Abstract
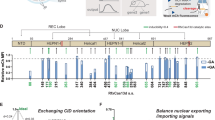
Molecular automata1,2,3 that combine sensing4,5,6, computation7,8,9,10,11,12 and actuation13,14 enable programmable manipulation of biological systems. We use RNA interference (RNAi)15 in human kidney cells to construct a molecular computing core that implements general Boolean logic1,3,8,9,10,11,12,16 to make decisions based on endogenous molecular inputs. The state of an endogenous input is encoded by the presence or absence of 'mediator' small interfering RNAs (siRNAs). The encoding rules, combined with a specific arrangement of the siRNA targets in a synthetic gene network17, allow direct evaluation of any Boolean expression in standard forms using siRNAs and indirect evaluation using endogenous inputs. We demonstrate direct evaluation of expressions with up to five logic variables. Implementation of the encoding rules through sensory up- and down-regulatory links between the inputs and siRNA mediators will allow arbitrary Boolean decision-making using these inputs.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Stojanovic, M.N. & Stefanovic, D. A deoxyribozyme-based molecular automaton. Nat. Biotechnol. 21, 1069–1074 (2003).
Benenson, Y., Gil, B., Ben-Dor, U., Adar, R. & Shapiro, E. An autonomous molecular computer for logical control of gene expression. Nature 429, 423–429 (2004).
Macdonald, J. et al. Medium scale integration of molecular logic gates in an automaton. Nano Lett. 6, 2598–2603 (2006).
Isaacs, F.J. et al. Engineered riboregulators enable post-transcriptional control of gene expression. Nat. Biotechnol. 22, 841–847 (2004).
Bayer, T.S. & Smolke, C.D. Programmable ligand-controlled riboregulators of eukaryotic gene expression. Nat. Biotechnol. 23, 337–343 (2005).
An, C.I., Trinh, V.B. & Yokobayashi, Y. Artificial control of gene expression in mammalian cells by modulating RNA interference through aptamer-small molecule interaction. RNA 12, 710–716 (2006).
Ezziane, Z. DNA computing: applications and challenges. Nanotechnology 17, R27–R39 (2006).
Basu, S. & Weiss, R. The device physics of cellular logic gates. First Workshop on Non-Silicon Computation Boston, February 3 2002 <http://www.cs.cmu.edu/∼phoenix/nsc1/paper/3-2.pdf> 54–61.
Balzani, V., Credi, A. & Venturi, M. Molecular logic circuits. ChemPhysChem 4, 49–59 (2003).
Kramer, B.P., Fischer, C. & Fussenegger, M. BioLogic gates enable logical transcription control in mammalian cells. Biotechnol. Bioeng. 87, 478–484 (2004).
Baron, R., Lioubashevski, O., Katz, E., Niazov, T. & Willner, I. Elementary arithmetic operations by enzymes: A model for metabolic pathway based computing. Angew. Chem. Int. Ed. 45, 1572–1576 (2006).
Seelig, G., Soloveichik, D., Zhang, D.Y. & Winfree, E. Enzyme-free nucleic acid logic circuits. Science 314, 1585–1588 (2006).
Kolpashchikov, D.M. & Stojanovic, M.N. Boolean control of aptamer binding sites. J. Am. Chem. Soc. 127, 11348–11351 (2005).
Beyer, S. & Simmel, F.C. A modular DNA signal translator for the controlled release of a protein by an aptamer. Nucleic Acids Res. 34, 1581–1587 (2006).
Eckstein, F. Small non-coding RNAs as magic bullets. Trends Biochem. Sci. 30, 445–452 (2005).
Penchovsky, R. & Breaker, R.R. Computational design and experimental validation of oligonucleotide-sensing allosteric ribozymes. Nat. Biotechnol. 23, 1424–1433 (2005).
Hasty, J., McMillen, D. & Collins, J.J. Engineered gene circuits. Nature 420, 224–230 (2002).
Wiener, N. Cybernetics (MIT Press, New York, 1948).
Soloveichik, D. & Winfree, E. The computational power of Benenson automata. Theor. Comp. Sci. 244, 279–297 (2005).
Weiss, R., Homsy, G. & Knight, T.F. Toward in vivo digital circuits. in Natural Computing Series (eds. Winfree, E. Landweber, L.F.) 275–295, (Springer, Heidelberg, 2002).
