Abstract
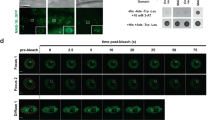
Bicoid (Bcd) is a morphogenetic protein that instructs patterning along the anterior–posterior (A–P) axis in Drosophila melanogaster embryos. Despite extensive studies, what controls the formation of a normal concentration gradient of Bcd remains an unresolved and controversial question. Here, we show that Bcd protein degradation is mediated by the ubiquitin-proteasome pathway. We have identified an F-box protein, encoded by fates-shifted (fsd), that has an important role in Bcd protein degradation by targeting it for ubiquitylation. Embryos from females lacking fsd have an altered Bcd gradient profile, resulting in a shift of the fatemap along the A–P axis. Our study is an experimental demonstration that, contrary to an alternative hypothesis, Bcd protein degradation is required for normal gradient formation and developmental fate determination.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Wolpert, L. Positional information and the spatial pattern of cellular differentiation. J. Theor. Biol 25, 1–47 (1969).
Kerszberg, M. & Wolpert, L. Specifying positional information in the embryo: looking beyond morphogens. Cell 130, 205–209 (2007).
Lander, A. D. Morpheus unbound: reimagining the morphogen gradient. Cell 128, 245–256 (2007).
Martinez Arias, A. & Hayward, P. Filtering transcriptional noise during development: concepts and mechanisms. Nat. Rev. Genet. 7, 34–44 (2006).
Wartlick, O., Kicheva, A. & Gonzalez-Gaitan, M. Morphogen gradient formation. Cold Spring Harb. Perspect. Biol. 1, a001255 (2009).
Ephrussi, A. & St. Johnston, D. Seeing is believing. The bicoid morphogen gradient matures. Cell 116, 143–152 (2004).
Driever, W. & Nüsslein-Volhard, C. A gradient of bicoid protein in Drosophila embryos. Cell 54, 83–93 (1988).
Struhl, G., Struhl, K. & Macdonald, P. The gradient morphogen bicoid is a concentration-dependent transcriptional activator. Cell 57, 1259–1273 (1989).
Driever, W., Thoma, G. & Nüsslein-Volhard, C. Determination of spatial domains of zygotic gene expression in the Drosophila embryo by the affinity of binding site for the bicoid morphogen. Nature 340, 363–367 (1989).
Deng, J., Wang, W., Lu, L. J. & Ma, J. A two-dimensional simulation model of the Bicoid gradient in Drosophila. PLoS ONE 5, e10275 (2010).
Bergmann, S. et al. Pre-steady-state decoding of the Bicoid morphogen gradient. PLoS biology 5, e46 (2007).
Gregor, T., Bialek, W., van Steveninck, R. R., Tank, D. W. & Wieschaus, E. F. Diffusion and scaling during early embryonic pattern formation. Proc. Natl Acad. Sci. USA 102, 18403–18407 (2005).
Gregor, T., Wieschaus, E. F., McGregor, A. P., Bialek, W. & Tank, D. W. Stability and nuclear dynamics of the bicoid morphogen gradient. Cell 130, 141–152 (2007).
Porcher, A. et al. The time to measure positional information: maternal hunchback is required for the synchrony of the Bicoid transcriptional response at the onset of zygotic transcription. Development 137, 2795–2804 (2010).
Zhao, C. et al. The activity of the Drosophila morphogenetic protein Bicoid is inhibited by a domain located outside its homeodomain. Development 129, 1669–1680 (2002).
Fenteany, G. et al. Inhibition of proteasome activities and subunit-specific amino-terminal threonine modification by lactacystin. Science 268, 726–731 (1995).
Meng, L. et al. Epoxomicin, a potent and selective proteasome inhibitor, exhibits in vivo antiinflammatory activity. Proc. Natl Acad. Sci. USA 96, 10403–10408 (1999).
Belle, A., Tanay, A., Bitincka, L., Shamir, R. & O'Shea, E. K. Quantification of protein half-lives in the budding yeast proteome. Proc. Natl Acad. Sci. USA 103, 13004–13009 (2006).
Pickart, C. M. Mechanisms underlying ubiquitination. Annu. Rev. Biochem. 70, 503–533 (2001).
Pickart, C. M. & Eddins, M. J. Ubiquitin: structures, functions, mechanisms. Biochim. Biophys. Acta 1695, 55–72 (2004).
Herrmann, J., Lerman, L. O. & Lerman, A. Ubiquitin and ubiquitin-like proteins in protein regulation. Circ. Res. 100, 1276–1291 (2007).
