianco, A., Townsley, F.M., Greiss, S., Lang, K. & Chin, J.W. Expanding the genetic code of Drosophila melanogaster. Nat. Chem. Biol. 8, 748–750 (2012).
Lang, K. et al. Genetic encoding of bicyclononynes and trans-cyclooctenes for site-specific protein labeling in vitro and in live mammalian cells via rapid fluorogenic Diels-Alder reactions. J. Am. Chem. Soc. 134, 10317–10320 (2012). This study describes the rapid and selective bioorthogonal reaction in cells using site-specific noncanonical amino acid incorporation and tetrazine reagents.
Johnson, J.A., Lu, Y.Y., Van Deventer, J.A. & Tirrell, D.A. Residue-specific incorporation of non-canonical amino acids into proteins: recent developments and applications. Curr. Opin. Chem. Biol. 14, 774–780 (2010).
Liu, J., Xu, Y., Stoleru, D. & Salic, A. Imaging protein synthesis in cells and tissues with an alkyne analog of puromycin. Proc. Natl. Acad. Sci. USA 109, 413–418 (2012).
Deal, R.B., Henikoff, J.G. & Henikoff, S. Genome-wide kinetics of nucleosome turnover determined by metabolic labeling of histones. Science 328, 1161–1164 (2010).
Eichelbaum, K., Winter, M., Diaz, M.B., Herzig, S. & Krijgsveld, J. Selective enrichment of newly synthesized proteins for quantitative secretome analysis. Nat. Biotechnol. 30, 984–990 (2012).
Howden, A.J. et al. QuaNCAT: quantitating proteome dynamics in primary cells. Nat. Methods 10, 343–346 (2013).
Dieterich, D.C. et al. In situ visualization and dynamics of newly synthesized proteins in rat hippocampal neurons. Nat. Neurosci. 13, 897–905 (2010).
Tcherkezian, J., Brittis, P.A., Thomas, F., Roux, P.P. & Flanagan, J.G. Transmembrane receptor DCC associates with protein synthesis machinery and regulates translation. Cell 141, 632–644 (2010).
Ngo, J.T. et al. Cell-selective metabolic labeling of proteins. Nat. Chem. Biol. 5, 715–717 (2009).
Grammel, M., Zhang, M.M. & Hang, H.C. Orthogonal alkynyl amino acid reporter for selective labeling of bacterial proteomes during infection. Angew. Chem. Int. Ed. Engl. 49, 5970–5974 (2010).
Grammel, M., Dossa, P.D., Taylor-Salmon, E. & Hang, H.C. Cell-selective labeling of bacterial proteomes with an orthogonal phenylalanine amino acid reporter. Chem. Commun. (Camb.) 48, 1473–1474 (2012).
Saxon, E. & Bertozzi, C.R. Cell surface engineering by a modified Staudinger reaction. Science 287, 2007–2010 (2000). This study describes the Staundinger ligation reaction between functionalized triarylphosphine reagents and cell-surface glycans that were metabolically labeled with azide-modified monosaccharide reporter. As triarylphosphine and alkyl azides are both abiotic, this study is the first example of a 'bioorthogonal' ligation reaction.
Laughlin, S.T. & Bertozzi, C.R. Imaging the glycome. Proc. Natl. Acad. Sci. USA 106, 12–17 (2009).
Yarema, K.J., Goon, S. & Bertozzi, C.R. Metabolic selection of glycosylation defects in human cells. Nat. Biotechnol. 19, 553–558 (2001). This study highlights the application of chemical reporter labeling and selection for identification of genetic mutations involved in human disease.
Laughlin, S.T., Baskin, J.M., Amacher, S.L. & Bertozzi, C.R. In vivo imaging of membrane-associated glycans in developing zebrafish. Science 320, 664–667 (2008). This study describes dynamic bioorthogonal labeling and imaging in whole animals.
Dumont, A., Malleron, A., Awwad, M., Dukan, S. & Vauzeilles, B. Click-mediated labeling of bacterial membranes through metabolic modification of the lipopolysaccharide inner core. Angew. Chem. Int. Ed. Engl. 51, 3143–3146 (2012).
Liu, F., Aubry, A.J., Schoenhofen, I.C., Logan, S.M. & Tanner, M.E. The engineering of bacteria bearing azido-pseudaminic acid–modified flagella. ChemBioChem 10, 1317–1320 (2009).
