Abstract
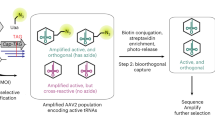
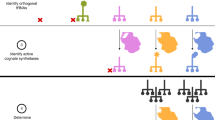
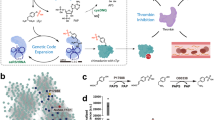
A variety of strategies to incorporate unnatural amino acids into proteins have been pursued1,2,3,4,5, but all have limitations with respect to technical accessibility, scalability, applicability to in vivo studies, or site specificity of amino acid incorporation. The ability to selectively introduce unnatural functional groups into specific sites within proteins, in vivo6,7, provides a potentially powerful approach to the study of protein function and to large-scale production of novel proteins. Here we describe a combined genetic selection and screen that allows the rapid evolution of aminoacyl-tRNA synthetase substrate specificity. Our strategy involves the use of an “orthogonal” aminoacyl-tRNA synthetase and tRNA pair that cannot interact with any of the endogenous synthetase–tRNA pairs in Escherichia coli8,9,10,11. A chloramphenicol-resistance (Cmr) reporter is used to select highly active synthetase variants, and an amplifiable fluorescence reporter is used together with fluorescence-activated cell sorting (FACS) to screen for variants with the desired change in amino acid specificity. Both reporters are contained within a single genetic construct, eliminating the need for plasmid shuttling and allowing the evolution to be completed in a matter of days. Following evolution, the amplifiable fluorescence reporter allows visual and fluorimetric evaluation of synthetase activity and selectivity. Using this system to explore the evolvability of an amino acid binding pocket of a tyrosyl-tRNA synthetase, we identified three new variants that allow the selective incorporation of amino-, isopropyl-, and allyl-containing tyrosine analogs into a desired protein. The new enzymes can be used to produce milligram-per-liter quantities of unnatural amino acid–containing protein in E. coli.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Merrifield, B. Solid phase synthesis. Science 232, 341–347 (1986).
Baca, M. & Kent, S.B.H. Catalytic contribution of flap-substrate hydrogen bonds in HIV-1 protease explored by chemical synthesis. Proc. Natl. Acad. Sci. USA 90, 11638–11642 (1993).
Anthonycahill, S.J., Griffith, M.C., Noren, C.J., Suich, D.J. & Schultz, P.G. Site-specific mutagenesis with unnatural amino acids. Trends Biochem. Sci. 14, 400–403 (1989).
Bain, J.D., Glabe, C.G., Dix, T.A., Chamberlin, A.R. & Diala, E.S. Biosynthetic site-specific incorporation of a non-natural amino acid into a polypeptide. J. Am. Chem. Soc. 111, 8013–8014 (1989).
Budisa, N. et al. Toward the experimental codon reassignment in vivo: protein building with an expanded amino acid repertoire. FASEB J. 13, 41–51 (1999).
Liu, D.R. & Schultz, P.G. Progress toward the evolution of an organism with an expanded genetic code. Proc. Natl. Acad. Sci. USA 96, 4780–4785 (1999).
Wang, L., Brock, A., Herberich, B. & Schultz, P.G. Expanding the genetic code of Escherichia coli. Science 292, 498–500 (2001).
Liu, D.R., Magliery, T.J. & Schultz, P.G. Characterization of an 'orthogonal' suppressor tRNA derived from E. coli tRNA(2)(Gln). Chem. Biol. 4, 685–691 (1997).
Liu, D.R., Magliery, T.J., Pasternak, M. & Schultz, P.G. Engineering a tRNA and aminoacyl-tRNA synthetase for the site-specific incorporation of unnatural amino acids into proteins in vivo. Proc. Natl. Acad. Sci. USA 94, 10092–10097 (1997).
Wang, L., Magliery, T.J., Liu, D.R. & Schultz, P.G. A new functional suppressor tRNA/aminoacyl-tRNA synthetase pair for the in vivo incorporation of unnatural amino acids into proteins. J. Am. Chem. Soc. 122, 5010–5011 (2000).