Ciechanover, A. Proteolysis: from the lysosome to ubiquitin and the proteasome. Nat. Rev. Mol. Cell Biol. 6, 79–87 (2005).
Hershko, A., Ganoth, D., Pehrson, J., Palazzo, R. E. & Cohen, L. H. Methylated ubiquitin inhibits cyclin degradation in clam embryo extracts. J. Biol. Chem. 266, 16376–16379 (1991).
Hsu, T., McRackan, D., Vincent, T. S. & Gert de Couet, H. Drosophila Pin1 prolyl isomerase Dodo is a MAP kinase signal responder during oogenesis. Nat. Cell Biol. 3, 538–543 (2001).
Deshaies, R. J. SCF and Cullin/Ring H2-based ubiquitin ligases. Annu. Rev. Cell Dev. Biol. 15, 435–467 (1999).
Conaway, R. C., Brower, C. S. & Conaway, J. W. Emerging roles of ubiquitin in transcription regulation. Science 296, 1254–1258 (2002).
Muratani, M., Kung, C., Shokat, K. M. & Tansey, W. P. The F box protein Dsg1/Mdm30 is a transcriptional coactivator that stimulates Gal4 turnover and cotranscriptional mRNA processing. Cell 120, 887–899 (2005).
von der Lehr, N. et al. The F-box protein Skp2 participates in c-Myc proteosomal degradation and acts as a cofactor for c-Myc-regulated transcription. Mol. Cell 11, 1189–1200 (2003).
Kornitzer, D., Raboy, B., Kulka, R. G. & Fink, G. R. Regulated degradation of the transcription factor Gcn4. EMBO J. 13, 6021–6030 (1994).
Jiang, J. & Struhl, G. Regulation of the Hedgehog and Wingless signalling pathways by the F-box/WD40-repeat protein Slimb. Nature 391, 493–496 (1998).
Kipreos, E. T. & Pagano, M. The F-box protein family. Genome Biol 1, reviews3002– reviews3002.7 (2000).
Ho, M. S., Tsai, P. I. & Chien, C. T. F-box proteins: the key to protein degradation. J. Biomed. Sci. 13, 181–191 (2006).
Frohnhöfer, H. G. & Nüsslein-Volhard, C. Organization of anterior pattern in the Drosophila embryo by the maternal gene bicoid. Nature 324, 120–125 (1986).
Berleth, T. et al. The role of localization of bicoid RNA in organizing the anterior pattern of the Drosophila embryo. EMBO J. 7, 1749–1756 (1988).
Driever, W., Siegel, V. & Nüsslein-Volhard, C. Autonomous determination of anterior structures in the early Drosophila embryo by the bicoid morphogen. Development 109, 811–820 (1990).
Driever, W. & Nüsslein-Volhard, C. The bicoid protein determines position in the Drosophila embryo in a concentration dependent manner. Cell 54, 95–104 (1988).
Small, S., Kraut, R., Hoey, T., Warrior, R. & Levine, M. Transcriptional regulation of a pair-rule stripe in Drosophila. Genes & Dev. 5, 827–839 (1991).
RiverA–Pomar, R. & Jackle, H. From gradients to stripes in Drosophila embryogenesis: filling in the gaps. Trends Genet. 12, 478–483 (1996).
Driever, W. & Nüsslein-Volhard, C. Bicoid protein is a positive regulator of hunchback transcription in the early Drosophila embryo. Nature 337, 138–143 (1989).
Perkins, T. J., Jaeger, J., Reinitz, J. & Glass, L. Reverse engineering the gap gene network of Drosophila melanogaster. PLoS Comput. Biol. 2, e51 (2006).
Schaeffer, V., Janody, F., Loss, C., Desplan, C. & Wimmer, E. A. Bicoid functions without its TATA-binding protein-associated factor interaction domains. Proc. Natl Acad. Sci. USA 96, 4461–4466 (1999).
Houchmandzadeh, B., Wieschaus, E. & Leibler, S. Establishment of developmental precision and proportions in the early Drosophila embryo. Nature 415, 798–802 (2002).
Crauk, O. & Dostatni, N. Bicoid determines sharp and precise target gene expression in the Drosophila embryo. Curr. Biol. 15, 1888–1898 (2005).
Rivera-Pomar, R., Lu, X., Taubert, H., Perrimon, N. & Jackle, H. Activation of posterior gap gene expression in the Drosophila blastoderm. Nature 376, 253–256 (1995).
He, F. et al. Probing intrinsic properties of a robust morphogen gradient in Drosophila. Dev. Cell 15, 558–567 (2008).
He, F. et al. Shaping a morphogen gradient for positional precision. Biophys. J. 99, 697–707 (2010).