Swarts, B.M. et al. Probing the mycobacterial trehalome with bioorthogonal chemistry. J. Am. Chem. Soc. 134, 16123–16126 (2012).
Anderson, C.T., Wallace, I.S. & Somerville, C.R. Metabolic click-labeling with a fucose analog reveals pectin delivery, architecture, and dynamics in Arabidopsis cell walls. Proc. Natl. Acad. Sci. USA 109, 1329–1334 (2012).
Hart, G.W., Slawson, C., Ramirez-Correa, G. & Lagerlof, O. Cross talk between O-GlcNAcylation and phosphorylation: roles in signaling, transcription, and chronic disease. Annu. Rev. Biochem. 80, 825–858 (2011).
Vocadlo, D.J., Hang, H.C., Kim, E.J., Hanover, J.A. & Bertozzi, C.R. A chemical approach for identifying O-GlcNAc–modified proteins in cells. Proc. Natl. Acad. Sci. USA 100, 9116–9121 (2003).
Zaro, B.W., Yang, Y.Y., Hang, H.C. & Pratt, M.R. Chemical reporters for fluorescent detection and identification of O-GlcNAc–modified proteins reveal glycosylation of the ubiquitin ligase NEDD4–1. Proc. Natl. Acad. Sci. USA 108, 8146–8151 (2011). This study describes a large-scale proteomic analysis of O-GlcNAcylation using chemical reporter labeling.
Rexach, J.E., Clark, P.M. & Hsieh-Wilson, L.C. Chemical approaches to understanding O-GlcNAc glycosylation in the brain. Nat. Chem. Biol. 4, 97–106 (2008).
Rexach, J.E. et al. Quantification of O-glycosylation stoichiometry and dynamics using resolvable mass tags. Nat. Chem. Biol. 6, 645–651 (2010).
Rexach, J.E. et al. Dynamic O-GlcNAc modification regulates CREB-mediated gene expression and memory formation. Nat. Chem. Biol. 8, 253–261 (2012). This study describes a large-scale chemoenzymatic proteomic analysis of dynamic O-GlcNAcylation in the brain.
Pratt, M.R. et al. Deconvoluting the functions of polypeptide N-α-acetylgalactosaminyltransferase family members by glycopeptide substrate profiling. Chem. Biol. 11, 1009–1016 (2004).
Zheng, T. et al. Tracking N-acetyllactosamine on cell-surface glycans in vivo. Angew. Chem. Int. Ed. Engl. 50, 4113–4118 (2011).
Resh, M.D. Trafficking and signaling by fatty-acylated and prenylated proteins. Nat. Chem. Biol. 2, 584–590 (2006).
Hang, H.C., Wilson, J.P. & Charron, G. Bioorthogonal chemical reporters for analyzing protein lipidation and lipid trafficking. Acc. Chem. Res. 44, 699–708 (2011).
Kho, Y. et al. A tagging-via-substrate technology for detection and proteomics of farnesylated proteins. Proc. Natl. Acad. Sci. USA 101, 12479–12484 (2004).
Charron, G., Tsou, L.K., Maguire, W., Yount, J.S. & Hang, H.C. Alkynyl-farnesol reporters for detection of protein S-prenylation in cells. Mol. Biosyst. 7, 67–73 (2011).
DeGraw, A.J. et al. Evaluation of alkyne-modified isoprenoids as chemical reporters of protein prenylation. Chem. Biol. Drug Des. 76, 460–471 (2010).
Charron, G., Li, M., MacDonald, M. & Hang, H.C. Prenylome profiling reveals S-farnesylation is crucial for membrane targeting and antiviral activity of ZAP long-isoform. Proc. Natl. Acad. Sci. USA http://dx.doi.org/10.1073/pnas.1302564110 (2013). This study describes the proteomic analysis of S-prenylated proteins in mammalian cells using chemical reporter labeling and the discovery of zinc-finger antiviral protein L (ZAPL) farnesylation–dependent antiviral activity.
Burnaevskiy, N. et al. Proteolytic elimination of N-myristoyl modifications by the Shigella virulence factor IpaJ. Nature 496, 106–109 (2013).
Fukata, Y. & Fukata, M. Protein palmitoylation in neuronal development and synaptic plasticity. Nat. Rev. Neurosci. 11, 161–175 (2010).
Martin, B.R. & Cravatt, B.F. Large-scale profiling of protein palmitoylation in mammalian cells. Nat. Methods 6, 135–138 (2009). This study describes a large-scale proteomic analysis of S-palmitoylated proteins in T cells using chemical reporter labeling.