Buchler, N.E., Gerland, U. & Hwa, T. On schemes of combinatorial transcription logic. Proc. Natl. Acad. Sci. USA 100, 5136–5141 (2003).
Isaacs, F.J., Dwyer, D.J. & Collins, J.J. RNA synthetic biology. Nat. Biotechnol. 24, 545–554 (2006).
Laslo, P. et al. Multilineage transcriptional priming and determination of alternate hematopoietic cell fates. Cell 126, 755–766 (2006).
Rosenwald, A. et al. Relation of gene expression phenotype to immunoglobulin mutation genotype in B cell chronic lymphocytic leukemia. J. Exp. Med. 194, 1639–1647 (2001).
Doench, J.G. & Sharp, P.A. Specificity of microRNA target selection in translational repression. Genes Dev. 18, 504–511 (2004).
Schwarz, D.S. et al. Asymmetry in the assembly of the RNAi enzyme complex. Cell 115, 199–208 (2003).
Caron, L. et al. The Lac repressor provides a reversible gene expression system in undifferentiated and differentiated embryonic stem cell. Cell. Mol. Life Sci. 62, 1605–1612 (2005).
Sullivan, C.S. & Ganem, D. A virus-encoded inhibitor that blocks RNA interference in mammalian cells. J. Virol. 79, 7371–7379 (2005).
Yokobayashi, Y., Weiss, R. & Arnold, F.H. Directed evolution of a genetic circuit. Proc. Natl. Acad. Sci. USA 99, 16587–16591 (2002).
Hooshangi, S., Thiberge, S. & Weiss, R. Ultrasensitivity and noise propagation in a synthetic transcriptional cascade. Proc. Natl. Acad. Sci. USA 102, 3581–3586 (2005).
Acknowledgements
We thank A. Murray, E. O'Shea, G. Church, D. Weitz, X. Xie, T. Hwa, C. Queitsch, A. Rivzi, G. Kudla and K. Foster for discussions; I. Benenson for the critical review of the manuscript; anonymous reviewers for insightful comments; and K. Thorn and B. Tilton for technical support. The EF1-α promoter was amplified by PCR from pLEIGW (a gift from Ihor Lemischka, Princeton University). The plasmid pCAGOP containing the synthetic chicken β-actin promoter with lac operators (CAGOP) was obtained from B. Binétruy27. pLV-tTRKRAB-Red was a gift of D. Trono, Ecole Polytechnique Fédérale de Lausanne (EPFL), Switzerland. The work was supported by the Bauer Fellows program and by the National Institute of General Medical Sciences (NIGMS) grant 5P50 GM068763-01. K.R. was partially supported by Harvard's Program for Research in Science and Engineering (PRISE).
Author information
Authors and Affiliations
Contributions
Y.B. and R.W. designed research; Y.B., K.R., L.B., R.M. and S.S. performed research; Y.B., R.W., K.R. and L.B. wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
There is a pending patent application that covers the results presented in this paper.
Supplementary information
Supplementary Fig. 1
Basic notions of the Boolean logic. (PDF 96 kb)
Supplementary Fig. 2
Crosstalk verification between siRNAs and their targets. (PDF 150 kb)
Supplementary Fig. 3
Influence of the secondary structure on siRNA efficiency. (PDF 142 kb)
Supplementary Fig. 4
The histogram of the expression levels. (PDF 63 kb)
Supplementary Fig. 5
Alternative sensory module designs. (PDF 54 kb)
Supplementary Table 1
Association between the literals and the siRNAs. (PDF 34 kb)
Supplementary Table 2
Sequences of the oligonucleotides employed in the circuit construction. (PDF 69 kb)
Rights and permissions
About this article
Cite this article
Rinaudo, K., Bleris, L., Maddamsetti, R. et al. A universal RNAi-based logic evaluator that operates in mammalian cells. Nat Biotechnol 25, 795–801 (2007). https://doi.org/10.1038/nbt1307
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt1307
This article is cited by
-
Synthetic morphology with agential materials
Nature Reviews Bioengineering (2023)
-
Robust and tunable signal processing in mammalian cells via engineered covalent modification cycles
Nature Communications (2022)
-
A synthetic circuit for buffering gene dosage variation between individual mammalian cells
Nature Communications (2021)
-
A biological multiplexer, designs, and simulations
The Journal of Supercomputing (2021)
-
Design and evaluation of biological gate circuits and their therapeutic applications in a model of multidrug resistant cancers
Biotechnology Letters (2020)