Pasternak, M., Magliery, T.J. & Schultz, P.G. A new orthogonal suppressor tRNA/aminoacyl-tRNA synthetase pair for evolving an organism with an expanded genetic code. Helv. Chim. Acta 83, 2277–2286 (2000).
Crameri, A., Whitehorn, E.A., Tate, E. & Stemmer, W.P.C. Improved green fluorescent protein by molecular evolution using DNA shuffling. Nat. Biotechnol. 14, 315–319 (1996).
Lorincz, M., Roederer, M., Diwu, Z., Herzenberg, L.A. & Nolan, G.P. Enzyme-generated intracellular fluorescence for single-cell reporter gene analysis utilizing Escherichia coli β-glucuronidase. Cytometry 24, 321–329 (1996).
Zlokarnik, G. et al. Quantitation of transcription and clonal selection of single living cells with β-lactamase as reporter. Science 279, 84–88 (1998).
Guzman, L.M., Belin, D., Carson, M.J. & Beckwith, J. Tight regulation, modulation, and high-level expression by vectors containing the arabinose pBAD promoter. J. Bacteriol. 177, 4121–4130 (1995).
Jeruzalmi, D. & Steitz, T.A. Structure of T7 RNA polymerase complexed to the transcriptional inhibitor T7 lysozyme. EMBO J. 17, 4101–4113 (1998).
Wang, L., Brock, A. & Schultz, P.G. Adding L-3-(2-naphthyl)alanine to the genetic code of E. coli. J. Am. Chem. Soc. 124, 1836–1837 (2002).
Brick, P., Bhat, T.N. & Blow, D.M. Structure of tyrosyl-transfer RNA synthetase refined at 2.3-Å resolution—interaction of the enzyme with the tyrosyl-adenylate intermediate. J. Mol. Biol. 208, 83–98 (1989).
Rath, V.L., Silvian, L.F., Beijer, B., Sproat, B.S. & Steitz, T.A. How glutaminyl-tRNA synthetase selects glutamine. Structure 6, 439–449 (1998).
He, B. et al. Rapid mutagenesis and purification of phage RNA polymerase. Protein Expr. Purif. 9, 142–151 (1997).
Matsoukas, J.M. et al. Differences in backbone structure between angiotensin II agonists and type I antagonists. J. Med. Chem. 38, 4660–4669 (1995).
Nakayama, T., Nomura, M., Haga, K. & Ueda, M. A new three-component photoresist based on calix[4]resorcinarene derivative, a cross-linker, and a photo-acid generator. Bull. Chem. Soc. Jpn. 71, 2979–2984 (1998).
Acknowledgements
We thank Thomas J. Magliery for helpful discussions, Andrew B. Martin for plasmid pBAD/JYAMB-4TAG, and Alan Saluk, Cheryl Silao, and Eric O'Connor of The Scripps Research Institute Flow Cytometry Core Facility. This work was supported by the National Institutes of Health (GM62159), the Department of Energy (DE-FG03-00ER45812), and the Skaggs Institute for Chemical Biology. S.W.S. is a fellow of the Jane Coffin Childs Memorial Fund for Medical Research. This is manuscript number 14972-CH of The Scripps Research Institute.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Rights and permissions
About this article
Cite this article
Santoro, S., Wang, L., Herberich, B. et al. An efficient system for the evolution of aminoacyl-tRNA synthetase specificity. Nat Biotechnol 20, 1044–1048 (2002). https://doi.org/10.1038/nbt742
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt742
This article is cited by
-
Breaking the deadlock in genetic code expansion
Nature Chemical Biology (2024)
-
Adding α,α-disubstituted and β-linked monomers to the genetic code of an organism
Nature (2024)
-
Biocatalysis
Nature Reviews Methods Primers (2021)
-
Engineering a Polyspecific Pyrrolysyl-tRNA Synthetase by a High Throughput FACS Screen
Scientific Reports (2019)
-
Site-specific incorporation of phosphotyrosine using an expanded genetic code
Nature Chemical Biology (2017)