Wilson, J.P., Raghavan, A.S., Yang, Y.Y., Charron, G. & Hang, H.C. Proteomic analysis of fatty-acylated proteins in mammalian cells with chemical reporters reveals S-acylation of histone H3 variants. Mol. Cell. Proteomics 10, M110.001198 (2011).
Yount, J.S. et al. Palmitoylome profiling reveals S-palmitoylation–dependent antiviral activity of IFITM3. Nat. Chem. Biol. 6, 610–614 (2010). This study describes the proteomic analysis of S-palmitoylated proteins in a dendritic cell line using chemical reporter labeling and the discovery that S-palmitoylation of IFITM3 is crucial for host resistance to influenza virus infection.
Li, Y., Martin, B.R., Cravatt, B.F. & Hofmann, S.L. DHHC5 protein palmitoylates flotillin-2 and is rapidly degraded on induction of neuronal differentiation in cultured cells. J. Biol. Chem. 287, 523–530 (2012).
Zhang, M.M., Tsou, L.K., Charron, G., Raghavan, A.S. & Hang, H.C. Tandem fluorescence imaging of dynamic S-acylation and protein turnover. Proc. Natl. Acad. Sci. USA 107, 8627–8632 (2010). This study describes dual chemical reporter labeling for the tandem detection of protein turnover and S-palmitoylation dynamics in mammalian cells, which revealed accelerated S-palmitoylation in activated T cells.
Martin, B.R., Wang, C., Adibekian, A., Tully, S.E. & Cravatt, B.F. Global profiling of dynamic protein palmitoylation. Nat. Methods 9, 84–89 (2012). This study describes large-scale profiling of dynamic S-palmitoylation in T cells using chemical reporter labeling with SILAC.
Zhang, C.H., Wu, P.-Y.J., Kelly, F.D., Nurse, P. & Hang, H.C. Quantitative control protein S-palmitoylation regulates meiotic entry in fission yeast. PLoS Biol. 11, e1001501 (2013). This study describes the application of palmitoylation reporter in fission yeast and the discovery that protein S-palmitoylation in cellular differentiation can be quantitatively regulated by a single palmitoyltransferase.
Ching, W., Hang, H.C. & Nusse, R. Lipid-independent secretion of a Drosophila Wnt protein. J. Biol. Chem. 283, 17092–17098 (2008).
Heal, W.P. et al. Bioorthogonal chemical tagging of protein cholesterylation in living cells. Chem. Commun. (Camb.) 47, 4081–4083 (2011).
Jiang, H. et al. SIRT6 regulates TNF-α secretion through hydrolysis of long-chain fatty acyl lysine. Nature 496, 110–113 (2013).
Jao, C.Y., Roth, M., Welti, R. & Salic, A. Metabolic labeling and direct imaging of choline phospholipids in vivo. Proc. Natl. Acad. Sci. USA 106, 15332–15337 (2009).
Yang, J., Seckute, J., Cole, C.M. & Devaraj, N.K. Live-cell imaging of cyclopropene tags with fluorogenic tetrazine cycloadditions. Angew. Chem. Int. Ed. Engl. 51, 7476–7479 (2012).
Milne, S.B. et al. Capture and release of alkyne-derivatized glycerophospholipids using cobalt chemistry. Nat. Chem. Biol. 6, 205–207 (2010). This study describes the analysis of lipid metabolism in mammalian cells using chemical reporter labeling and cobalt-alkyne chemistry for the capture and release of labeled lipids.
Thiele, C. et al. Tracing fatty acid metabolism by click chemistry. ACS Chem. Biol. 7, 2004–2011 (2012).
Rangan, K.J., Yang, Y.Y., Charron, G. & Hang, H.C. Rapid visualization and large-scale profiling of bacterial lipoproteins with chemical reporters. J. Am. Chem. Soc. 132, 10628–10629 (2010).
Babu, M.M. et al. A database of bacterial lipoproteins (DOLOP) with functional assignments to predicted lipoproteins. J. Bacteriol. 188, 2761–2773 (2006).
Lin, H., Su, X. & He, B. Protein lysine acylation and cysteine succination by intermediates of energy metabolism. ACS Chem. Biol. 7, 947–960 (2012).
Yang, Y.Y., Ascano, J.M. & Hang, H.C. Bioorthogonal chemical reporters for monitoring protein acetylation. J. Am. Chem. Soc. 132, 3640–3641 (2010). This study describes chemical reporters for protein acetylation that function in vitro and in mammalian cells.