Coppey, M., Berezhkovskii, A. M., Kim, Y., Boettiger, A. N. & Shvartsman, S. Y. Modeling the bicoid gradient: diffusion and reversible nuclear trapping of a stable protein. Dev. Biol. 312, 623–630 (2007).
Spirov, A. et al. Formation of the bicoid morphogen gradient: an mRNA gradient dictates the protein gradient. Development 136, 605–614 (2009).
He, F. et al. Distance measurements via the morphogen gradient of Bicoid in Drosophila embryos. BMC Dev. Biol. 10, 80 (2010).
Jaeger, J. et al. Dynamic control of positional information in the early Drosophila embryo. Nature 430, 368–371 (2004).
Manu et al. Canalization of gene expression in the Drosophila blastoderm by gap gene cross regulation. PLoS biology 7, e1000049 (2009).
Lucchetta, E. M., Vincent, M. E. & Ismagilov, R. F. A precise Bicoid gradient is nonessential during cycles 11–13 for precise patterning in the Drosophila blastoderm. PLoS One 3, e3651 (2008).
Hecht, I., Rappel, W. J. & Levine, H. Determining the scale of the Bicoid morphogen gradient. Proc. Natl Acad. Sci. USA 106, 1710–1715 (2009).
Gregor, T., McGregor, A. P. & Wieschaus, E. F. Shape and function of the Bicoid morphogen gradient in dipteran species with different sized embryos. Dev. Biol. 316, 350–358 (2008).
Grimm, O. & Wieschaus, E. The Bicoid gradient is shaped independently of nuclei. Development 137, 2857–2862 (2010).
Fu, D. & Ma, J. Interplay between positive and negative activities that influence the role of Bicoid in transcription. Nucleic acids research 33, 3985–3993 (2005).
Fu, D., Wen, Y. & Ma, J. The co-activator CREB-binding protein participates in enhancer-dependent activities of Bicoid. J. Biol. Chem. 279, 48725–48733 (2004).
Forler, D. et al. An efficient protein complex purification method for functional proteomics in higher eukaryotes. Nat. Biotechnol. 21, 89–92 (2003).
Crevel, G. & Cotterill, S. DNA replication in cell-free extracts from Drosophila melanogaster. EMBO J. 10, 4361–4369 (1991).
Clemens, J. C. et al. Use of double-stranded RNA interference in Drosophila cell lines to dissect signal transduction pathways. Proc. Natl Acad. Sci. USA 97, 6499–6503 (2000).
Tautz, D. & Pfeifle, C. A non-radioactive in situ hybridization method for the localization of specific RNAs in Drosophila embryos reveals translational control of the segmentation gene hunchback. Chromosoma 98, 81–85 (1989).
Kosman, D. et al. Multiplex detection of RNA expression in Drosophila embryos. Science 305, 846 (2004).
Surkova, S. et al. Characterization of the Drosophila segment determination morphome. Developmental biology 313, 844–862 (2008).
Lattin, J., Carroll, J. D. & Green, P. E. Analyzing multivariate data. (Thompson Books/Cole, 2003).
Acknowledgements
We thank members of our groups at CCHMC, in particular F. He, D. Cheung, W. Dui, and J. Deng, for discussion and assistance, and we thank Xinhua Lin's lab for some of the primers used in our dsRNAi screening. This work was supported in part by grants from NIH and NSF (to J.M.).
Author information
Authors and Affiliations
Contributions
J.L. and J.M. conceived and designed the study. J.L. performed all experiments and analysis. J.L. and J.M. interpreted the data, J.L. generated all figures and J.L. and J.M. wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
Supplementary Information (PDF 5349 kb)
Rights and permissions
About this article
Cite this article
Liu, J., Ma, J. Fates-shifted is an F-box protein that targets Bicoid for degradation and regulates developmental fate determination in Drosophila embryos. Nat Cell Biol 13, 22–29 (2011). https://doi.org/10.1038/ncb2141
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ncb2141
This article is cited by
-
Seeing is believing: the Bicoid protein reveals its path
Hereditas (2018)
-
Noise transmission during the dynamic pattern formation in fly embryos
Quantitative Biology (2018)
-
No preliminary evidence of differences in astrocyte density within the white matter of the dorsolateral prefrontal cortex in autism
Molecular Autism (2017)
-
The novel cannabinoid receptor GPR55 mediates anxiolytic-like effects in the medial orbital cortex of mice with acute stress
Molecular Brain (2017)
-
Synaptic Loss and the Pathophysiology of PTSD: Implications for Ketamine as a Prototype Novel Therapeutic
Current Psychiatry Reports (2017)