Yang, Y.Y., Grammel, M. & Hang, H.C. Identification of lysine acetyltransferase p300 substrates using 4-pentynoyl-coenzyme A and bioorthogonal proteomics. Bioorg. Med. Chem. Lett. 21, 4976–4979 (2011); erratum 21, 6613 (2011).
Arnesen, T. et al. Proteomics analyses reveal the evolutionary conservation and divergence of N-terminal acetyltransferases from yeast and humans. Proc. Natl. Acad. Sci. USA 106, 8157–8162 (2009).
Mukherjee, S., Hao, Y.H. & Orth, K. A newly discovered post-translational modification—the acetylation of serine and threonine residues. Trends Biochem. Sci. 32, 210–216 (2007).
Bao, X., Zhao, Q., Yang, T., Fung, Y.M. & Li, X.D. A chemical probe for lysine malonylation. Angew. Chem. Int. Ed. Engl. 52, 4883–4886 (2013).
Greer, E.L. & Shi, Y. Histone methylation: a dynamic mark in health, disease and inheritance. Nat. Rev. Genet. 13, 343–357 (2012).
Dalhoff, C., Lukinavicius, G., Klimasauskas, S. & Weinhold, E. Direct transfer of extended groups from synthetic cofactors by DNA methyltransferases. Nat. Chem. Biol. 2, 31–32 (2006). This study describes the SAM chemical reporter that can be efficiently used by a DNA methyltransferase.
Peters, W. et al. Enzymatic site-specific functionalization of protein methyltransferase substrates with alkynes for click labeling. Angew. Chem. Int. Ed. Engl. 49, 5170–5173 (2010).
Islam, K., Zheng, W., Yu, H., Deng, H. & Luo, M. Expanding cofactor repertoire of protein lysine methyltransferase for substrate labeling. ACS Chem. Biol. 6, 679–684 (2011).
Wang, R., Zheng, W., Yu, H., Deng, H. & Luo, M. Labeling substrates of protein arginine methyltransferase with engineered enzymes and matched S-adenosyl-L-methionine analogues. J. Am. Chem. Soc. 133, 7648–7651 (2011).
Wang, R. et al. Profiling genome-wide chromatin methylation with engineered posttranslation apparatus within living cells. J. Am. Chem. Soc. 135, 1048–1056 (2013). This study describes the application of a protein methylation reporter in living cells.
Willnow, S., Martin, M., Luscher, B. & Weinhold, E. A selenium-based click AdoMet analogue for versatile substrate labeling with wild-type protein methyltransferases. ChemBioChem 13, 1167–1173 (2012).
Bothwell, I.R. et al. Se-adenosyl-L-selenomethionine cofactor analogue as a reporter of protein methylation. J. Am. Chem. Soc. 134, 14905–14912 (2012).
Gibson, B.A. & Kraus, W.L. New insights into the molecular and cellular functions of poly(ADP-ribose) and PARPs. Nat. Rev. Mol. Cell Biol. 13, 411–424 (2012).
Jiang, H., Kim, J.H., Frizzell, K.M., Kraus, W.L. & Lin, H. Clickable NAD analogues for labeling substrate proteins of poly(ADP-ribose) polymerases. J. Am. Chem. Soc. 132, 9363–9372 (2010). This study describes in vitro chemical reporters for protein ADP-ribosylation.
Shogren-Knaak, M.A., Alaimo, P.J. & Shokat, K.M. Recent advances in chemical approaches to the study of biological systems. Annu. Rev. Cell Dev. Biol. 17, 405–433 (2001).
Yarbrough, M.L. et al. AMPylation of Rho GTPases by Vibrio VopS disrupts effector binding and downstream signaling. Science 323, 269–272 (2009).
Woolery, A.R., Luong, P., Broberg, C.A. & Orth, K. AMPylation: something old is new again. Front Microbiol. 1, 113 (2010).
Müller, M.P. et al. The Legionella effector protein DrrA AMPylates the membrane traffic regulator Rab1b. Science 329, 946–949 (2010).
Tan, Y. & Luo, Z.Q. Legionella pneumophila SidD is a deAMPylase that modifies Rab1. Nature 475, 506–509 (2011).
Grammel, M., Luong, P., Orth, K. & Hang, H.C. A chemical reporter for protein AMPylation. J. Am. Chem. Soc. 133, 17103–17105 (2011). This study describes a chemical reporter for protein AMPylation.
Macek, B., Mann, M. & Olsen, J.V. Global and site-specific quantitative phosphoproteomics: principles and applications. Annu. Rev. Pharmacol. Toxicol. 49, 199–221 (2009).
Allen, J.J. et al. A semisynthetic epitope for kinase substrates. Nat. Methods 4, 511–516 (2007). This study describes the union of a 'bumped' ATP analog and mutant kinase pair with two-step alkylation and antibody detection of specific kinase substrates.
Blethrow, J.D., Glavy, J.S., Morgan, D.O. & Shokat, K.M. Covalent capture of kinase-specific phosphopeptides reveals Cdk1-cyclin B substrates. Proc. Natl. Acad. Sci. USA 105, 1442–1447 (2008).
Paulsen, C.E. et al. Peroxide-dependent sulfenylation of the EGFR catalytic site enhances kinase activity. Nat. Chem. Biol. 8, 57–64 (2012). This study describes the proteomic analysis of sulfenylation with an alkyne chemical probe and discovery of oxidation-enhanced kinase activity.
Blackman, M.L., Royzen, M. & Fox, J.M. Tetrazine ligation: fast bioconjugation based on inverse-electron-demand Diels-Alder reactivity. J. Am. Chem. Soc. 130, 13518–13519 (2008). The tetrazine ligation has afforded a rapid and selective bioorthogonal ligation reaction for live-cell imaging studies.
Devaraj, N.K. & Weissleder, R. Biomedical applications of tetrazine cycloadditions. Acc. Chem. Res. 44, 816–827 (2011). The tetrazine ligation has afforded a rapid and selective bioorthogonal ligation reaction for live-cell imaging studies.
Pezacki, J.P. et al. Chemical contrast for imaging living systems: molecular vibrations drive CARS microscopy. Nat. Chem. Biol. 7, 137–145 (2011).
Chang, P.V., Dube, D.H., Sletten, E.M. & Bertozzi, C.R. A strategy for the selective imaging of glycans using caged metabolic precursors. J. Am. Chem. Soc. 132, 9516–9518 (2010).
Xie, R., Hong, S., Feng, L., Rong, J. & Chen, X. Cell-selective metabolic glycan labeling based on ligand-targeted liposomes. J. Am. Chem. Soc. 134, 9914–9917 (2012).
Meldal, M. & Tornøe, C.W. Cu-catalyzed azide-alkyne cycloaddition. Chem. Rev. 108, 2952–3015 (2008). Studies by the Meldal and Sharpless laboratories demonstrated that azide-alkyne cycloadditions could be accelerated with Cu( I ) catalysts for bioorthogonal labeling applications.
Kolb, H.C., Finn, M.G. & Sharpless, K.B. Click chemistry: diverse chemical function from a few good reactions. Angew. Chem. Int. Ed. Engl. 40, 2004–2021 (2001). Studies by the Meldal and Sharpless laboratories demonstrated azide-alkyne cycloadditions could be accelerated with Cu( I ) catalysts for bioorthogonal labeling applications.
Huisgen, R. 1,3-Dipolar cycloadditions. past and future. Angew. Chem. Int. Ed. Engl. 2, 565–598 (1963). Early studies of cycloaddition reactions by Huisgen and coworkers provided the foundation for future bioorthogonal reactions.
Acknowledgements
We thank K. Rangan and N. Westcott for helpful comments on the manuscript. H.C.H. acknowledges support from Ellison Medical Foundation and US National Institutes of Health–National Institute of General Medical Sciences (1R01GM087544).
Author information
Authors and Affiliations
Contributions
M.G. and H.C.H. wrote this review.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Grammel, M., Hang, H. Chemical reporters for biological discovery. Nat Chem Biol 9, 475–484 (2013). https://doi.org/10.1038/nchembio.1296
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchembio.1296
This article is cited by
-
Label-free detection of biomolecules using inductively coupled plasma mass spectrometry (ICP-MS)
Analytical and Bioanalytical Chemistry (2024)
-
Kinetic resolution of cyclic benzylic azides enabled by site- and enantioselective C(sp3)–H oxidation
Nature Communications (2022)
-
A modification-centric assessment tool for the performance of chemoproteomic probes
Nature Chemical Biology (2022)
-
Click-ExM enables expansion microscopy for all biomolecules
Nature Methods (2021)
-
Bioorthogonal chemistry
Nature Reviews Methods Primers (2021)